Abstract
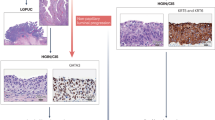
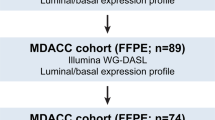
We used gene expression profiling, mutation analyses of FGFR3 and TP53, and LOH analyses of chromosome 9 and the TP53 region on chromosome arm 17p, to molecularly characterize 75 Ta and T1 bladder carcinomas. We identified four major cellular processes related to cell cycle, protein synthesis, immune response, and extra cellular components that contribute to the expressional heterogeneity of early-stage urothelial cell carcinoma (UCC). Activating FGFR3 mutations were found at the highest frequency in G1 tumors (80%), and showed a strong correlation with FGFR3 expression. In contrast, G3 tumors displayed mutations in less than 10% of the cases and a low level of FGFR3 expression. Even though LOH on chromosome 9 was not associated with any specific expression pattern, our data indicate that loss of chromosome 9 is associated with tumor development rather than initiation. The combined analyses suggest the existence of two types of UCC tumors, one which is characterized by FGFR3 mutation or expression, high expression of protein synthesis genes, and low expression of cell cycle genes. Furthermore, the presented data underscore FGFR3 receptor involvement in urothelial cell transformation as the presence of FGFR3 mutations has a major impact on the global gene expression profile of bladder carcinomas.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bakkar AA, Wallerand H, Radvanyi F, Lahaye JB, Pissard S, Lecerf L et al. (2003). Cancer Res 63: 8108–8112.
Baud E, Catilina P, Bignon YJ . (1999). Int J Oncol 14: 441–445.
Belkin AM, Stepp MA . (2000). Microsc Res Tech 51: 280–301.
Billerey C, Chopin D, Aubriot-Lorton MH, Ricol D, Gil Diez de Medina S, Van Rhijn B et al. (2001). Am J Pathol 158: 1955–1959.
Blaveri E, Simko JP, Korkola JE, Brewer JL, Baehner F, Mehta K et al. (2005). Clin Cancer Res 11: 4044–4055.
Brausi M, Collette L, Kurth K, van der Meijden AP, Oosterlinck W, Witjes JA et al. (2002). Eur Urol 41: 523–531.
Cappellen D, De Oliveira C, Ricol D, de Medina S, Bourdin J, Sastre-Garau X et al. (1999). Nat Genet 23: 18–20.
Cordon-Cardo C . (1998). Cancer Surv 32: 115–131.
Duggan B, Williamson K . (2004). Curr Opin Urol 14: 277–286.
Dyrskjøt L, Kruhøffer M, Thykjaer T, Marcussen N, Jensen JL, Møller K et al. (2004). Cancer Res 64: 4040–4048.
Dyrskjøt L, Thykjaer T, Kruhøffer M, Jensen JL, Marcussen N, Hamilton-Dutoit S et al. (2003). Nat Genet 33: 90–96.
Dyrskjøt L, Zieger K, Kruhøffer M, Thykjaer T, Jensen JL, Primdahl H et al. (2005). Clin Cancer Res 11: 4029–4036.
Hafner C, Knuechel R, Stoehr R, Hartmann A . (2002). Int J Cancer 101: 1–6.
Haukaas S, Daehlin L, Maartmann-Moe H, Ulvik NM . (1999). BJU Int 83: 957–963.
Herr HW . (2000). J Urol 163: 60–61; discussion 61–62.
Heyer LJ, Kruglyak S, Yooseph S . (1999). Genome Res 9: 1106–1115.
Höglund M, Säll T, Heim S, Mitelman F, Mandahl N, Fadl-Elmula I . (2001). Cancer Res 61: 8241–8246.
Kim JH, Tuziak T, Hu L, Wang Z, Bondaruk J, Kim M et al. (2005). Lab Invest 85: 532–549.
Knowles MA . (1999). Electrophoresis 20: 269–279.
Modlich O, Prisack HB, Pitschke G, Ramp U, Ackermann R, Bojar H et al. (2004). Clin Cancer Res 10: 3410–3421.
Mor O, Nativ O, Stein A, Novak L, Lehavi D, Shiboleth Y et al. (2003). Oncogene 22: 7702–7710.
Mostofi FK, Davies CJ, Sesterhenn I . (1999). Histological Typing Of Urinary Bladder Tumors. International Classification Of Tumors, 2nd edn. Springer: Berlin.
Prout GRJ, Barton BA, Griffin PP, Friedell GH . (1992). J Urol 148: 1413–1419.
Richter J, Beffa L, Wagner U, Schraml P, Gasser TC, Moch H et al. (1998). Am J Pathol 153: 1615–1621.
Saal LH, Troein C, Vallon-Christersson J, Gruvberger S, Borg A, Peterson C . (2002). Genome Biol 3: SOFTWARE0003.
Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N et al. (2003). Biotechniques 34: 374–378.
Sanchez-Carbayo M, Socci ND, Lozano JJ, Li W, Charytonowicz E, Belbin TJ et al. (2003). Am J Pathol 163: 505–516.
Sobin LH, Wittekind CL . (2002). TNM Classification Of Malignant Tumors, 6th edn. John Wiley & Sons: New York.
Spruck III CH, Ohneseit PF, Gonzalez-Zulueta M, Esrig D, Miyao N, Tsai YC et al. (1994). Cancer Res 54: 784–788.
Thykjaer T, Workman C, Kruhøffer M, Demtroder K, Wolf H, Andersen LD et al. (2001). Cancer Res 61: 2492–2499.
Trudel S, Li ZH, Wei E, Wiesmann M, Chang H, Chen C et al. (2005). Blood 105: 2941–2948.
Tusher VG, Tibshirani R, Chu G . (2001). Proc Natl Acad Sci USA 98: 5116–5121.
Vajo Z, Francomano CA, Wilkin DJ . (2000). Endocr Rev 21: 23–39.
van Rhijn BW, van der Kwast TH, Vis AN, Kirkels WJ, Boeve ER, Jobsis AC et al. (2004). Cancer Res 64: 1911–1914.
van Rhijn BW, van Tilborg AA, Lurkin I, Bonaventure J, de Vries A, Thiery JP et al. (2002). Eur J Hum Genet 10: 819–824.
van Rhijn BW, Vis AN, van der Kwast TH, Kirkels WJ, Radvanyi F, Ooms EC et al. (2003). J Clin Oncol 21: 1912–1921.
Wary KK, Mariotti A, Zurzolo C, Giancotti FG . (1998). Cell 94: 625–634.
Wild PJ, Herr A, Wissmann C, Stoehr R, Rosenthal A, Zaak D et al. (2005). Clin Cancer Res 11: 4415–4429.
Yang YH, Dudoit S, Luu P, Lin DM, Peng V, Ngai J et al. (2002). Nucleic Acids Res 30: e15.
Acknowledgements
This study was supported by the Swedish Cancer Society, The Gunnar, Arvid and Elisabeth Nilsson Foundation, The Crafoord Foundation, the John and Augusta Persson Foundation, the IngaBritt and Arne Lundberg Foundation, the Maud and Birger Gustavsson Foundation, and the Petrus and Augusta Hedlund Foundation. The microarray facility was supported by the Knut and Alice Wallenberg Foundation via the Swegene Program.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Rights and permissions
About this article
Cite this article
Lindgren, D., Liedberg, F., Andersson, A. et al. Molecular characterization of early-stage bladder carcinomas by expression profiles, FGFR3 mutation status, and loss of 9q. Oncogene 25, 2685–2696 (2006). https://doi.org/10.1038/sj.onc.1209249
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209249