Abstract
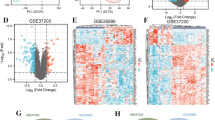
To investigate the relationship between Barrett's esophagus (BE) and esophageal adenocarcinoma (EAC), we determined gene expression profiles of discrete pathological stages of esophageal neoplasia using a sequence-verified human cDNA microarray. Fifty one RNAs, comprising 24 normal esophagi (NE), 18 BEs, and nine EACs were hybridized to cDNA microarrays. Five statistical analyses were used for the data analysis. Genes showing significantly different expression levels among the three sample groups were identified. Genes were grouped into functional categories based on the Gene Ontology Consortium. Surprisingly, the expression pattern of BE was significantly more similar to EAC than to NE, notwithstanding the known histopathologic differences between BE and EAC. The pattern of NE was clearly distinct from that of EAC. Thirty-six genes were the most differentially modulated, according to these microarray data, in BE-associated neoplastic progression. Twelve genes were significantly differentially expressed in cancer-associated BE's plus EAC (as a single combined tissue group) vs noncancer-associated BE's. These genes represent potential biomarkers to diagnose EAC at its early stages. Our results demonstrate that molecular events at the transcriptional level in BE are remarkably similar to BE's-associated adenocarcinoma of the esophagus. This finding alarmingly implies that BE is biologically closer to cancer than to normal esophagus, and that the cancer risk of BE is perhaps higher than we had imagined. These findings suggest that changes modulated at the molecular biologic level supervene earlier than histologic changes, and that BE is an early intermediate stage in the process of EAC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Abraham JM, Wang S, Suzuki H, Jiang HY, Rosenblum-Vos LS, Yin J et al. (1996). Cell Growth Differ 7: 855–860.
Blot WJ, McLaughlin JK . (1999). Semin Oncol 26 ( Suppl 15): 2–8.
Boynton RF, Huang Y, Blount PL, Reid BJ, Raskind WH, Haggitt RC et al. (1991). Cancer Res 51: 5766–5769.
Brabender J, Marjoram P, Salonga D, Metzger R, Schneider PM, Park JM et al. (2004). Oncogene 23: 4780–4788.
Burdiles P, Csendes A, Smok G, Braghetto I, Korn O . (2003). Rev Med Chil 131: 587–596.
Buttar NS, Wang KK, Leontovich O, Westcott JY, Pacifico RJ, Anderson MA et al. (2002). Gastroenterology 122: 1101–1112.
Cameron AJ . (2002). Dis Esophagus 15: 106–108.
Cesar AC, Borim AA, Caetano A, Cury PM, Silva AE . (2004). Cancer Genet Cytogenet 153: 127–132.
Chang Y, Gong J, Liu B, Zhang J, Dai F . (2004). World J Gastroenterol 10: 3194–3196.
Chen X, Yang CS . (2001). Carcinogenesis 22: 1119–1129.
Chen X, Ding YW, Yang G, Bondoc F, Lee MJ, Yan CS . (2000). Carcinogenesis 21: 257–263.
Chen X, Li N, Wang S, Hong J, Fang M, Yousselfson J et al. (2002). Carcinogenesis 23: 2095–2102.
Chung GG, Zerkowski MP, Ocal IT, Dolled-Filhart M, Kang JY, Psyrri A et al. (2004). Cancer 100: 2084–2092.
Cossentino MJ, Wong RK . (2003). Semin Gastrointest Dis 14: 128–135.
Ferguson MK, Durkin A . (2002). J Gastrointest Surg 6: 29–35; discussion 36.
Fujii T, Nakagawa S, Hanzawa M, Sueyoshi S, Fujita H, Shirouzu K et al. (2003). Oncol Rep 10: 427–431.
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S et al. (2004). Genome Biol 5: R80.
Going JJ, Keith WN, Neilson L, Stoeber K, Stuart RC, Williams GH . (2002). Gut 50: 373–377.
Harris MA, Clark J, Ireland A, Lomax J, Ashburner M, Foulger R et al. (2004). Nucleic Acids Res 32 (Database issue): D258–D261.
Hosack DA, Dennis Jr G, Sherman BT, Lane HC, Lempicki RA . (2003). Genome Biol 4: R70.
Hovanes K, Li TW, Munguia JE, Truong T, Milovanovic T, Lawrence MJ et al. (2001). Nat Genet 28: 53–57.
Hyun DH, Gray DA, Halliwell B, Jenner P . (2004). J Neurochem 90: 422–430.
Kimos MC, Wang S, Borkowski A, Yang GY, Yang CS, Perry K et al. (2004). Int J Cancer 111: 415–417.
Li A, Varney ML, Singh RK . (2004). Clin Exp Metastasis 21: 571–579.
Li H, Walsh TN, O'Dowd G, Gillen P, Byrne PJ, Hennessy TP . (1994). Surgery 115: 176–181.
Matsushima T, Mori M, Kido A, Adachi Y, Sugimachi K . (1998). Oncol Rep 5: 73–76.
McManus DT, Olaru A, Meltzer SJ . (2004). Cancer Res 64: 1561–1569.
Meighan-Mantha RL, Hsu DK, Guo Y, Brown SA, Feng SL, Peifley KA et al. (1999). J Biol Chem 274: 33166–33176.
Meltzer SJ, Yin J, Manin B, Rhyu MG, Cottrell J, Hudson E et al. (1994). Cancer Res 54: 3379–3382.
Mori M, Mimori K, Yoshikawa Y, Shibuta K, Utsunomiya T, Sadanaga N et al. (2002). Surgery 131 ( Suppl): S39–S47.
Mori Y, Selaru FM, Sato F, Yin J, Simms LA, Xu Y et al. (2003). Cancer Res 63: 4577–4582.
Offner FA, Lewin KJ, Weinstein WM . (1996). Hum Pathol 27: 885–889.
Powell J, McConkey CC . (1992). Eur J Cancer Prev 1: 265–269.
Raskind WH, Norwood T, Levine DS, Haggitt RC, Rabinovitch PS, Reid BJ . (1992). Cancer Res 52: 2946–2950.
Reid BJ, Haggitt RC, Rubin CE, Rabinovitch PS . (1987). Gastroenterology 93: 1–11.
Sanz-Ortega J, Hernandez S, Saez MC, Sierra E, Sanz-Ortega G, Torres A et al. (2003). Hepatogastroenterology 50: 404–407.
Schwartz DR, Wu R, Kardia SL, Levin AM, Huang CC, Shedden KA et al. (2003). Cancer Res 63: 2913–2922.
Selaru FM, Zou T, Xu Y, Shustova V, Yin J, Mori Y et al. (2002). Oncogene 21: 475–478.
Shirvani VN, Ouatu-Lascar R, Kau BS, Omary MB, Triadafilopoulos G . (2000). Gastroenterology 118: 487–496.
Spechler SJ . (2002). Med Clin N Am 86: 1423–1445, vii.
Stoltzing O, Schneider PM, Becker K, Wegerer S, Siewert JR, Holscher AH . (1998). Langenbecks Arch Chir Suppl Kongressbd 115 (Suppl I): 485–489.
Suspiro A, Pereira AD, Afonso A, Albuquerque C, Chaves P, Soares J et al. (2003). Am J Gastroenterol 98: 728–734.
Tamoto E, Tada M, Murakawa K, Takada M, Shindo G, Teramoto K et al. (2004). Clin Cancer Res 10: 3629–3638.
Tibshirani R, Hastie T, Narasimhan B, Chu G . (2002). Proc Natl Acad Sci USA 99: 6567–6572.
Tusher VG, Tibshirani R, Chu G . (2001). Proc Natl Acad Sci USA 98: 5116–5121.
Umansky M, Yasui W, Hallak A, Brill S, Shapira I, Halpern Z et al. (2001). Oncogene 20: 7987–7991.
von Rahden BH, Stein HJ, Siewert JR . (2003). Curr Oncol Rep 5: 203–209.
Wang S, Mori Y, Sato F, Yin J, Xu Y, Zou TT et al. (2003). Oncogene 22: 467–470.
Wiley SR, Winkles JA . (2003). Cytokine Growth Factor Rev 14: 241–249.
Wilson KT, Fu S, Ramanujam KS, Meltzer SJ . (1998). Cancer Res 58: 2929–2934.
Xu Y, Selaru FM, Yin J, Zou TT, Shustova V, Mori Y et al. (2002). Cancer Res 62: 3493–3497.
Yamamoto H, Horiuchi S, Adachi Y, Taniguchi H, Nosho K, Min Y et al. (2004). Carcinogenesis 25: 325–332.
Zhai Y, Wu R, Schwartz DR, Darrah D, Reed H, Kolligs FT et al. (2002). Am J Pathol 160: 1229–1238.
Acknowledgements
This work was supported by the grants CA85069, CA01808, CA95323, DK67872, CA10676.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, S., Zhan, M., Yin, J. et al. Transcriptional profiling suggests that Barrett's metaplasia is an early intermediate stage in esophageal adenocarcinogenesis. Oncogene 25, 3346–3356 (2006). https://doi.org/10.1038/sj.onc.1209357
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209357
Keywords
This article is cited by
-
Comprehensive RNA dataset of tissue and plasma from patients with esophageal cancer or precursor lesions
Scientific Data (2022)
-
Alteration of protein expression and spliceosome pathway activity during Barrett’s carcinogenesis
Journal of Gastroenterology (2021)
-
Low blood levels of sTWEAK are related to locoregional failure in head and neck cancer
European Archives of Oto-Rhino-Laryngology (2015)
-
Next-generation sequencing of endoscopic biopsies identifies ARID1A as a tumor-suppressor gene in Barrett’s esophagus
Oncogene (2014)
-
The actin-myosin regulatory MRCK kinases: regulation, biological functions and associations with human cancer
Journal of Molecular Medicine (2014)