Abstract
In fragile X syndrome, the most common cause of inherited mental retardation, phenotypic expression has been linked to a region containing a repetitive sequence, (CGG)n, that appears to lengthen dramatically in fagile X patients and to show length variation in normal individuals. In order to investigate possible mechanisms responsible for further expansion of CGG in the normal population, we selected 31 normal unrelated X chromosomes carrying either the high-risk DX204-AC15 5 or DX196-AC 151 haplotypes, as defined by the flanking microsatellites, DXS548 and FRAXAC2. Nearly 100% of CGGs with more than 35 repeats were found on DX204-AC155 haplotypes, with a mean length significantly higher and much more variable than in normal individuals carrying other haplotypes including the high-risk haplotype DX196-AC151. These findings suggest that the transition from the normal to the abnormal range occurs by a multistep process, a primary event, such as unequal crossing-over, leading to increased size and moderate instability of the repeat, and from which DNA polymerase slippage could lead to recurrent premutations. Our results also suggest that the upper limit of the normal range is roughly 35 repeats in the fragile X gene. The 36–54 repeats range would define an intermediate allele only observed, up to now, in DX204-AC155 fragile X chromosomes.
Similar content being viewed by others
Introduction
Fragile X syndrome is the most common cause of inherited mental retardation, with an incidence of about 1 in 1,500 males and 1 in 2,500 females. It is associated with a rare, fragile site at Xq27.3 (FRAXA). In males, the syndrome is associated with moderate to severe mental retardation, typical facies, post-pubertal macroorchidism and a folate-sensitive fragile site. Phenotypic expression has been linked to abnormal cytosine methylation of a single CpG island [1–3]. This region contains a repetitive sequence (CGG)n that appears to lengthen dramatically in fragile X patients and has been identified as the first exon of the FMR1 gene [4]. Analysis of length variation in the (CGG) repeat in normal individuals has shown a range of allele sizes extending from 6 to 54 repeats [5, 6]. These normal alleles are stable when segregating in families. Premutations showing no phenotypic effect in fragile X families range in size from approximately 54 to over 200 repeats, and are unstable when segregating in families. Alleles with more than 200 repeats correspond to the full mutation with meiotic and mitotic instability [5]. Expansion of premutations to full mutations only occurs in female meiotic transmission, accounting for the observation that daughters of normal transmitting males (NTMs) never show the fragile X phenotype [2].
Three CA repeats have been described flanking the CGG locus: DXS548 located 150 kb from the CGG repeat on the centromeric side [7], and FRAXAC1 and FRAX-AC2, located 10 kb on the centromeric and telomeric sides, respectively [8]. Analysis of these CA repeats in normal and fragile X chromosomes showed significant differences in allele and haplotype distributions, indicating that a limited number of primary events may have been at the origin of most present-day fragile X chromosomes in Caucasian populations [9–12]. In a recent report [13], the FRAXAC2 locus was shown to be a hypermutable DNA sequence composed of three variable subregions. Inheritance studies showed that this microsatellite was stable in normal families but unstable in fragile-X-derived meioses. However, the mutation rate of this locus does not seem high enough to blur completely the original allele combinations, as was shown by analysis of a BanI RFLP [10] and by the strong linkage disequilibria found on both normal and fragile X chromosomes [9, 10, 12, and this work].
Among the founder chromosomes at the origin of fragile X chromosomes, two are strongly associated with the fragile X syndrome: DX204-AC155, a haplotype composed of the 204-bp allele at the DXS548 locus and of the 155-bp allele at the FRAXAC2 locus, and DX196-AC151, a haplotype composed of the 196-bp allele at DXS548 and of the 151-bp allele at FRAXAC2. To explain the existence of such chromosomes associated with fragile X syndrome, it has been suggested that the stability of normal CGGs may depend on their length or on possible differential interspersion of AGG triplets, as uninterrupted exact repeats are expected to be more unstable [10]. High-risk haplotypes would carry longer CGG and/or more perfect repeats. In order to investigate such hypotheses, we compared the size of the CGG repeat in normal chromosomes carrying high-risk haplotypes with that in other chromsomes. Our findings show a clear-cut difference in the distribution of the CGG repeat in DX204-AC155 chromosomes, for which the larger size of the CGG accounts for the observed intermediate 36–45 repeats range. This result suggests that the transition from the normal (<36) to the abnormal (>54) range occurred by a multistep process, for instance unequal crossing-over followed by DNA polymerase slippage.
Methods
Subjects
Typing of the DXS548 CA repeat was performed on a sample of 554 unrelated French individuals from the general population. Typing of FRAXAC2 and sizing the CGG repeat were performed on selected individuals carrying the expected alleles and haplotypes.
Analysis of CA Repeats
Polymerase chain reaction (PCR) amplifications were performed using the oligonucleotide primers described by Verkerk et al. [4] for DXS548, and by Richards et al. [8], for the FRAXAC2 locus. PCR products were migrated on a sequencing gel, transferred onto a nylon membrane, and hybridized with a 32P-5′-end-labeled oligonucleotide (CA)10. Hybridization was performed at 50 °C for 2 h and the membranes were washed in 2 × SSC/0.1 % SDS for 10 min at room temperature and 5 min at 50 °C.
Sizing of CGG Repeats
PCR of the CGG repeats was performed according to Fu et al. [5], except that we used 200 µM 7-deaza-dGTP instead of 50 µM dGTP plus 150 µM 7-deaza-dGTP. PCR products were comigrated with a sequencing reaction of exon 11 of the CFTR gene [14] as size marker. In some cases, PCR of the CGG repeats was performed according to Erster et al. [15]: PCR products were migrated on a sequencing gel, transferred onto a nylon membrane and hybridized with a 32P-end-labeled oligonucleotide (CGG)7. Hybridization was performed at 55°C for 2 h and the membranes were washed in 2 × SSC/0.1% SDS for 10 min at room temperature and 10 min at 55 °C.
Statistical Methods
Analysis of variance (ANOVA) including all three groups was performed and the significance was tested by the F test. The mean sizes in the different groups were compared by a Student t test, after a common estimation of the overall variance.
Results
Of the 554 chromosomes tested with DXS548, 52 (9%) carried the DX204 allele, 66 (12%) carried the DX196 allele and 436 (79%) carried other alleles, mostly DX194. These proportions are similar to those found in a previous study of 188 unrelated normal chromosomes: 9, 14 and 77%, respectively [10]. The 118 chromosomes carrying DX204 or DX196 were analyzed at the FRAXAC2 locus. Finally, 31 chromosomes were identified carrying one of the two selected high-risk haplotypes, DX204-AC155 (3%) or DX196-AC151 (3%), and were subsequently analysed at the CGG locus. These percentages were also not significantly different from those found in a previous study of 153 unrelated normal chromosomes [10].
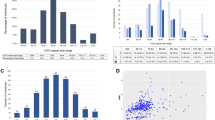
The sizes of the CGG repeats in these 31 high-risk haplotypes are shown in table 1. Thirty-two chromosomes carrying haplotypes other than DX204-AC155 or DX196-AC151 were also analyzed at the CGG locus.
Comparison of the CGG in the three groups by analysis of variance showed strong heterogeneity (F[2,61] = 12.6; p ≪ 10−3). In the 17 DX204-AC155 haplotypes, the mean size of the CGG locus was 38.4 ± 7.6 repeats. This value is significantly higher than the mean size in the 14 DX196-AC151 haplotypes (28.3 ± 5.1 repeats; p < 10−3) or the 32 control haplotypes (29.4 ± 6.7 repeats; p < 10−3). There was no difference between DX196-AC151 and control haplotypes (p ≈ 0.4).
In three chromosomes (individuals 7, 9 and 51), we found CGGs of 50, 55 and 59 repeats, respectively. These values are very close to 54, the normal premutation limit defined by Fu et al. [5]. Two of these chromosomes carried a DX204-AC155 high-risk haplotype, but the other was found in the third group and carried the DX196-AC155 haplotype. This haplotype is rare in the general and fragile X populations (its frequency was found to be approximately 1 and 3%, respectively) and may derive by crossing-over from a DX204-AC155 haplotype previously enlarged at the CGG repeat. However, this haplotype may also derive from another at-risk haplotype by mutation at FRAXAC2 or DXS548.
When possible, the stability of the CGG alleles was examined in siblings of individuals carrying high-risk haplotypes. The CGG alleles were stably inherited except in individual 9, a male carrying a premutated allele (55 repeats) transmitted to his daughter with one more repeat (56 repeats).
Discussion
The existence of a founder effect in fragile X syndrome suggests that an initial genetic event has occurred on a few occasions in human history and has been responsible for the risk of developing further expansion of the CGG repeat. Among the genetic events able to generate instability of CGG and consequently to increase the risk of expansion, the increased size of the repeat was expected to be very likely. Our results show that this is the case in DX204-AC155 haplotypes, where the mean CGG is approximately 10 repeats longer than in other haplotypes. A DX196-AC155 chromosome has been found with a CGG of 59 repeats, a value in the premutation range. This chromosome may be derived by crossing-over from a DX204-AC155 haplotype previously expanded at the CGG repeat. If we assume that this chromosome was a former DX204-AC155 haplotype, 100% of the CGG >35 repeats were found associated with this haplotype. This suggests that a primary genetic event occurred on the DX204-AC155 chromosome and generated a new enlarged CGG allele. Such a genetic event differed from the normal production of a polymorphism (for instance by DNA polymerase slippage), because it disrupted the distribution of alleles by generating a new mode with a dramatically high variance, reflecting great instability (fig. 1). Unequal crossing-over, creating the DX204-AC155 at-risk haplotype, would account for this disruption that has generated a reservoir of large CGG repeats associated with the DX204-AC155 haplotype, from which DNA polymerase slippage could lead to recurrent premutations as well as shorter alleles ranging in the normal size. This intermediate range, from which fragile chromosomes may be derived, was previously postulated on the basis of population data, to reconcile the high incidence of fragile X syndrome with the absence of de novo mutations [16,17].
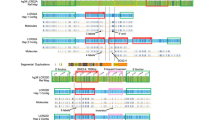
Distribution of the number of repeats in DX204-AC155 haplotypes and others, including DX196-AC151, in the 63 individuals tested. The clear departure (p < 10−3) between DX204-AC155 (mean size 38.4 ± 7.6 repeats) and other haplotypes (mean size 29.1 ± 6.2 repeats) suggests that a primary event, probably unequal crossing-over, disrupted the bimodal distribution of normal haplotypes. It is also clear that this primary event increased the instability of the repeats, as shown by the spread of the variance.
Because DX204-AC155 haplotypes account for only 14% of fragile X chromosomes [10], they cannot be considered to be the reservoir of all the fragile X mutations, as has been shown in myotonic dystrophy (MD) for the alleles carrying CTG repeats ranging from 19 to 30 triplets that represent 78% of MD chromosomes [18]. Moreover, at least six founder chromosomes must be hypothesized to account for present-day fragile X chromosomes [10]. It is thus clear that most of the fragile X chromosomes are derived from ancestral haplotypes other than DX204-AC155.
We observed no CGG with more than 35 repeats in the high-risk DX196-AC151 haplotype or any other haplotype. Two hypotheses may account for this observation. Firstly, the fragile X mutations not associated with DX204-AC155 may result from a one-step process, i.e. an unequal crossing-over leading directly from the normal (<35) to the premutation range. The absence of interspersed AGG triplets in these founder chromosomes could make them so unstable that the size jumps straight from the normal 6–35 range to more than 54 repeats, as has been suggested by the observation that exact repeats are less stable than interrupted repeats [2–4]. Secondly, it is possible that the intermediate range was not observed because of the small number of chromosomes analyzed, especially if the unequal crossing-over generated the high-risk haplotype in a pool of DX196-AC151 haplotypes that preexisted in the normal population. In this case, the high-risk DX196-AC151 chromosomes would be blended in low-risk DX196-AC151 chromosomes, resulting in the nondetection of increased CGGs. In contrast, we were able to document an increase in the CGG in DX204-AC155 because the crossing-over probably created this founder chromosome.
Mutations caused by expanding trinucleotide repeats have also been discovered in MD [19, 20], in Kennedy’s disease [21] and more recently in Huntington’s disease [22] and spinocerebellar ataxia type 1 [23]. Analysis of an insertion/deletion polymorphism close to the MD locus showed a clear-cut difference in the distribution of an allele for which the larger size of the CTG accounts for the observed intermediate 19–30 repeats range [18]. As in fragile X syndrome, this finding also suggests that the transition from the normal (< 18) to the abnormal (>36) range occurred by a multistep process, for instance unequal crossing-over followed by DNA polymerase slippage. In these two repeat-expansion disorders, there is an intermediate status that constitutes a reservoir of further pathological chromosomes. Analyses of linkage disequilibria in Huntington’s chorea, Kennedy’s syndrome and spinocerebellar ataxia I are not yet documented but perhaps they will lead to similar observations. If confirmed, our finding of an intermediate range, subject to further expansion, may lead to the development of more appropriate screening and counseling of the general population.
The DX204-AC155 chromosomes carry a CGG that corresponds to the S allele described by Morton and Macpherson [17], an allele differing from the unstable premutated Z allele and from the normal stable N allele, by its moderate instability. The absence of an S allele in fragile X families may be due to the time (in generations) needed by the CGG to expand and reach a size ≥ 54 repeats.
References
Vincent A, Heitz D, Petit C, Kretz C, Oberlé I, Mandel JL: Abnormal pattern detected in fragile X patients by pulse field gel electrophoresis. Nature 1991;349:624–626
Oberlé I, Rousseau F, Heitz D, Kretz C, Devys D, Hanauer A, Boué J, Bertheas MF, Mandel JL: Instability of a 550 bp DNA segment and abnormal methyiation in fragile X syndrome. Science 1991;252:1097–1102
Kremer EJ, Pritchard M, Lynch M, Yu S, Holman K, Baker E, Warren ST, Schlessinger D, Sutherland GR, Richards RI: Mapping of DNA instability at the fragile X to a trinucleotide repeat sequence p(CGG)n. Science 1991;252:1711–1714
Verkerk AJMH, Pieretti M, Sutcliffe JS, Fu YH, Kuhl DPA, Pizzuti A, Reiner O, Richards S, Victoria MF, Zhang F, Eussen BE, van Om-men GJB, Blonden LA, Riggins GJ, Chastain JL, Kuns CB, Galjaard H, Caskey CT, Nelson DL, Oostra BA, Warren ST: Identification of a gene (FMR1) containing a CGG repeat coincident with a breakpoint cluster region exhibiting length variation in fragile X syndrome. Cell 1991;65:905–914
Fu YH, Kuhl DPA, Pizzuti A, Pieretti M, Sutcliffe JS, Richards S, Verkerk AMJH, Holden JJA, Fenwick RG Jr, Warren ST, Oostra BA, Nelson DL, Caskey CT: Variation of the CGG repeat at the fragile X site results in genetic instability: Resolution of the Sherman paradox. Cell 1991;67:1047–1058
Brown WT, Houck GE, Jeziorowska A, Levinson FN, Ding X, Dobkin C, Zhong N, Henderson J, Brooks SS, Jenkins EC: Rapid fragile X carrier screening and prenatal diagnosis using a nonradioactive PCR test. JAMA 1993;270:1569–1575
Riggins GJ, Sherman SL, Oostra BA, Sutcliffe JS, Feitell D, Nelson DL, van Oost BA, Smits APT, Feliciano JR, Pfendner E, Kuhl DPA, Caskey CT, Warren ST: Characterization of a highly polymorphic dinucleotide repeat 150 kb proximal to the fragile X site. Am J Med Genet 1991;43:237–243
Richards RI, Holman K, Kremer E, Lynch M, Pritchard M, Yu S, Mulley J, Sutherland GR: Fragile X syndrome: Genetic localisation by linkage mapping of two microsatellite repeats FRAXAC1 and FRAXAC2 which immediately flank the fragile site. J Med Genet 1991;28:818–823
Richards RI, Holman K, Friend K, Kremer E, Hillen D, Staples A, Brown WT, Goonewardena P, Tarleton J, Schwartz C, Sutherland GR: Evidence of founder chromosomes in fragile X syndrome. Nature Genet 1992;1:257–260
Oudet C, Mornet E, Serre JL, Thomas F, Lentes-Zengerling S, Kretz C, Deluchat C, Tejada I, Boué J, Boué A, Mandel JL: Linkage disequilibrium between the fragile X mutation and two closely linked CA repeats suggests that fragile X chromosomes are derived from a small number of founder chromosomes. Am J Hum Genet 1993;52:297–304
Buyle S, Reyniers E, Vits L, De Boulle K, Handing I, Wuyts FLE, Deelen W, Halley DJJ, Oostra BA, Willems PJ: Founder effect in a Belgian-Dutch fragile X population. Hum Genet 1993;92:269–272
Oudet C, von Koskull H, Peippo M, Mandel JL: Striking founder effect for the fragile X syndrome in Finland. Eur J Hum Genet 1993;1:181–189
Zhong N, Dobkin C, Brown WT: A complex mutable polymorphism located within the fragile X gene. Nature Genet 1993;5:248–312
Kerem B, Zielenski J, Markiewicz D, Bozon D, Gazit E, Yahav J, Kennedy D, Riordan JR, Collins FS, Rommens JM, Tsui LC: Identification of mutations in regions corresponding to the two putative nucleotide (ATP)-binding folds of the cystic fibrosis gene. Proc Natl Acad Sci USA 1989;87:8447–8451
Erster SH, Brown WT, Goonewardena P, Dobkins CS, Jenkins EC, Pergolizzi RG: Polymerase chain reaction analysis of fragile X mutations. Hum Genet 1992;90:55–61
Chakravarti A: Fragile X founder effect? Nature Genet 1992; 1:237–238.
Morton NE, Macpherson JN: Population genetics of the fragile X syndrome: Multiallelic model for the FMR1 locus. Proc Natl Acad Sci USA 1992;89:4215–4217
Imbert G, Kretz C, Johnson K, Mandel JL: Origin of the expansion mutation in myotonic dystrophy. Nature Genet 1993;4:72–76
Mahadevan M, Tsilfidis C, Sabourin L, Shutler G, Amemiya C, Jansen G, Neville C, Narang M, Barcelo J, O’Hoy K, Leblond S, Earle-Macdonald J, de Jong PJ, Wieringa B, Korneluk RG: Myotonic dystrophy mutation: An unstable CTG repeat in the 3′ untranslated region of the gene. Science 1992;255:1253–1255
Fu YH, Pizzuti R, Fenwick RG, King J, Rajnarayan S, Dunne PW, Dubel J, Nasser GA, Ashizawa T, de Jong P, Wieringa B, Korneluk R, Perryman MB, Epstein HF, Caskey CT: An unstable triplet repeat in a gene related to myotonic muscular dystrophy. Science 1992;255:1256–1258
La Spada AR, Wilson EM, Lubahn DB, Harding AE, Fischbeck H: Androgen receptor gene mutations in X-linked spinal and bulbar muscular atrophy. Nature 1991;352:77–79
Huntington’s Disease Collaborative Group: A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell 1993;72:971–983
Orr HT, Chung MY, Banfi S, Kwiatkowski TJ Jr, Servadio A, Beaudet AL, McCall AE, Duvick LA, Ranum LPW, Zoghbi HY: Expansion of an unstable trinucleotide CAG repeat in spinocerebellar ataxia type 1. Nature Genet 1993;4:221–226
Acknowledgements
This work was supported by the Association Française contre les Myopathies (support to MM, AB and EM).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Montagnon, M., Bogyo, A., Deluchat, C. et al. Transition from Normal to Premutated Alleles in Fragile X Syndrome Results from a Multistep Process. Eur J Hum Genet 2, 125–131 (1994). https://doi.org/10.1159/000472352
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1159/000472352