Abstract
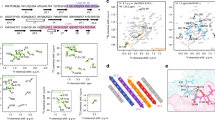
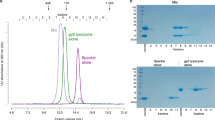
POLYOMA virus is a small oncogenic virus containing a circular DNA the identification and location of defined regions of which provides physical reference points which are important for correlation in genetic and biochemical studies. The protein coded by gene 32 of bacteriophage T4 binds tightly to single-stranded DNA and RNA and facilitates denaturation and renaturation of double-stranded DNA1,2. Using glutaraldehyde fixation gene 32 protein denaturation of double-stranded DNA can be observed using electron microscopy at physiological temperatures and denaturation maps are produced which closely resemble those obtained with heat and alkaline denaturation2. Use of gene 32 protein in binding experiments with circular sutpercoiled DNA molecules from polyoma virus and simian virus 40 (SV40) has already shown that only one denaturation loop is formed per molecule2,4 reflecting possible structural constraints in the DNA. The characterisation and mapping of the preferred denaturation site(s)4 is also of interest since these putative A + T-rich region(s) are likely to be related to the binding sites of enzymes using DNA as a template. In the present study the use of two different specific restriction enzymes (endonucleases) has enabled us to identify five regions of polyoma DNA which are preferentially denatured by phage T4 gene 32 protein and the determination of their precise locations on the circular viral DNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Alberts, B., and Frey, L. J., Nature, 227, 1313–1317 (1970).
Delius, H., Mantell, N. J., and Alberts, B., J. molec. Biol., 67, 341–350 (1972).
Morrow, J. F., and Berg, P., Proc. natn. Acad. Sci. U.S.A., 69, 3365–3369 (1972).
Yaniv, M., Croissant, O., and Cuzin, F., Biochem. biophys. Res. Commun., 57, 1074–1079 (1974).
Fried, M., J. Virol., 13, 939–946 (1974).
Hirt, B., J. molec. Biol., 26, 365–369 (1967).
Griffin, B., Fried, M., and Cowie, A., Proc. natn. Acad. Sci. U.S.A., 71, 2077–2081 (1974).
Davis, R. W., Simon, M., and Davidson, N., Meth. Enzym., 21, 413–428 (1971).
Robberson, D., and Fried, M., Proc. natn. Acad. Sci. U.S.A., 71, 3497–3501 (1974).
Crawford, L. V., Robbins, A. K., and Nicklin, P. M., J. gen. Virol., 25, 133–142 (1974).
Follet, E. A. C., and Crawford, L. V., J. molec. Biol., 34, 565–573 (1968).
Kunito, Y., Furuno, A., and Suzuki, K., J. molec. Biol., 70, 415–423 (1972).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
MONJARDINO, J., JAMES, A. Denaturation of polyoma DNA by phage T4 gene 32 protein. Nature 255, 249–252 (1975). https://doi.org/10.1038/255249a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/255249a0
This article is cited by
-
Circular superhelical DNA
Nature (1977)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.