Abstract
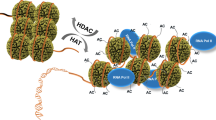
Modification of histones, DNA-binding proteins found in chromatin, by addition of acetyl groups occurs to a greater degree when the histones are associated with transcriptionally active DNA1,2. A breakthrough in understanding how this acetylation is mediated was the discovery that various transcriptional co-activator proteins have intrinsic histone acetyltransferase activity (for example, Gcn5p (ref. 3), PCAF4, TAFII250 (ref. 5) and p300/CBP6,7). These acetyltransferases also modify certain transcription factors (TFIIEβ, TFIIF, EKLF and p53 (8–10)). GATA-1 is an important transcription factor in the haematopoietic lineage11 and is essential for terminal differentiation of erythrocytes and megakaryocytes12,13. It is associated in vivo with the acetyltransferase p300/CBP14. Here we report that GATA-1 is acetylated in vitro by p300. This significantly increases the amount of GATA-1 bound to DNA and alters the mobility of GATA-1–DNA complexes, suggestive of a conformational change in GATA-1. GATA-1 is also acetylated in vivo and acetylation directly stimulates GATA-1-dependent transcription. Mutagenesis of important acetylated residues shows that there is a relationship between the acetylation and in vivo function of GATA-1. Wepropose that acetylation of transcription factors can alter interactions between these factors and DNA and among different transcription factors, and is an integral part of transcription and differentiation processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Grunstein, M. Histone acetylation in chromatin structure and transcription. Nature 389, 349–352 (1997).
Hebbes, T. R., Thorne, A. W. & Crane-Robinson, C. Adirect link between core histone acetylation and transcriptionally active chromatin. EMBO J. 7, 1395–1402 (1988).
Brownell, J. E. et al. Tetrahymena histone acetyltransferase A: a homolog to yeast Gcn5p linking histone acetylation to gene activation. Cell 84, 843–851 (1996).
Yang, X.-J., Ogryzko, V. V., Nishikawa, J., Howard, B. & Nakatani, Y. Ap300/CBP-associated factor that competes with the adenoviral oncoprotein E1a. Nature 382, 319–324 (1996).
Mizzen, C. A. et al. The TAFII250 subunit of TFIID has histone acetyltransferase activity. Cell 87, 1261–1270 (1996).
Ogryzko, V. V., Schiltz, R. L., Russanova, V., Howard, B. H. & Nakatani, Y. The transcriptional coactivators p300 and CBP are histone acetyltransferases. Cell 87, 953–959 (1996).
Bannister, A. J. & Kouzarides, T. The CBP coactivator is a histone acetyltransferase. Nature 384, 641–643 (1996).
Imhof, A. et al. Acetylation of general transcription factors by histone acetyltransferases. Curr. Biol. 7, 689–692 (1997).
Gu, W. & Roeder, R. G. Activation of p53 sequence-specific DNA binding by acetylation of the p53 C-terminal domain. Cell 90, 595–606 (1997).
Zhang, W. & Bieker, J. J. Acetylation and modulation of erythroid Kruppel-like factor (EKLF) activity by interaction with histone acetyltransferases. Proc. Natl Acad. Sci. USA 95, 9855–9860 (1998).
Orkin, S. H. GATA-binding transcription factors in hematopoietic cells. Blood 80, 575–581 (1992).
Pevny, L. et al. Erythroid differentiation in chimeric mice blocked by a targeted mutation in the gene for transcription factor GATA-1. Nature 349, 257–260 (1991).
Shivdasani, R. A., Fujiwara, Y., McDevitt, M. A. & Orkin, S. H. Alineage-selective knockout establishes the critical role of transcription factor GATA-1 in megakaryocyte growth and platelet development. EMBO J. 16, 3965–3973 (1997).
Blobel, G. A., Nakajima, T., Eckner, R., Montminy, M. & Orkin, S. H. CREB-binding protein cooperates with transcription factor GATA-1 and is required for erythroid differentiation. Proc. Natl Acad. Sci. USA 95, 2061–2066 (1998).
Trainor, C. D. et al. Apalindromic regulatory site within vertebrate GATA-1 promoters requires both zinc fingers of the GATA-1 DNA-binding domain for high-affinity interaction. Mol. Cell. Biol. 16, 2238–2247 (1996).
Evans, T. & Felsenfeld, G. The erythroid-specific transcription factor Eryf1: a new finger protein. Cell 68, 597–612 (1989).
Omichinski, J. G. et al. Asmall single-“finger” peptide from the erythroid transcription factor GATA-1 binds specifically to DNA as a zinc or iron complex. Proc. Natl Acad. Sci. USA 90, 1676–1680 (1993).
Yang, H.-Y. & Evans, T. Distinct roles for the two cGATA-1 finger domains. Mol. Cell. Biol. 12, 4562–4570 (1992).
Martin, D. I. K. & Orkin, S. H. Transcriptional activation and DNA binding by the erythroid factor GF-1/NF-E1/Eryf1. Genes Dev. 4, 1886–1898 (1990).
Merika, M. & Orkin, S. H. Functional synergy and physical interactions of the erythroid transcription factor GATA-1 with the Kruppel family proteins Sp1 and EKLF. Mol. Cell. Biol. 15, 2437–2447 (1995).
Tsang, A. P. et al. FOG, a multiple zinc finger protein acts as a cofactor for transcription factor GATA-1 in erythroid and megakaryocyte differentiation. Cell 90, 109–119 (1997).
Visvader, J. E., Crossley, M., Hill, J., Orkin, S. H. & Adams, J. M. The C-terminal zinc finger of GATA-1 or GATA-2 is sufficient to induce megakaryocyte differentiation of an early myeloid cell line. Mol. Cell. Biol. 15, 634–641 (1995).
Pikaart, M. J. & Felsenfeld, G. Expression and codon usage optimisation of the erythroid-specific transcription factor cGATA-1 in baculoviral and bacterial systems. Protein Expr. Purif. 8, 469–475 (1996).
Towatari, M. et al. Regulation of GATA-2 phosphorylation by mitogen-activated protein kinase and interleukin-3. J. Biol. Chem. 270, 4101–4107 (1995).
Briegel, K. et al. Regulation and function of transcription factor GATA-1 during red blood cell differentiation. Development 122, 3839–3850 (1996).
Evans, T. & Felsenfeld, G. Trans-activation of a globin promoter in nonerythroid cells. Mol. Cell. Biol. 11, 843–853 (1991).
Omichinski, J. G. et al. NMR structure of a specific DNA complex of Zn-containing DNA binding domain of GATA-1. Science 261, 438–446 (1993).
Acknowledgements
We thank N. Dillon and M. Greaves, in whose laboratories some of this work was done, for support and comments; C. Crane-Robinson for the anti-acetyl-lysine antisera; J. Omichinski for the GATA-1 peptides; M. Zenke for anti-cGATA-1 antisera; M. Pikaart and M. Towatari for plasmids; M.Antoniou for MEL cells; and A. Ashworth, T. Enver, G. Felsenfeld, A. Fisher, J. Mermoud and E. Milot for discussions and comments on the manuscript. J.B. acknowledges support from the Kay Kendall Leukaemia Fund.
Author information
Authors and Affiliations
Corresponding author
Supplementary Information
Rights and permissions
About this article
Cite this article
Boyes, J., Byfield, P., Nakatani, Y. et al. Regulation of activity of the transcription factor GATA-1 by acetylation. Nature 396, 594–598 (1998). https://doi.org/10.1038/25166
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/25166
This article is cited by
-
In the quest for histone deacetylase inhibitors: current trends in the application of multilayered computational methods
Amino Acids (2023)
-
Modulation of cellular processes by histone and non-histone protein acetylation
Nature Reviews Molecular Cell Biology (2022)
-
GATA-3 is a proto-oncogene in T-cell lymphoproliferative neoplasms
Blood Cancer Journal (2022)
-
Interplay between p300 and HDAC1 regulate acetylation and stability of Api5 to regulate cell proliferation
Scientific Reports (2021)
-
Inhibition of CBP synergizes with the RNA-dependent mechanisms of Azacitidine by limiting protein synthesis
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.