Abstract
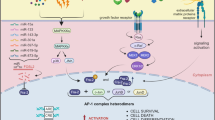
Numerous studies have revealed distinct functions of Fos proteins in different mouse tissues and cell lines. Here, we perform a direct comparison of the features of exogenous c-Fos, Fra-1 and Fra-2 proteins expressed in murine tumor cells of epithelial origin, CSML0. Although transactivation potential of c-Fos is much stronger than that of Fra-1 and Fra-2, all three proteins are capable of modulating transcription of target genes. Moreover, there is a certain degree of specificity in the induction of the transcription of AP-1-responsive genes by different Fos proteins. For instance, c-Fos and Fra-1 but not Fra-2 activated genes of the urokinase system. Additionally, not only a strong transcriptional activator c-Fos, but also Fra-1 induced morphological alterations in CSML0 cells. N-terminal domain of Fra-1 was required for this function. On the other hand, Fra-2 failed to change morphology of CSML0 cells. We therefore conclude that c-Fos, Fra-1 and Fra-2 differently activate transcription of target genes and induce morphological changes in epithelioid carcinoma cells in a manner not directly linked to their transactivation potentials.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Angel P, Karin M . 1991 Biochim. Biophys. Acta 1072: 129–157
Battista S, de Nigris F, Fedele M, Chiappetta G, Scala S, Vallone D, Pierantoni G, Megar T, Santoro M, Viglietto G, Verde P, Fusco A . 1998 Oncogene 17: 377–385
Bergers G, Graninger P, Braselmann S, Wrighton C, Busslinger M . 1995 Mol. Cell. Biol. 15: 3748–3758
Brown J, Ye H, Bronson R, Dikkes P, Greenberg M . 1996 Cell 86: 297–309
Chiappetta G, Tallini G, De Biasio MC, Pentimalli F, de Nigris F, Losito S, Fedele M, Battista S, Verde P, Santoro M, Fusco A . 2000 Clin. Cancer Res. 6: 4300–4307
Chomczynski P, Sacchi N . 1987 Analytical Biochem. 162: 156–159
Debinski W, Slagle-Webb B, Achen M, Stacker S, Tulchinsky E, Gillespie G, Gibo D . 2001 Mol. Med. 7: 598–608
Grigoriadis E, Wang Z-Q, Cecchini M, Hoffstette W, Felix R, Fleisch H, Wagner E . 1994 Science 266: 443–447
Jehn B, Costello E, Marti A, Keon N, Deane R, Li F, Friis R, Burry P, Martin F, Jaggi R . 1992 Mol. Cell. Biol. 12: 3890–3902
Jehn J, Wisdom R, Tratner I, Verma IM . 1991 Proc. Natl. Acad. Sci. USA 88: 5077–5081
Karin M, Liu Z-g, Zandi E . 1997 Curr. Opin. Cell. Biol. 9: 240–246
Kirschmeier P, Housey G, Johnson M, Perkins A, Weinstein I . 1988 DNA 7: 219–225
Kovary K, Bravo R . 1992 Mol. Cell. Biol. 12: 5015–5023
Kustikova O, Kramerov D, Grigorian M, Berezin V, Bock E, Lukanidin E, Tulchinsky E . 1998 Mol. Cell. Biol. 18: 7095–7105
Lepekhin EA, Walmod PS, Berezin A, Berezin V, Bock E . 2000 Methods Mol. Biol. 161: 85–100
Markowitz D, Goff S, Bank A . 1989 J. Virol. 62: 1120–1124
Matsuo K, Owens J, Tonko M, Elliot C, Chambers T, Wagner E . 2000 Nature Gen. 24: 184–187
Mechta F, Lallemand D, Pfarr C, Yaniv M . 1997 Oncogene 14: 837–847
Murakami M, Sonobe M, Ui M, Kabuyama Y, Watanabe H, Wada T, Handa H, Iba H . 1997 Oncogene 14: 2435–2444
Murakami M, Ui M, Iba H . 1999 Cell Growth Differ. 10: 333–342
Ozanne B, McGarry L, Spence H, Jonston I, Winnie J, Meagher L, Stapleton G . 2000 Eur. J. Cancer 36: 1640–1648
Qian X, Wang T, Rothman V, Nicosia R, Tuszynski G . 1997 Exp. Cell. Res. 235: 403–412
Risse-Hackl G, Adamkiewicz J, Wimmel A, Schuermann M . 1998 Oncogene 16: 3057–3068
Saksela K, Baltimore D . 1993 Mol. Cell. Biol. 13: 3698–3705
Schreiber M, Wang Z, Jochum W, Fetka I, Elliot C, Wagner E . 2000 Development 127: 4937–4948
Suzuki T, Okuno H, Yoshido T, Endo T, Nishina H, Iba H . 1991 Nucl. Acids Res. 19: 5537–5542
Tulchinsky E . 2000 Histol. Histopathol. 15: 921–928
Vallone D, Battista S, Pierantoni G, Fedele M, Casalino L, Santoro M, Viglietto G, Fusco A, Verde P . 1997 EMBO J. 16: 5310–5321
Wisdom R . 1999 Exp. Cell. Res. 253: 180–185
Wisdom R, Verma I . 1993 Mol. Cell. Biol. 13: 7429–7438
Zajchowski D, Bartholdi M, Gong Y, Webster L, Lui H, Munishkin A, Beauheim C, Harvey S, Ethier S, Johnson P . 2001 Cancer Res. 61: 5168–5178
Acknowledgements
This work was supported by grants from Danish Cancer Society, Danish Medical Research Council and the Novo Nordisk foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Andersen, H., Mahmood, S., Tkach, V. et al. The ability of Fos family members to produce phenotypic changes in epithelioid cells is not directly linked to their transactivation potentials. Oncogene 21, 4843–4848 (2002). https://doi.org/10.1038/sj.onc.1205590
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1205590
Keywords
This article is cited by
-
Role of Fra-2 in cancer
Cell Death & Differentiation (2024)
-
Fra-1 controls motility of bladder cancer cells via transcriptional upregulation of the receptor tyrosine kinase AXL
Oncogene (2012)
-
Nuclear deadenylation/polyadenylation factors regulate 3′ processing in response to DNA damage
The EMBO Journal (2010)
-
Combination of osteopontin and activated leukocyte cell adhesion molecule as potent prognostic discriminators in HER2- and ER-negative breast cancer
British Journal of Cancer (2010)
-
Role of Fra-2 in breast cancer: influence on tumor cell invasion and motility
Breast Cancer Research and Treatment (2008)