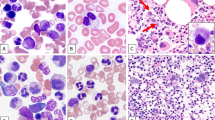
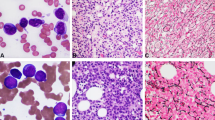
Abstract
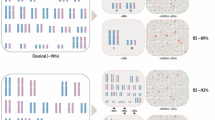
A series of 38 patients with acute myeloblastic leukemia (AML) with 49 or more chromosomes and without structural abnormalities was selected within the Groupe Francophone de Cytogénétique Hématologique (GFCH) to better define their characteristics. The median age of the patients was 65 years, and all FAB subtypes were represented. Although all chromosomes were gained, some seems to prevail: chromosome 8 (68%), 21 (47%), 19 (37%), and 13 and 14 (34% each). Since MLL rearrangement leads patients in a group with an unfavorable prognosis, search for cryptic rearrangements of MLL was performed in 34 patients and showed abnormalities in 5 (15%). When we applied the most frequent definition of complex karyotypes (three or more abnormalities), all patients with high hyperdiploid AML fall in the unfavorable category. Among the 18 patients without MLL rearrangement receiving an induction therapy, 16 (89%) reached CR and 6 (33%) were still alive after a 31-month median follow-up (14–61 months). Although this study was retrospective, these results suggest that high hyperdiploid AML without chromosome rearrangement seems to be a subgroup of uncommon AML (less than 1%), and may be better classified in the intermediate prognostic group.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Brunning RD, Matutes E, Harris NL, Flandrin G, Vardiman J, Bennett J et al. Acute myeloid leukaemia. In: Jaffe ES, Harris NL, Stein H, Vardiman JW (eds). World Health Organization of Tumors. Pathology and Genetics of Tumors of Haematopoietic and Lymphoid Tissues. IARC Press: Lyon, France, 2001 pp 75–108.
Grimwade D, Walker H, Oliver F, Wheatley K, Harrison C, Harrison G, et al., On behalf of the Medical Research Council Adult and Children's Leukemia Working Parties. The importance of diagnosis cytogenetics on outcome in AML: analysis of 1,612 patients entered into the MRC AML 10 trial. Blood 1998; 92: 2322–2333.
Slovak ML, Kopecky KJ, Cassileth PA, Harrington DH, Theil KS, Mohamed A, et al., For the Southwest Oncology Group and the Eastern Cooperative Oncology Group. Karyotypic analysis predicts outcome of preremission and postremission therapy in adult acute myeloid leukaemia: a Southwest Oncology Group/Eastern Cooperative Oncology Group study. Blood 2000; 96: 4075–4083.
Byrd JC, Mrozek K, Dodge RK, Carroll AJ, Edwards CG, Arthur DC et al. Pretreatment cytogenetic abnormalities are predictive of induction success, cumulative incidence of relapse, and overall survival in adult patients with de novo acute myeloid leukaemia: results from Cancer and Leukaemia Group B (CALGB 8461). Blood 2002; 100: 4325–4336.
Kern W, Haferlach T, Schoch C, Löffler H, Gassmann W, Heinecke A et al. Early blast clearance by remission induction therapy is a major independent prognostic factor for both achievement of complete remission and long term outcome in acute myeloid leukemia: data from the German AML Cooperative Group (AMLCG) 1992 Trial. Blood 2003; 101: 64–70.
Schoch C, Kern W, Kohlmann A, Hiddemann W, Schnittger S, Haferlach T . Acute myeloid leukemia with a complex karyotype is a distinct biological entity characterized by genomic imbalances and a specific gene expression profile. Genes Chromosomes Cancer 2005; 43: 227–238.
ISCN. An International System for Human Cytogenetic Nomenclature. In: Shaffer LG, Tommerup N (eds). S Karger: Basel, 2005.
Leymarie V, Flandrin G, Noguera ME, Leymarie F, Lioure B, Daliphard S, And the GOELAMS cytologists (Groupe ouest-est des leucémies aiguës et autres maladies du sang). Telehematology: a pilot experience of cytological diagnosis of acute myeloid leukemia via the internet. A GOELAMS study. Haematologica 2005; 91: 1285–1286.
Schoch C, Haferlach T, Bursch S, Gerstner D, Schnittger S, Dugas M et al. Loss of genetic material is more common than gain in acute myeloid leukemia with complex aberrant karyotype: a detailed analysis of 125 cases using conventional chromosome analysis and fluorescence in situ hybridization including 24-color FISH. Genes Chromosomes Cancer 2002; 35: 20–29.
Schoch C, Kohlmann A, Dugas M, Kern W, Schnittger S, Haferlach T . Impact of trisomy 8 on expression of genes located on chromosome 8 in different AML subgroups. Genes Chromosomes Cancer 2006; 45: 1164–1168.
Paulsson K, Heidenblad M, Strombeck B, Staaf J, Jonsson G, Borg A et al. High-resolution genome wide-array-based comparative genome hybridization reveals cryptic chromosome changes in AML and MDS cases with trisomy 8 as the sole cytogenetic aberration. Leukemia 2006; 20: 840–846.
Paulsson K, Panagopoulos I, Knuutila S, Jee KJ, Garwicz S, Fioretos T et al. Formation of trisomies and their parental origin in hyperdiploid childhood acute lymphoblastic leukemia. Blood 2003; 102: 3010–3015.
Virtaneva K, Wright FA, Tanner SM, Yuan B, Lemon WJ, Caliguiri MA et al. Expression profiling reveals fundamental biological differences in acute myeloid leukaemia with isolated trisomy 8 and normal cytogenetics. Proc Natl Acad Sci USA 2001; 98: 1124–1129.
Schoch C, Kohlmann A, Dugas M, Kern W, Hiddemann W, Schnittger S et al. Genomic gains and losses influence expression levels of genes located within the affected regions: a study on acute myeloid leukemias with trisomy 8, 11 or 13 and monosomy 7 or deletion 5q. Leukemia 2005; 19: 1224–1228.
Zhang FF, Murata-Collins JL, Gaytan P, Forman SJ, Kopecky KJ, Wilman CL et al. Twenty-four-color spectral karyotyping reveals chromosome aberrations in ‘cytogenetically normal’ acute myeloid leukemia. Genes Chromosomes Cancer 2000; 28: 318–328.
Vledman T, Vignon C, Schrock E, Rowley JD, Ried T . Hidden chromosome abnormalities in haematological malignancies detected by multicolour spectral karyotyping. Nat Genet 1997; 15: 406–410.
Caligiuri MA, Strout MP, Lawrence D, Arthur DC, Baer MR, Yu F et al. Rearrangement of ALL1 (MLL) in acute myeloid leukemia with normal cytogenetics. Cancer Res 1998; 58: 55–59.
von Bergh A, Emmanuel B, van Zelderen-Bhola S, Smetsers T, van Soest R, Stul M et al. A DNA probe combination for improved detection of MLL/11q23 breakpoints by double-color interphase-FISH in acute leukemias. Genes Chromosomes Cancer 2000; 28: 14–22.
Mitelman F, Johansson B, Mertens F (eds). Mitelman Database of Chromosomes Aberrations in Cancer [Internet] 2007. available from http://cgap.nci.nih.gov/Chromosomes/Mitelman,accessed on January 2007.
Schoch C, Schnittger S, Klaus M, Hiddemann W, Haferlach T . AML with 11q23/MLL abnormalities as defined by the WHO classification: incidence, partner chromosomes, FAB subtype, age distribution and prognostic impact in an unselected series of 1897 cytogenetically analyzed AML cases. Blood 2003; 102: 2395–2402.
Bloomfield CD, Archer KJ, Mrozek K, Lillington DM, Kancho Y, Head DR et al. 11q23 balanced chromosomes aberrations in treatment related myelodysplastic syndromes and acute leukaemia: report from an international workshop. Genes Chromosomes Cancer 2002; 33: 362–378.
Paulsson K, Heidenblad M, Mörse H, Borg A, Fioretos T, Johansson B . Identification of cryptic aberrations and characterization of translocation breakpoints using array CGH in high hyperdiploid childhood acute lymphoblastic leukemia. Leukemia 2006; 20: 2002–2007.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luquet, I., Laï, J., Barin, C. et al. Hyperdiploid karyotypes in acute myeloid leukemia define a novel entity: a study of 38 patients from the Groupe Francophone de Cytogenetique Hematologique (GFCH). Leukemia 22, 132–137 (2008). https://doi.org/10.1038/sj.leu.2404974
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2404974
Keywords
This article is cited by
-
Allogeneic hematopoietic cell transplantation for acute myeloid leukemia with hyperdiploid complex karyotype
Bone Marrow Transplantation (2024)
-
The complex karyotype in hematological malignancies: a comprehensive overview by the Francophone Group of Hematological Cytogenetics (GFCH)
Leukemia (2022)
-
A new adult AML case with an extremely complex karyotype, remission and relapse combined with high hyperdiploidy of a normal chromosome set in secondary AML
BMC Hematology (2018)
-
Karyotype complexity and prognosis in acute myeloid leukemia
Blood Cancer Journal (2016)
-
Hyperdiploidy with 49–65 chromosomes represents a heterogeneous cytogenetic subgroup of acute myeloid leukemia with differential outcome
Leukemia (2014)