Abstract
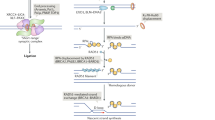
The maintenance of genomic stability is one of the most important defenses against neoplastic transformation. This objective must be accomplished despite a constant barrage of spontaneous DNA double strand breaks. These dangerous lesions are corrected by two primary pathways of double strand break repair; non homologous end joining and homologous recombination. Recent studies employing mouse models have shown that absence of either pathway leads to genomic instability, including potentially oncogenic translocations. Because translocations involve the union of different chromosomes, cellular machinery must exist that creates these structures in the context of unrepaired double strand breaks. Evidence is mounting that the pathways of double strand break repair that are so important for survival may themselves be the culprits that generate potentially fatal translocations. Evidence and models for the dual roles of double strand break repair in both preventing, and generating, oncogenic karyotypic changes are discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Artandi SE, Chang S, Lee SL, Alson S, Gottlieb GJ, Chin L, DePinho RA . 2000 Nature 406: 641–645
Bailey SM, Meyne J, Chen DJ, Kurimasa A, Li GC, Lehnert BE, Goodwin EH . 1999 Proc. Natl. Acad. Sci. USA 96: 14899–14904
Baylin SB, Herman JG . 2000 Trends Genet. 16: 168–174
Blasco MA, Lee HW, Hande MP, Samper E, Lansdorp PM, DePinho RA, Greider CW . 1997 Cell 91: 25–34
Bosco G, Haber JE . 1998 Genetics 150: 1037–1047
Coleman AE, Kovalchuk AL, Janz S, Palini A, Ried T . 1999 Blood 93: 4442–4444
Connor F, Bertwistle D, Mee PJ, Ross GM, Swift S, Grigorieva E, Tybulewicz VL, Ashworth A . 1997 Nat. Genet. 17: 423–430
Davies AA, Masson JY, McIlwraith MJ, Stasiak AZ, Stasiak A, Venkitaraman AR, West SC . 2001 Mol. Cell. 7: 273–282
Difilippantonio MJ, Zhu J, Chen HT, Meffre E, Nussenzweig MC, Max EE, Ried T, Nussenzweig A . 2000 Nature 404: 510–514
Ferguson DO, Holloman WK . 1996 Proc. Natl. Acad. Sci. USA 93: 5419–5424
Ferguson DO, Sekiguchi JM, Chang S, Frank KM, Gao Y, DePinho RA, Alt FW . 2000 Proc. Natl. Acad. Sci. USA 97: 6630–6633
Frank KM, Sharpless NE, Gao Y, Sekiguchi JM, Ferguson DO, Zhu C, Manis JP, Horner J, DePinho RA, Alt FW . 2000 Mol. Cell. 5: 993–1002
Fugmann SD, Lee AI, Shockett PE, Villey IJ, Schatz DG . 2000 Annu. Rev. Immunol. 18: 495–527
Gao Y, Chaudhuri J, Zhu C, Davidson L, Weaver DT, Alt FW . 1998 Immunity 9: 367–376
Gao Y, Ferguson DO, Xie W, Manis J, Sekiguchi J, Frank KM, Chaudhuri J, Horner J, DePinho RA, Alt FW . 2000 Nature 404: 897–900
Guidos CJ, Williams CJ, Grandal I, Knowles G, Huang MT, Danska JS . 1996 Genes Dev. 10: 2038–2054
Haber JE . 1999 Trends Biochem. Sci. 24: 271–275
Haber JE . 2000 Curr. Opin. Cell. Biol. 12: 286–292
Hilgenfeld E, Padilla-Nash H, Schrock E, Ried T . 1999 Curr. Top. Microbiol. Immunol. 246: 169–174
Holliday R . 1967 Mutat. Res. 4: 275–288
Holmes AM, Haber JE . 1999 Cell 96: 415–424
Howard-Flanders P, Boyce RP . 1966 Radiat. Res supp 16, 156
Jeggo PA . 1998 Radiat. Res. 150: S80–S91
Johnson RD, Jasin M . 2000 EMBO J. 19: 3398–3407
Karanjawala ZE, Grawunder U, Hsieh CL, Lieber MR . 1999 Curr. Biol. 9: 1501–1504
Komori T, Okada A, Stewart V, Alt FW . 1993 Science 261: 1171–1175
Korsmeyer SJ . 1992 Annu. Rev. Immunol. 10: 785–807
Lieber MR . 1999 Genes Cells 4: 77–85
Liyanage M, Coleman A, du Manoir S, Veldman T, McCormack S, Dickson RB, Barlow C, Wynshaw-Boris A, Janz S, Wienberg J, Ferguson-Smith MA, Schrock E, Ried T . 1996 Nat. Genet. 14: 312–315
Loeb KR, Loeb LA . 2000 Carcinogenesis 21: 379–385
Malkova A, Ivanov EL, Haber JE . 1996 Proc. Natl. Acad. Sci. USA 93: 7131–7136
McClintock B . 1941 Genetics 26: 234–282
McEachern MJ, Krauskopf A, Blackburn EH . 2000 Annu. Rev. Genet. 34: 331–358
Moshous D, Callebaut I, de Chasseval R, Corneo B, Cavazzana-Calvo M, Le Deist F, Tezcan I, Sanal O, Bertrand Y, Philippe N, Fischer A, de Villartay, JP . 2001 Cell 105: 177–186
Moynahan ME, Pierce AJ, Jasin M . 2001 Mol. Cell. 7: 263–272
Nacht M, Strasser A, Chan YR, Harris AW, Schlissel M, Bronson RT, Jacks T . 1996 Genes Dev. 10: 2055–2066
Patel KJ, Vu VP, Lee H, Corcoran A, Thistlethwaite FC, Evans MJ, Colledge WH, Friedman LS, Ponder BA, Venkitaraman AR . 1998 Mol. Cell. 1: 347–357
Petrini JH . 2000 Curr. Opin. Cell. Biol. 12: 293–296
Pfeiffer P, Goedecke W, Obe G . 2000 Mutagenesis 15: 289–302
Richardson C, Jasin M . 2000a Mol. Cell. Biol. 20: 9068–9075
Richardson C, Jasin M . 2000b Nature 405: 697–700
Rudolph KL, Chang S, Lee HW, Blasco M, Gottlieb GJ, Greider C, DePinho RA . 1999 Cell 96: 701–712
Scully R, Livingston DM . 2000 Nature 408: 429–432
Sekiguchi JM, Gao Y, Gu Y, Frank K, Sun Y, Chaudhuri J, Zhu C, Cheng H-L, Manis J, Ferguson D, Davidson L, Alt FW . 1999 Cold Spring Harbor Symp. Quant. Biol. 64: 169–181
Shinohara A, Ogawa T . 1999 Mutat. Res. 435: 13–21
Sonoda E, Sasaki MS, Buerstedde JM, Bezzubova O, Shinohara A, Ogawa H, Takata M, Yamaguchi-Iwai Y, Takeda S . 1998 EMBO J. 17: 598–608
Taccioli GE, Rathbun G, Oltz E, Stamato T, Jeggo PA, Alt FW . 1993 Science 260: 207–210
Takata M, Sasaki MS, Sonoda E, Morrison C, Hashimoto M, Utsumi H, Yamaguchi-Iwai Y, Shinohara A, Takeda S . 1998 EMBO J. 17: 5497–5508
Vanasse GJ, Halbrook J, Thomas S, Burgess A, Hoekstra MF, Disteche CM, Willerford DM . 1999 J. Clin. Invest. 103: 1669–1675
Welcsh PL, King MC . 2001 Hum. Mol. Genet. 10: 705–713
Wood RD, Mitchell M, Sgouros J, Lindahl T . 2001 Science 291: 1284–1289
Xu X, Wagner KU, Larson D, Weaver Z, Li C, Ried T, Hennighausen L, Wynshaw-Boris A, Deng CX . 1999 Nat. Genet. 22: 37–43
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Ferguson, D., Alt, F. DNA double strand break repair and chromosomal translocation: Lessons from animal models. Oncogene 20, 5572–5579 (2001). https://doi.org/10.1038/sj.onc.1204767
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1204767
Keywords
This article is cited by
-
Increased double-strand breaks in aged mouse male germ cells may result from changed expression of the genes essential for homologous recombination or nonhomologous end joining repair
Histochemistry and Cell Biology (2023)
-
Early neuronal accumulation of DNA double strand breaks in Alzheimer’s disease
Acta Neuropathologica Communications (2019)
-
Protein arginine methylation: an emerging regulator of the cell cycle
Cell Division (2018)
-
Significant Role of Segmental Duplications and SIDD Sites in Chromosomal Translocations of Hematological Malignancies: A Multi-parametric Bioinformatic Analysis
Interdisciplinary Sciences: Computational Life Sciences (2018)
-
Control of DNA integrity in skeletal muscle under physiological and pathological conditions
Cellular and Molecular Life Sciences (2017)