Abstract
The main protease (Mpro) of SARS-CoV-2 is critical for viral function and a key drug target. Mpro is only active when reduced; turnover ceases upon oxidation but is restored by re-reduction. This suggests the system has evolved to survive periods in an oxidative environment, but the mechanism of this protection has not been confirmed. Here, we report a crystal structure of oxidized Mpro showing a disulfide bond between the active site cysteine, C145, and a distal cysteine, C117. Previous work proposed this disulfide provides the mechanism of protection from irreversible oxidation. Mpro forms an obligate homodimer, and the C117-C145 structure shows disruption of interactions bridging the dimer interface, implying a correlation between oxidation and dimerization. We confirm dimer stability is weakened in solution upon oxidation. Finally, we observe the protein’s crystallization behavior is linked to its redox state. Oxidized Mpro spontaneously forms a distinct, more loosely packed lattice. Seeding with crystals of this lattice yields a structure with an oxidation pattern incorporating one cysteine-lysine-cysteine (SONOS) and two lysine-cysteine (NOS) bridges. These structures further our understanding of the oxidative regulation of Mpro and the crystallization conditions necessary to study this structurally.
Similar content being viewed by others
Introduction
During the COVID-19 pandemic, the SARS-CoV-2 main protease (Mpro, nsp5 or 3CLpro) emerged as a key antiviral target and focus of intense study1,2,3. Mpro plays a central role in the SARS-CoV-2 replication cycle, as the viral genome codes for polyproteins that must be cleaved into individual protein units to support viral function. Mpro processes at least 11 known sites along polyproteins 1a and 1ab, including its own N- and C-termini1, and is therefore essential for viral replication. This key role in replication, along with the historical success of viral protease inhibitors, the lack of any similar human protein, and prior work on SARS-CoV-1 Mpro, has made SARS-CoV-2 Mpro the target of several drug discovery programs. These efforts have already yielded an FDA approved molecule, nirmatrelvir4. Given the persistence of the COVID-19 virus and the possible emergence of future pathogenic coronaviruses, it is imperative we develop a deeper understanding of Mpro and its role in viral function.
Mpro’s activity is regulated by multiple mechanisms, though we have a poor understanding of how these support viral fitness. Most prominently, at sufficiently high concentrations, the enzyme forms a homodimer. Dimerization enhances the catalytic rate, effectively turning Mpro from an inactive form into an active one5. Structural work suggests this concentration-dependent regulation is not an evolutionary accident. Specifically, Mpro adopts a chymotrypsin-like fold but has a distinct dimerization domain at its C-terminus that many other chymotrypsin-like enzymes lack6. Studies of the truncated enzyme without this domain, as well as of the domain in isolation, have demonstrated it is both necessary and sufficient for dimer formation7. This suggests that this dimerization domain enables regulation of Mpro’s catalytic rate based on the concentration of free enzyme in the cell.
In addition to regulation via dimerization, Mpro has been shown to be sensitive to the local redox environment. Including the active site cysteine, the protein sequence contains 12 cysteine residues (~4% of total), an unusually high number8. Under mildly reductive conditions all cysteines are reduced, and the protein’s catalytic rate is maximized, suggesting this is the active form of the enzyme found in a cellular context9. Upon oxidation, a remarkable and growing number of modifications have been reported by both structural and mass spectrometry studies, including glutathionylated C30010, a peroxy-C145, an N-ethylmaleimide modified C145 and C15611, a SONOS bridge between C22, C44, and K6112,13, and a disulfide link between C117 and C1459,14.
Given the complexity of viral replication in human hosts, the prevalence or role of these modifications in the viral replication cycle has remained unclear. Oxidative stress in the cell has been shown to regulate the function of other viruses15, most notably HIV16,17,18, and early in the pandemic oxidative stress was hypothesized to play a central role in COVID-19 pathogenesis19. It has even been speculated that robustness to oxidative environments might enable corona or other viruses to survive in bat hosts, which are known to exhibit unusual oxidative cellular conditions10.
Among the known oxidative modifications of Mpro, the C117-C145 disulfide modification particularly is notable. Funk and colleagues recently performed a systematic study of the behavior of Mpro under oxidative conditions and highlighted this modification as uniquely functional9. They produced single-point cysteine-to-serine mutants for each cysteine in Mpro. Of all these mutants, they found C117S was the only mutant that did not recover activity after being exposed to H2O2 and then re-reduced with DTT. This suggests C117 may have a special role in protecting the active site C145 from oxidative damage. Supporting this, Tran et al. recently reported the structure of an oxidized mutant of Mpro, the H163A variant, that contained a disulfide bond linking C117 to C14514. The H163A modification inactivates the enzyme, even under reducing conditions. Removal of the H163 sidechain disrupts a π-stacking interaction with F140, which is part of the loop that forms the transition state-stabilizing oxyanion hole. Disruption of the oxyanion hole was observed in structures of both the oxidized (C117-C145) and reduced (no disulfide) enzyme, explaining the loss in activity. However, it was unclear how the C117-C145 disulfide in the H163A variant related to the behavior of the wild type.
To address these questions, we determined the structure of Mpro with the C117-C145 disulfide modification under mildly oxidizing conditions, providing a structural understanding of how this disulfide can protect the enzyme from irreversible oxidation. We find that oxidized protein only crystallizes in a more loosely packed, orthorhombic lattice, whereas the reduced protein forms a monoclinic lattice under the same crystallization conditions. Seeding with these orthorhombic crystals enabled us to crystallize Mpro exhibiting a set of NOS and SONOS oxidative modifications.
Results
An orthorhombic lattice is flexible enough to produce crystals with oxidative modifications
By delivering streams of microcrystals into the x-ray focus of the SPB/SFX instrument of the European XFEL, we obtained diffraction data yielding two crystal structures of Mpro, one active/reduced structure and one inactive/oxidized structure (supplementary Table S1). The crystals merged to determine these two structures were grown in two separate batch crystallization experiments. In the first, the resulting structure was fully reduced due to the presence of 1 mM TCEP during purification. In the second experiment, TCEP was omitted, and Mpro exposed to air over time spontaneously crystallized into a different space group, despite being crystallized under otherwise the same conditions (buffer, temperature, concentrations). Specifically, our reduced-Mpro crystals formed a monoclinic lattice with C2 symmetry. These crystals contain the native homodimer, with a single protomer in the asymmetric unit and the dimer completed by crystallographic symmetry. Protein subjected to oxidation by exposure to air exhibits a covalent disulfide bond between C117 and C145 and forms crystals in space group P212121, with the asymmetric unit consisting of the entire homodimer (protomer A-to-B all-atom RMSD: 0.96 Å). The orthorhombic lattice exhibits a looser overall packing and higher solvent content (Fig. 1, supplementary Table S1). Both datasets were collected at room temperature.
A Reduced protein under our crystallization conditions, containing TCEP, results in monoclinic (C2) protein crystals (PDB ID: 7PXZ). In a separate batch crystallization experiment, after oxidation by air and in the absence of reducing agent, the same protein spontaneously forms crystals with an orthorhombic (P212121) lattice and exhibits a disulfide link between C117 and C145 (PDB ID: 7PZQ). By producing seed crystals from this oxidized protein, however, we were able to obtain two further structures in the same orthorhombic lattice: first, using reduced protein, a structure with reduced C117/C145 (PDB ID: 7Z2K), and second, using the same reduced protein but with the addition of a sulfonated calpeptin ligand, a structure exhibiting NOS and SONOS crosslinks (PDB ID: 7Z3U, which is published in ref. 20). B Visualization of the orthorhombic and monoclinic lattices, with the solvent content highlighted by map-channels47. The packing and crystal contact patterns are substantially altered, with the orthorhombic lattice exhibiting significantly larger solvent channels and an overall higher solvent content.
While crystallization conditions for the oxidized and reduced crystals are the same, the lattices and crystal morphologies obtained differ (Fig. 1, supplementary Fig. S1). This makes a direct comparison of reduced and oxidized structures challenging, as we could not control for differences due to oxidation state vs. crystal packing. Therefore, we attempted to obtain a reduced structure in the orthorhombic lattice seen in our oxidized crystals. By seeding reduced protein with crystals of the oxidized form, we were successfully able to generate crystals of reduced protein in the orthorhombic lattice (Fig. 1). As our XFEL beamtime had concluded by this time, data for these crystals were collected under cryogenic conditions at PETRA-III beamline P11 (supplementary Table S1). The cryogenic conditions cause a contraction of the lattice and reduction of the solvent content by 4–5% as compared to room temperature collection (supplementary Table S1). The molecular structure of the enzyme in the reduced state is similar in both the monoclinic (XFEL/RT) and the orthorhombic (synchrotron/100 K) lattices (all-atom RMSD: 1.56 Å). Both structures are used here as a basis of comparison to elucidate changes due to oxidation.
Finally, in conjunction with our ongoing work to develop Mpro inhibitors, we employed our oxidized orthorhombic seeds in a co-crystallization experiment with Mpro bound to a sulfonated calpeptin derivative. The resulting structure was reported in a paper describing the ligand binding pose20. Unexpectedly, the same structure exhibits a rich pattern of oxidative modifications, which we discuss here. Protomer A contains a SONOS bridge involving C22, C44, and K61, whereas protomer B shows only a NOS bridge involving C22 and K61 at the same site. Both modifications are consistent with previous reports12,13. In addition, 2mFo-DFc maps unambiguously show a NOS bridge between K102 and C156 in protomer B, not previously described in the literature, and suggest partial occupancy of the same modification in protomer A.
Disulfide formation in Mpro precludes catalysis and disrupts the dimer interface
By oxidizing Mpro via air exposure, we obtained structures with a disulfide bond between C117 and C145. To understand the structural changes that occur upon formation of the C117-C145 disulfide bond (Fig. 2), we determined two reduced reference structures. The first was obtained at room temperature with XFEL radiation, identical to the data collection conditions of our C117-C145 structure, but crystallized in a different space group (C2). The second reference structure is in the same space group as the C117-C145 structure (P212121), following seeding with crushed oxidized crystals, and was obtained at 100 K.
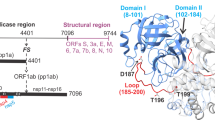
A aligned overlay of reduced (orange: monoclinic/room temperature, PDB ID: 7PXZ; yellow: orthorhombic/100 K, PDB ID: 7Z2K) and oxidized (cyan: orthorhombic/room temperature, PDB ID: 7PZQ) structures, with one monomer of the Mpro dimer shown as surface. Oxidation of the active site cysteine, C145, results in B disulfide bridge formation with C117 and displacement of N28 (density: oxidized 2mFo-DFc at 1 RMSD). Colocalization of C117 and C145 requires C displacement of C-terminal residues 301–306 from the dimer interface and is correlated with D a shift of the dimerization domain and disruption of the stabilizing interactions between the two protomers. E MSA showing N28, C117, and C145 are conserved across related coronaviruses. N28 and C145 are absolutely conserved in the set studied. C117 is partially conserved, but where it is not, another cysteine is present in either position 116 or 142 (magenta) that could conceivably fulfill the same role.
In the reduced form, the catalytically active cysteine C145 sits on a loop in the active site pocket, while C117 forms part of a β-hairpin about 8 Å away (Cα-to-Cα). Oxidative crosslinking of these residues relocates both to a location approximately in the middle of their reduced positions (5.1 Å Cα-to-Cα). This disrupts the β-hairpin motif containing C117 and displaces the conserved N28, which in the reduced structure sits between C145 and C117 but in the oxidized structure undergoes a rotamer shift to make space for the disulfide bridge (Fig. 2). This residue was identified as essential for dimerization and enzymatic activity in SARS-CoV-1 Mpro 21. The rotameric change of N28 was predicted by MD simulations performed by Funk et al. and is confirmed by our structures9. Moreover, the position of the loop containing the active cysteine, spanning S139–S147, rearranges upon disulfide formation (supplementary Fig. S2). Finally, disulfide formation partially buries the active site cysteine, which has a solvent-exposed area of 24.3 Å2 in the reduced structure but 17.5 Å2 and 13.9 Å2 for the two molecules in the asymmetric unit, A and B respectively, in the oxidized structure.
In protomer A of our oxidized structure, C145 shows the residual population at the same position it occupies in the reduced structure. The electron density of this residual population was sufficient to model an alternative conformation, resulting in a refined structure with 55% occupancy of the disulfide conformer and 45% population of the reduced conformation. In protomer B, the reduced conformation is insufficiently populated to generate a confident model, and our structure contains a fully occupied disulfide. No evidence of other oxidative modifications was observed in our electron density maps.
The structural rearrangements required to bring C117 and C145 together require a series of long-range structural changes that disrupt the dimer interface (Fig. 2). In the reduced state, the C-termini form part of the dimer interface adjacent to the β-hairpin containing C117, but upon crosslinking of C117 and C145, C-terminal residues 301–306 become disordered due to the loop rearrangements necessary to bring the two cysteines together.
This ejection of the C-termini from the dimer interface is accompanied by a shift in the entire dimerization domain, which contains contacts that bridge the two protomers. As a result, the entire dimer interface is less well-packed in the oxidized structure as compared to its reduced counterpart. The surface area that forms the dimer interface is estimated to be 1301 Å2 and 1283 Å2 for protomers A and B, respectively, in the orthorhombic reduced structure. This interfacial area decreases to 1198 Å2 and 1259 Å2 in the disulfide-containing structure. Notably, the loop formed by residues S284, A285, and L286, which packs tightly with the same residues on the symmetric protomer in the reduced state, is disrupted in the oxidized structure (Fig. 3). In the reduced structure, this loop forms a tight zipper-like packing interface with the opposite protomer, but in the oxidized structure this zipper is out of register and does not form a tight interface (Fig. 3). This disruption of the dimer interface suggests Mpro’s dimer affinity is weakened upon oxidation, as recently reported by Funk et al.9.
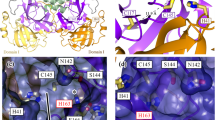
Shown is the region where the loop containing S284, A285, and L286 forms a hydrophobic zipper with the same residues on the opposite dimer-forming protomer. All four structures reported are drawn, A reduced/C2 (orange, PDB ID: 7PXZ), reduced/P212121 (yellow, PDB ID: 7Z2K), NOS/SONOS (purple, PDB ID: 7Z3U), and B C117-C145 (teal, PDB ID: 7PZQ). Only the C117-C145 structure shows a disruption of this dimer interface. For these three residues, the buried surface area decreases from ~110 Å2 for the structures in A to 87 and 85 Å2 for molecules A and B of the C117-C145 structure shown in B, respectively. Structures were aligned by minimizing all heavy atom RMSD prior to visualization. Surfaces are the solvent-accessible (Connolly) surface computed with the PyMOL Molecular Graphics System (2.0, Schrödinger LLC)48.
Analytical size exclusion chromatography confirms weakened dimer affinity
To test if the disruption of the dimer interface observed in our C117-C145 structure translates into a reduction of the dimerization affinity, we performed analytical size exclusion chromatography (Fig. 4). Freshly purified protein, exposed to air for only a few hours, exhibited a dimerization dissociation constant (KD) of 2.3 µM. Addition of 1 mM TCEP in the running buffer produced no significant change, with an estimated KD of 2.1 µM. However, incubating the protein in 1 mM hydrogen peroxide prior to injection significantly decreased the dimer stability, resulting in an order of magnitude increase of the measured KD to 19 µM. Further, upon exposure to peroxide, the observed monomer peak elutes notably earlier, at approximately 0.49 column volumes vs. 0.47, suggesting oxidation has produced a less compact monomer species. Our results agree with analytical ultracentrifugation performed by Zhang et al. (KD~2.5 µM, reduced)2 and the SAXS measurements of Silvestrini et al. (KD~7 µM, reduced)22. They are qualitatively consistent with the analytical centrifugation measurements performed by Funk et al.9, who determined absolute KD values that are a factor of 10 smaller, but with the same relative order of magnitude decrease upon oxidation.
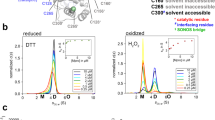
A Mpro dimerization was analyzed under three conditions: first, immediately after purification, without oxidizing or reducing agent (native, top); second, in the presence of 1 mM TCEP in the running buffer (middle), or after the protein was incubated with 1 mM peroxide for 5 h (bottom). Protein was logarithmically diluted between 100 µM and 780 nM and assayed by size exclusion. SEC traces are normalized by protein concentration so that they appear on the same scale; lower concentrations therefore exhibit visible baseline drift and noise. Distinct monomer and dimer peaks are seen for each sample/concentration combination, eluting at approximately 0.48 and 0.43 column volumes, respectively. The concentration of these species was estimated by the peak height at the positions indicated by vertical dashed lines. The peroxide-exposed monomer (middle dashed line) elutes earlier than the monomer peaks for native and TCEP-exposed protein (rightmost dashed line), suggesting this species is modified and may be structurally less compact than fully reduced monomer. B The monomer and dimer concentrations from A were fit to a two-state equilibrium model, \(2M\rightleftharpoons {D}\) (solid lines), with a single parameter, the KD. The determined KD parameters show that oxidation of Mpro by 1 mM peroxide (teal, KD 19 ± 14.8 µM) significantly weakens the dimer interface as compared to reduced protein (blue, KD of 2.1 ± 0.50 µM, 1 mM TCEP in the running buffer) or freshly purified “native” protein in the absence of exogenous reducing or oxidizing agents (black, KD 2.3 ± 0.63 µM). Errors reported are 95% confidence intervals assuming a Gaussian error model. The x-axis reports the total concentration of single Mpro protein chains. The y-axis reports the fraction of monomeric chains as determined from the traces in (A).
Observation of NOS and SONOS modifications upon co-crystallization with a sulfonated calpeptin ligand
During our ongoing structural studies of Mpro ligands, we sought to obtain a structure of Mpro bound to a ligand of interest, a sulfonated calpeptin derivative that reacts covalently with the catalytic cysteine in Mpro. Co-crystallization attempts of this ligand with reduced, monoclinic seeds failed to yield a high-resolution structure in our hands, instead forming small clusters of crystals that diffracted to low resolution (~5 Å) and could not be indexed. We hypothesized that the looser packing of the orthorhombic lattice provided by our oxidized seeds might better accommodate structural rearrangements caused by ligand binding. Subsequently, we attempted crystallization with our orthorhombic, oxidized seeds and obtained a high-resolution structure clearly showing bound ligand, which we refined against data up to 1.72 Å. Ligand density consistent with full occupancy was present in the active site of both monomers. Unexpectedly, however, the structure shows multiple NOS and SONOS modifications (Fig. 5).
Oxidative NOS and SONOS bridges are seen at four sites in the A two dimeric protomers that form both the asymmetric and biological unit (all panels, purple: oxidized NOS/SONOS structure in P212121, PDB ID: 7Z3U, published in citation 21; yellow: reduced reference in P212121 PDB ID: 7Z2K). B Protomer A exhibits a SONOS linkage between C22, K61, and C44 that distorts the structure from the reduced form. In contrast, C protomer B shows only a NOS linkage between C22 and K61 at this site, with the overall structure differing little from the reduced reference. D The electron density on protomer A between K102 and C156 is ambiguous, consistent with but not conclusively showing a NOS linkage at partial occupancy. In contrast, E the density at the same site on protomer B clearly shows a NOS bond at partial occupancy. F None of the residues participating in these linkages are strongly conserved besides C44. Densities shown as blue volumes are the 2mFo-DFc map at 1 RMSD. The same 2mFo-DFc map at 0.5 RMSD is overlaid as a light mesh for the K102-C156 NOS figures, to show partial occupancies more clearly.
The NOS and SONOS modifications exhibit a distinct asymmetry between the monomers that form the dimer in the asymmetric unit. Protomer A shows a SONOS linkage between C22, K61, and C44 (Fig. 5), which has been previously reported12,13. This modification results in a shift of the α-helix between E55 and K61 and disrupts the position of a loop between C44 and Y54, which is ordered in the reduced structure but becomes disordered upon SONOS formation. This disorder may be in part because the shifted loop can no longer form a backbone H-bonding contact between M49 and Q189, the latter of which sits on a flexible domain-connecting loop consisting of the residues V186 to G195.
In contrast, protomer B more closely resembles the reduced structure. It exhibits a NOS bridge between C22 and K61 (Fig. 5). The effect of this modification is less dramatic, with C22 and K61 separated by 7.2 Å in the reduced structure (Cα-to-Cα) but only 7.4 Å with the NOS bridge present.
Finally, both protomers A and B show evidence for a NOS bridge between K102 and C156 that connects neighboring β-sheets (Fig. 5, see supplementary Fig. S3 for supporting OMIT and isomorphous difference maps). This modification is clear in the density for protomer B, while the density in protomer A is ambiguous but consistent with a NOS bridge at low occupancy. The NOS modification at this site induces essentially no deviation from the reduced structure in the same space group, where K102 and C156 are in close proximity.
Discussion
Mpro appears to exhibit an unusually rich set of oxidation modifications, which have been revealed by structural and biochemical methods. While a response to oxidative stress has been implicated in virus biology in general, the possible physiological relevance of each of the observed oxidized states of Mpro remains a topic of ongoing investigation.
Our structure of C117-C145 modified Mpro provides a mechanistic model for several key observations regarding Mpro’s behavior upon change of redox state. Most notably, our structures provide a simple explanation as to why Mpro’s dimer affinity decreases by about an order of magnitude upon oxidation (Fig. 4)9. Our structure further confirms a key role of N28, which rotates to allow space for the C117-C145 disulfide bridge. N28 is highly conserved (Fig. 1), suggesting asparagine at this position is essential for viral fitness21. We assume that the small volume and hydrophilic nature of the carboxamide sidechain facilitates this conformational change, enabling Mpro’s ability to toggle between reduced and oxidized states.
While we have been able to grow large single crystals from air-oxidized protein in the orthorhombic space group (supplementary Fig. S1) and have studied these with synchrotron radiation, we have not observed clear C117-C145 disulfide density in those studies. In contrast, the diffraction of XFEL light from microcrystals clearly shows the C117-C145 disulfide. We considered the hypothesis that XFEL radiation may have allowed us to observe this modification via “radiation damage-free” data collection, as the x-ray exposure (<100 fs) is much more rapid than the nuclear motions required for the two cysteine sidechains to adopt significantly different positions following x-ray induced reduction23,24,25. In this model, structures obtained at the synchrotron would appear fully reduced, because during data collection at the synchrotron, x-ray-induced cleavage of the disulfide would reverse the structural changes that accompany C117-C145 crosslinking. This would require an implausibly large rearrangement of the structure under cryogenic conditions. The success of Tran et al. in capturing this disulfide with synchrotron radiation in the H163A variant14 is further evidence against this model. Their ability to obtain a structure clearly showing the C117-C145 disulfide suggests that this modification is not uniquely sensitive to reduction by X-rays.
Alternatively, it may be that the large crystals containing significant C117-C145 population are less well-ordered than their microcrystalline counterparts, and XFEL radiation allowed us to study such microcrystals. Our data only show that XFEL radiation is sufficient to observe the C117-C145 disulfide in wild-type Mpro, but we cannot conclude that such radiation is necessary to preserve the C117-C145 modification.
By studying the H163A Mpro mutant, Tran and colleagues were able to crystallographically observe the same C117-C145 disulfide bond we found in wild-type Mpro 14. In that work, they observed that the loss of the imidazole sidechain in H163 removed a favorable π-stacking interaction with F140, resulting in displacement of a mobile loop containing F140 (supplementary Fig. S2). Further, this loop motion displaces the N and C termini, resulting in dimer interface disruption. Importantly, the loop containing F140 forms the oxyanion hole, which stabilizes the transition state during proteolysis. In our disulfide-containing structure, the F140 loop does not reach the same extreme position as observed by Tran et al. but is disrupted from its conformation in the corresponding reduced structures (supplementary Fig. S2). Our work therefore supports the hypothesis of Tran et al., that C117-C145 can form in a similar fashion in both the wild type and H163A enzymes but demonstrates that the extensive remodeling of the F140 loop in the H163A structure is particular to that variant.
Complementing these structures of the C117-C145 disulfide, Funk et al. reported that Mpro C117S was the only C-to-S mutant that failed to recover activity after exposure to oxidative conditions followed by reduction9. Our structures illustrate how, upon oxidation, the catalytic C145 moves from a solvent-exposed conformation to a buried, disulfide conformation. Our structure, alongside these previous findings and the conserved nature of cysteines at positions 117 and 145, implicate this modification in a regulatory response to an oxidative environment. Our results support the idea that the C117-C145 disulfide provides a protective mechanism against oxidative damage by making harsher, irreversible oxidation of the catalytic C145 to sulfinic and sulfonic acids impossible26.
Seeding with crystals containing the C117-C145 disulfide enables kinetic control over the crystallization lattice, allowing us to obtain a ligand-bound structure that shows NOS and SONOS bridges. We considered the hypothesis that ligand binding might facilitate these modifications. Yang and colleagues, however, recently presented nine Mpro structures exhibiting the C22-K61-C44 SONOS bond. Five contain bound inhibitors but four show no ligand of interest13, demonstrating that ligand binding is not necessary to observe SONOS modifications in Mpro crystals. Further, the NOS bond at K102-C156, which has not been reported previously, is far from the active site and seems unlikely to be influenced by the binding of a ligand in that pocket. As no oxidizing agents were added to the crystallization experiment, we attribute NOS/SONOS formation to molecular oxygen introduced by air exposure. The frequency and diversity of NOS/SONOS modifications observed in Mpro suggest these crosslinks may have a functional role in regulating the enzyme’s function in oxidative environments.
The fact that Mpro is the premier target for anti-COVID-19 small molecule therapeutics warrants further investigation into the possible impact of these oxidative modifications on viral fitness. While our structures provide mechanistic insight into the regulation of the enzyme by oxidation and dimerization, we must now understand how regulation based on oxidative stress or protein concentration impacts the viral lifecycle in vivo. Such work will provide deeper insight into viral biology and – hopefully – open new opportunities to disrupt that biology with life-preserving medicines.
Methods
Protein production and purification
The gene encoding Mpro was overexpressed in E. coli and purified for subsequent crystallization using previously published protocols and plasmid constructs2. Specifically, cell pellets containing recombinant protein were lysed in 20 mM TRIS buffer, pH 7.8, supplemented with 150 mM NaCl and 10 mM imidazole using a homogenizer. After removal of insoluble cell matter by ultracentrifugation, a nickel NTA column was used to purify the Mpro -histidine-tag fusion protein. Following imidazole elution, the protein buffer was changed to 20 mM TRIS, pH 7.8, 150 mM NaCl, 1 mM TCEP using a PD10 column, and the histidine tag was cleaved by 3C protease overnight. Subsequently, the histidine tag and the 3C protease were removed using a nickel NTA column. For the reduced form of Mpro a final size exclusion chromatography was performed with an S200 Superdex column using 20 mM TRIS, pH 7.8, 150 mM NaCl, 1 mM TCEP, and 1 mM EDTA, while for the oxidized form TCEP was omitted.
Crystallization experiments
Mpro microcrystals were grown using seeded batch crystallization in the XBI laboratories of the European XFEL27. Seed crystals were grown from protein purified in the presence of 0.5 mM TCEP, using a sitting drop geometry by combining 250 nL Mpro protein solution (6.25 mg/ml) and 250 nL precipitant (25% PEG1500, 0.1 M MIB buffer pH 7.5, 5% DMSO), as reported previously28. A seed stock was produced by adding the resulting Mpro crystals to a reaction tube containing a glass bead (Beads-for-Seeds, Jena Bioscience) and vortexing periodically for 5 s with subsequent incubation at room temperature. For the microcrystal batch crystallization, 250 μL glass seed beads were added to a 1.5 mL reaction tube, which was then filled with 900 μL precipitant solution (25% PEG1500, 0.1 M MIB buffer pH 7.5, 5% DMSO) mixed with 100 μL seed stock and 100 μL Mpro protein solution (35 mg/ml). Subsequently, crystals were grown in a shaker at 18 °C at 900 rpm overnight. The reduced and oxidized forms were crystallized separately. Resulting crystals were thin plates with a size ranging from 3–15 μm. Crystal concentration was adjusted by allowing the crystals to settle overnight and removing supernatant accordingly. Final crystal slurry was filtered through a 30 μm mesh gravity filter (Sysmex CellTrics) before injection.
Protein crystals for single-crystal rotation experiments were produced as previously reported28, using orthorhombic seeds and reduced protein at 6.25 mg/mL. For the ligand free and S-calpeptin containing crystallization experiments, the same reduced protein batch was used. The S-calpeptin compound was dried in the well prior to crystallization mixture addition, yielding a maximum concentration of 5 mM.
Instrumentation
SFX experiments (7PXZ, 7PZQ) were performed at the SPB/SFX instrument29 in April 2021 as a part of proposal 2696. The size of the mirror-focused focal spot in the interaction region was estimated to be 4 × 4 µm2 FWHM diameter based on optical imaging of single shots using a 20 µm thick Ce:YAG screen. The x-ray pulse energy was in the range of 1.2–3.5 mJ at 9.3 keV. Diffraction from the sample was measured using an AGIPD30 of 1 megapixel located 117.7–118.6 mm downstream of the sample interaction region, with the unused direct beam passing through a central hole in the detector to a beam stop further downstream. The resolution at the edge of the AGIPD was 1.8 Å, and 1.6 Å data were obtained by integrating Bragg reflections up to the detector corner. Experiment control was provided by Karabo31.
We used double-flow focusing nozzles (DFFN) for sample delivery32,33. The DFFN had an inner diameter of 75 μm and a liquid jet was established by applying 35 mg/min helium flow, 25 μL/min ethanol flow, and 15–20 μL/min sample flow. We measured the jet diameter to be about 4.5 μm, with a flow rate of 40–45 μL/min under identical conditions to those used for the experiment. This translates into a jet speed of approximately 43 m/s34. During injection, sample was at room temperature, approximately 20 °C.
Rotation experiments (7Z2K, 7Z3U) were performed at PETRA-III beamline P11, delivering a 100 µm beam of 12 keV x-rays focused by a paired KB mirror system attenuated to 30% transmission35. Crystals were mounted robotically on a single-axis goniometer and held at 100 K using a cryojet (Oxford). During data collection, samples were rotated 200 degrees with frames read out from a DECTRIS Eiger detector at a distance of 200 mm every 0.2 degrees, for a total of 1000 images per crystal. Total dose per collection was approximately 1.05 MGy as determined by a calibrated diode measurement of x-ray flux (0.7•1012 ph/s at 100% transmission).
Data analysis
During SFX experiments, online monitoring of the running experiment was performed with Karabo31 v2 and OnDA36 v22. The AGIPD geometry was refined against lysozyme data taken at the beginning and end of every shift using geoptimiser37. Preprocessing of images was performed with Cheetah38 v2019.1 and subsequent crystallographic analysis was done with CrystFEL v0.9.139. The MOSFLM algorithm was used for preliminary indexing40, but all reported results used xgandalf 41. Serial data merging was performed with partialator using the unity model. Data from rotation experiments with single crystals were processed with XDS42 v10.01.2022. Prior to model building and refinement, XFEL datasets were resolution-truncated when CC* fell below 0.543; rotation datasets employed CC1/2 > 0.5 as a cutoff. Following preliminary refinement, the resolution of 7PZQ was manually cut by an additional ~0.1 Å, which improved the resulting refinement statistics and maps. All surface area calculations were performed with PISA44 v1.48.
Structure determination
Structures were determined by iterative rounds of model building in Coot45 v0.9 and refinement with phenix.refine46 v1.20.1, after molecular replacement with search models 6YNQ (for 7PXZ & 7Z3U), 7AKU (for 7PZQ) and 7AR5 (for 7Z2K). Disulfide, NOS, and SONOS bonds were enforced using bonding restraints generated by phenix. The rotameric states of the catalytic His41 were chosen based analyzing refined B factors of the sidechain in two possible conformations (supplementary Fig. S4, supplementary Table S2). All structures were refined with riding hydrogens.
Analytical SEC
Mpro was prepared in 20 mM Tris (pH 7.8) buffer supplemented with 150 mM NaCl and 1 mM EDTA, and for the indicated sample, 1 mM TCEP. The peroxide sample was produced by incubating protein with 1 mM hydrogen peroxide for 5 h prior to injection. Subsequently, protein solutions were spun down at 16,000 g for 5 min and applied to a Cytiva Superdex 75 10/300 increase column using a ÄKTA Pure system from Cytiva. Two peaks are observed in the resulting chromatograms at elution volumes consistent with dimer and monomer species. Concentrations were estimated by peak height at the position of peak maxima, and these were used to fit a two-state dimer dissociation model by least-squares regression. Raw data and processing scripts available upon request.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data that support this study are available from the corresponding author upon request. Structural models, structure factor data, and associated metadata are available from the Protein Data Bank under PDB accession codes: 7PXZ (monoclinic XFEL reduced), 7PZQ (orthorhombic XFEL disulfide), 7Z2K (orthorhombic synchrotron reduced) & 7Z3U (orthorhombic synchrotron NOS/SONOS). Raw diffraction data are available from EuXFEL [https://doi.org/10.22003/XFEL.EU-DATA-002696-00]. Analytical SEC data and processing code are available via Zenodo [https://doi.org/10.5281/zenodo.10616060]. The source data underlying Fig. 4 is provided as a Source Data file. Source data are provided with this paper.
Code availability
The versions of Cheetah and CrystFEL used in this work are available from the respective websites: [https://www.desy.de/~barty/cheetah] and [https://www.desy.de/~twhite/crystfel].
References
Hu, Q. et al. The SARS-CoV-2 main protease (Mpro): structure, function, and emerging therapies for COVID-19. MedComm 3, e151 (2022).
Zhang, L. et al. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved a-ketoamide inhibitors. Science 368, 409–412 (2020).
Dai, W. et al. Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease. Science 368, 1331–1335 (2020).
U.S. Food and Drug Administration (FDA). Coronavirus (COVID-19) update: FDA authorizes first oral antiviral for treatment of COVID-19 (Food and Drug Administration, 2021).
Ding, L., Zhang, X. X., Wei, P., Fan, K. & Lai, L. The interaction between severe acute respiratory syndrome coronavirus 3C-like proteinase and a dimeric inhibitor by capillary electrophoresis. Anal. Biochem. 343, 159–65 (2005).
Anand, K. et al. Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra α-helical domain. EMBO J. 21, 3213–3224 (2002).
Zhong, N. et al. C-terminal domain of SARS-CoV main protease can form a 3D domain-swapped dimer. Protein Sci. 18, 839–44 (2009).
Miseta, A. & Csutora, P. Relationship between the occurrence of cysteine in proteins and the complexity of organisms. Mol. Biol. Evol. 17, 1232–1239 (2000).
Funk, L.-M. et al. Multiple redox switches of the SARS-CoV-2 main protease in vitro provide opportunities for drug design. Nat. Commun. 15, 411 (2024).
Davis, D. A. et al. Regulation of the dimerization and activity of SARS-CoV-2 main protease through reversible glutathionylation of cysteine 300. mBio 12, 1–21 (2021).
Kneller, D. W. et al. Room-temperature X-ray crystallography reveals the oxidation and reactivity of cysteine residues in SARS-CoV-2 3CL Mpro: insights into enzyme mechanism and drug design. IUCrJ 7, 1028–1035 (2020).
Rabe von Pappenheim, F. et al. Widespread occurrence of covalent lysine–cysteine redox switches in proteins. Nat. Chem. Biol. 18, 368–375 (2022).
Yang, K. S. et al. A novel Y-shaped, S-O-N-O-S-bridged cross-link between three residues C22, C44, and K61 is frequently observed in the SARS-CoV-2 main protease. ACS Chem. Biol. 18, 449–455 (2022).
Tran, N. et al. The H163A mutation unravels an oxidized conformation of the SARS-CoV-2 main protease. Nat. Commun. 14, 5625 (2023).
Schwarz, K. B. Oxidative stress during viral infection: a review. Free Radic. Biol. Med. 21, 641–649 (1996).
Davis, D. A. et al. Regulation of HIV-1 protease activity through cysteine modification. Biochemistry 35, 2482–2488 (1996).
Davis, D. A. et al. Reversible oxidative modification as a mechanism for regulating retroviral protease dimerization and activation. J. Virol. 77, 3319–3325 (2003).
Daniels, S. I. et al. The initial step in human immunodeficiency virus type 1 GagProPol processing can be regulated by reversible oxidation. PLoS ONE 5, 1–9 (2010).
Cecchini, R. & Cecchini, A. L. SARS-CoV-2 infection pathogenesis is related to oxidative stress as a response to aggression. Med. Hypotheses 143, 110102 (2020).
Reinke, P. Y. A. et al. Calpeptin is a potent cathepsin inhibitor and drug candidate for SARS-CoV-2 infections. Commun. Biol. 6, 1–13 (2023).
Barrila, J., Gabelli, S. B., Bacha, U., Amzel, L. M. & Freire, E. Mutation of Asn28 disrupts the dimerization and enzymatic activity of SARS 3CLpro. Biochemistry 49, 4308–4317 (2010).
Silvestrini, L. et al. The dimer-monomer equilibrium of SARS-CoV-2 main protease is affected by small molecule inhibitors. Sci. Rep. 11, 1–16 (2021).
Caleman, C., Junior, F. J., Grånäs, O. & Martin, A. V. A perspective on molecular structure and bonding-breaking in radiation damage in serial femtosecond crystallography. Crystals 10, 1–16 (2020).
Chapman, H. N., Caleman, C. & Timneanu, N. Diffraction before destruction. Philos. Trans. R. Soc. B Biol. Sci. 369, 1–13 (2014).
Nass, K. Radiation damage in protein crystallography at X-ray free-electron lasers. Acta Crystallogr. D Struct. Biol. 75, 211–218 (2019).
Ulrich, K. & Jakob, U. The role of thiols in antioxidant systems. Free Radic. Biol. Med. https://doi.org/10.1016/j.freeradbiomed.2019.05.035 (2019).
Han, H. et al. The XBI BioLab for life science experiments at the European XFEL. J. Synchrotron Radiat. 54, 7–21 (2021).
Günther, S. et al. X-ray screening identifies active site and allosteric inhibitors of SARS-CoV-2 main protease. Science 372, 642–646 (2021).
Mancuso, A. P. et al. The single particles, clusters and biomolecules and serial femtosecond crystallography instrument of the European XFEL: initial installation. J. Synchrotron Radiat. 26, 660–676 (2019).
Henrich, B. et al. The adaptive gain integrating pixel detector AGIPD a detector for the European XFEL. Nucl. Instrum. Methods Phys. Res. A 633, 1–4 (2011).
Fangohr, H. et al. Data analysis support in Karabo at European XFEL. In 16th International Conference on Accelerator and Large Experimental Control Systems 245–252 (Joint Accelerator Conferences Website, 2018).
Oberthuer, D. et al. Double-flow focused liquid injector for efficient serial femtosecond crystallography. Sci. Rep. 7, 1–12 (2017). 2017 7:1.
Knoška, J. et al. Ultracompact 3D microfluidics for time-resolved structural biology. Nat. Commun. 11, 657 (2020).
Vakili, M. et al. 3D printed devices and infrastructure for liquid sample delivery at the European XFEL. J. Synchrotron Radiat. 29, 331–346 (2022).
Burkhardt, A. et al. Status of the crystallography beamlines at PETRA III. Eur. Phys. J. Plus 131, 1–9 (2016).
Mariani, V. et al. OnDA: online data analysis and feedback for serial X-ray imaging. J. Appl. Crystallogr. 49, 1073–1080 (2016).
Yefanov, O. et al. Accurate determination of segmented X-ray detector geometry. Opt. Express 23, 28459–28470 (2015).
Barty, A. et al. Cheetah: software for high-throughput reduction and analysis of serial femtosecond X-ray diffraction data. J. Appl. Crystallogr. 47, 1118–1131 (2014).
White, T. A. et al. CrystFEL: a software suite for snapshot serial crystallography. J. Appl. Crystallogr. 45, 335–341 (2012).
Leslie, A. G. W. & Powell, H. R. Processing diffraction data with mosflm. Evol. Methods Macromol. Crystallogr. 245, 41–51 (2007).
Gevorkov, Y. et al. XGANDALF - extended gradient descent algorithm for lattice finding. Acta Crystallogr. A Found. Adv. 75, 694–704 (2019).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Juers, D. H. & Ruffin, J. MAP-CHANNELS: a computation tool to aid in the visualization and characterization of solvent channels in macromolecular crystals. J. Appl. Crystallogr. 47, 2105–2108 (2014).
Delano, W. L. The PyMOL molecular graphics system. CCP4 Newslett. 40, 1–9 (2002).
Acknowledgements
We acknowledge T. White for assistance during the XFEL experiment and C. Uetrecht for valuable discussions. T.J.L. was supported by a Helmholtz Young Investigator Award. H.B., C.B., S.B., H.N.C., T.J.L. acknowledge financial support obtained from the Cluster of Excellence ‘Advanced Imaging of Matter’ of the Deutsche Forschungsgemeinschaft (DFG) - EXC 2056 - project ID 390715994. In addition, H.B. and C.B. received support from BMBF via projects 05K19GU4 and 05K20GUB. The Helmholtz association funded S.F., J.L. and A.M. through the project FISCOV and P.Y.A.R. and A.M. through FragX. The Helmholtz association further supported the project by financing P.Y.A.R, J.L. and A.M. via the Impulse and Networking funds InternLabs-0011 ‘HIR3X’. W.E. was supported through the BMBF-funded project “ConScience” (project 16GW0277) and the Röntgen-Angstrom cluster project “X-ray drug design platform” (13K22CHB). D.T. was supported by the Slovenian Research Agency (ARRS; research program P1-0048, Infrastructural program IO-0048). L.R. was financed by BMBF project 13K18FLA. We acknowledge European XFEL in Schenefeld, Germany, for provision of x-ray free-electron laser beamtime at SPB/SFX as part of proposal 2696 and would like to thank the staff for their assistance. The authors are indebted to the XFEL Biology Infrastructure (XBI) for the provision of instrumentation that has enabled this experiment. Sample reservoirs and the anti-settling device employed in parts of the measurements presented here were designed and fabricated by the Max Planck Institute for Medical Research, Heidelberg, which also provided instruction in its use. This research was supported through computational resources (Maxwell cluster) and experimental facilities (PETRA-III beamline P11) operated by Deutsches Elektronen-Synchrotron DESY, Hamburg, Germany, a member of the Helmholtz Association HGF. We acknowledge the P11 staff for their invaluable help. All authors are indebted to Galen Correy, as well as an anonymous reviewer, for constructive criticism during the review process.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Crystallization was performed by P.Y.A.R. and R.S. Size exclusion chromatography experiments were conducted by P.Y.A.R. Crystallographic refinement was performed by P.Y.A.R., R.S., D.O., D.T. and W.H. Authors M.G., A.R.M., S.G., A.C., A.R., B.C.S., B.N., C.K., C.S., F.H.M.K., A.T., W.E., G.E.P., G.M., H.K., H.B., H.H., J.Koliyadu, J.B., J.L., J.M., J.Knoška, K.L., L.B., M.S., M.K., M.V., P.V., P.M., R.d.W., R.B., R.L., S.H., S.F., T.G., T.S., V.S., Y.K., and O.M.Y contributed to the technical conduct of XFEL-based serial crystallography experiments. L.G., J.S., T.B., A.S.D., A.P.M., C.B., S.B., L.R., H.N.C., A.M., D.T., W.H. and T.J.L. conceived of the work, provided management oversight and technical comments on the manuscript. The manuscript, including figures and relevant data analysis, was drafted, and then revised, by P.Y.A.R., R.S., D.O., D.T., W.H. and T.J.L. The project was coordinated by T.J.L.
Corresponding author
Ethics declarations
Competing interests
T.J.L. and A.S.D. are employees and shareholders of CHARM Therapeutics. T.G. is an employee of Sosei Heptares and A.S.D. was an employee of Sosei Heptares at the time the work was conducted. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Galen Correy and the other, anonymous, reviewer for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Reinke, P.Y.A., Schubert, R., Oberthür, D. et al. SARS-CoV-2 Mpro responds to oxidation by forming disulfide and NOS/SONOS bonds. Nat Commun 15, 3827 (2024). https://doi.org/10.1038/s41467-024-48109-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-48109-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.