Abstract
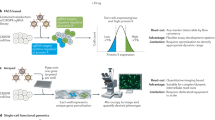
High-throughput genomics, transcriptomics, proteomics and metabolomics have the potential to identify the functional consequences of induced and natural genetic variation. Surprisingly, the experiments of most genomics researchers still mainly involve perturbing a biological system of interest by modifying either one factor or one gene at a time. By contrast, this article argues that multifactorial experimentation would allow the study of many more biologically relevant questions in parallel at the same or lower cost.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Chong, L. & Ray, L. B. Whole-istic biology. Science 295, 1661 (2002).
Kitano, H. Systems biology. Science 295, 1662–1668 (2002).
Knowlton, R. G. et al. A polymorphic DNA marker linked to cystic fibrosis is located on chromosome 7. Nature 318, 380–385 (1985).
Kerem, B. -S. et al. Identification of the cystic fibrosis gene: genetic analysis. Science 245, 1073–1080 (1989).
Gilliam, T. C. et al. Localization of the Huntington's disease gene to a small segment of chromosome 4 flanked by D4S10 and the telomere. Cell 50, 565–571 (1987).
Huntington's Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable in Huntington's disease chromosomes. Cell 72, 971–983 (1993).
Hughes, T. R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000).
Ideker, T. et al. Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292, 929–934 (2001).
Davidson, E. H. et al. A genomic regulatory network for development. Science 295, 1669–1678 (2002).
Ernest, S. et al. Genetic and molecular control of folate-homocysteine metabolism in mutant mice. Mamm. Genome 13, 259–267 (2002).
Jansen, R. C. & Nap, J. P. H. Genetical genomics: the added value from segregation. Trends Genet. 17, 388–391 (2001).
Brem, R. B., Yvert, G., Clinton, R. & Kruglyak, L. Genetic dissection of transcriptional regulation in budding yeast. Science 296, 752–756 (2002).
Klose, J. et al. Genetic analysis of the mouse brain proteome. Nature Genet. 30, 385–393 (2002).
Wayne, M. L. & McIntyre, L. M. Combining mapping and arraying: an approach to candidate gene identification. Proc. Natl Acad. Sci. USA 99, 14903–14906 (2002).
Demant, P. & Hart, A. A. M. Recombinant congenic strains — a new tool for analyzing genetic traits determined by more than one gene. Immunogenetics 24, 416–422 (1986).
Fijneman, R. J. A., Ophoff, R. A., Hart, A. A. M. & Demant, P. Kras-2 alleles, mutations, and lung tumor susceptibility in the mouse — an evaluation. Oncogene 9, 1417–1421 (1994).
Darvasi, A. Experimental strategies for the genetic dissection of complex traits in animal models. Nature Genet. 18, 19–24 (1998).
Nadeau, J. H., Singer, J. B., Matin, A. & Lander, E. S. Analysing complex genetic traits with chromosome substitution strains. Nature Genet. 24, 221–225 (2000).
Threadgill, D. W., Hunter, K. W. & Williams, R. W. Genetic dissection of complex and quantitative traits: from fantasy to reality via a community effort. Mamm. Genome 13, 175–178 (2002).
Jannink, J. L. & Jansen, R. C. Mapping epistatic quantitative trait loci with one-dimensional genome searches. Genetics 6, 337–342 (2001).
Gygi, S. P., Rochon, Y., Franza, B. R. & Aebersold, R. Correlation between protein and mRNA abundance in yeast. Mol. Cell. Biol. 19, 1720–1730 (1999).
Jansen, R. C., Nap, J. P. H. & Mlynarova, L. Errors in genomics and proteomics. Nature Biotechnol. 20, 19 (2002).
Mather, K. & Jinks, J. L. Biometrical Genetics 3rd edn (Chapman & Hall, London, 1982).
Brenner, S. et al. Gene expression analysis by massively parallel signature sequencing (MPSS) on microbead arrays. Nature Biotechnol. 18, 630–634 (2000).
Nap, J. P., Conner, A. J., Mlynarova, M., Stiekema, W. J. & Jansen, R. C. Dissection of a synthesized quantitative trait to characterize transgene interactions. Genetics 147, 315–320 (1997).
Mlynarova, L., Loonen, A., Mietkiewska, E., Jansen, R. C. & Nap, J. P. Assembly of two transgenes in an artificial chromatin domain gives highly coordinated expression in tobacco. Genetics 160, 727–740 (2002).
Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002).
Claverie, J. M. Gene number — what if there are only 30,000 human genes? Science 291, 1255–1257 (2001).
Carlborg, O., Andersson, L. & Kinghorn, B. The use of genetic algorithm for simultaneous mapping of multiple interacting quantitative trait loci. Genetics 155, 2003–2010 (2000).
Broman, K. W. & Speed, T. A model selection approach for the identification of quantitative trait loci in experimental crosses. J. R. Statist. Soc. B 64, 1–6 (2002).
Ritchie, M. D. et al. Multifactorial-dimensionality reduction reveals high-order interactions among estrogen-metabolism genes in sporadic breast cancer. Am. J. Hum. Genet. 69, 138–147 (2001).
Nelson, M. R., Kardia, S. L. R., Ferrell, R. E. & Sing, C. F. A combinatorial partitioning method to identify multilocus genotypic partitions that predict quantitative trait variation. Genome Res. 11, 458–470 (2001).
Fijneman, R. J. A., de Vries, S. S., Jansen, R. C. & Demant, P. Complex interaction of new quantitative trait loci, Sluc1, Sluc2, Sluc3, and Sluc4, that influence the susceptibility to lung cancer in the mouse. Nature Genet. 14, 465–467 (1996).
de Boer, M. P., ter Braak, C. J. F. & Jansen, R. C. A penalized likelihood method for mapping epistatic quantitative trait loci with one-dimensional genome searches. Genetics 162, 951–960 (2002).
Wagner, A. How to construct a large genetic network from n gene perturbations in fewer than n2 easy steps. Bioinformatics 17, 1183–1197 (2001).
de la Fuente, A., Brazhnik, P. & Mendez, P. Linking the genes: inferring quantitative gene networks from microarray data. Trends Genet. 18, 395–398 (2002).
Stoll, M. et al. A genomic-systems biology map for cardiovascular function. Science 294, 1723–1726 (2001).
Kerr, M. K. & Churchill, G. A. Statistical design and the analysis of gene expression microarray data. Genet. Res. 77, 123–128 (2001).
Fisher, R. A. The Design of Experiments (Oxford Univ. Press, Oxford, UK, 1935).
Kitami, T. & Nadeau, J. H. Biochemical networking contributes more to genetic buffering in human and mouse metabolic networks than does gene duplication. Nature Genet. 32, 191–194 (2002).
Yuh, C. H., Bolouri, H. & Davidson, E. H. Genomic cis-regulatory logic: experimental and computational analysis of a sea urchin gene. Science 279, 1896–1902 (1998).
Fessele, S., Maier, H., Zischek, C., Nelson, P. J. & Werner, T. Regulatory context is a crucial part of gene function. Trends Genet. 18, 60–63 (2002).
Frankel, W. N. & Schork, N. J. Who is afraid of epistasis? Nature Genet. 14, 371–373 (1996).
Templeton, A. R. in Epistasis and the Evolutionary Process (ed. Wolf, J.B.) 41–57 (Oxford Univ. Press, Oxford, UK, 2000).
Acknowledgements
This article is dedicated to my former room-mate and bioinformatics colleague J. (Hans) M. Sandbrink, who recently passed away. I am grateful to M. P. de Boer for carrying out the simulations presented in Box 1, to R. W. Williams for providing early access to his mouse work, and to three reviewers for their constructive comments.
Author information
Authors and Affiliations
Related links
Related links
Databases
OMIM
Saccharomyces Genome Database
FURTHER INFORMATION
Glossary
- ADVANCED INTERCROSS LINES
-
Subsequent generations (F3, F4, and so on) of an intercross pedigree that are used for the high-resolution mapping of trait loci.
- CHROMOSOME SUBSTITUTION STRAINS
-
(CSS). Each CSS contains an entire chromosome of a donor parent placed in the genetic background of the recipient parent.
- COMBINATORIAL PARTITIONING
-
A computational strategy that consists of pooling genotypes from multiple loci into a smaller number of classes, thereby avoiding the increased dimensionality that is associated with modelling interactions between loci or between loci and the environment.
- COMPLEX TRAITS BIOLOGY
-
The study of traits that are determined by many genes, which almost always interact with environmental factors.
- EPISTASIS
-
In the context of quantitative genetics, epistasis refers to any genetic interaction in which the combined phenotypic effect of two or more loci is less than (negative epistasis) or greater than (positive epistasis) the sum of the effects at individual loci.
- GENETIC ALGORITHM
-
A numerical optimization procedure that is based on evolutionary principles such as mutation, deletion and selection.
- GENETICAL GENOMICS
-
The process that uses gene expression profiling and marker-based fingerprinting of each individual in a segregating population to analyse the cis- and trans-acting factors that underlie variation in gene expression. This information can then be used to reconstruct a gene network.
- MARKOV CHAIN MONTE CARLO STRATEGIES
-
A randomized computational approach for identifying the most likely among many possible models.
- MULTICOLLINEARITY
-
The situation in which two or more predictors (or subsets of predictors) are strongly (but not perfectly) correlated to one other, making it difficult to interpret the strength of the effect of each predictor (or predictor subset). For example, it would be hard to detect a gene if its effect is 'absorbed' (or masked) by combinations of genetic background action/interaction parameters in the model.
- QUANTITATIVE TRAIT LOCI
-
(QTL). Genetic loci or chromosomal regions that contribute to variability in complex quantitative traits (such as plant height or body weight), as identified by statistical analysis. Quantitative traits are typically affected by several genes and by the environment.
- RECOMBINANT CONGENIC STRAIN
-
(RCS). A population of fully homozygous individuals, each of which contains a restricted part of one of the two genomes from which the inbred lines were created.
- RECOMBINANT INBRED LINES
-
(RILs). A population of fully homozygous individuals that is obtained through the repeated selfing of an F1 hybrid, and that comprises 50% of each parental genome in different combinations.
- SYSTEMS BIOLOGY
-
The study of the complex interactions that occur at all levels of biological information — from whole-genome sequence interactions to developmental and biochemical networks — and their functional relationship to organism-level phenotypes.
- VARIANCE
-
A statistic that quantifies the dispersion of data about the mean.
Rights and permissions
About this article
Cite this article
Jansen, R. Studying complex biological systems using multifactorial perturbation. Nat Rev Genet 4, 145–151 (2003). https://doi.org/10.1038/nrg996
Issue Date:
DOI: https://doi.org/10.1038/nrg996
This article is cited by
-
Inferring cell-type-specific causal gene regulatory networks during human neurogenesis
Genome Biology (2023)
-
The phosphatidylinositide 3-kinase (PI3K) signaling pathway is a determinant of zileuton response in adults with asthma
The Pharmacogenomics Journal (2018)
-
Quantitative trait loci from identification to exploitation for crop improvement
Plant Cell Reports (2017)
-
The wright stuff: reimagining path analysis reveals novel components of the sex determination hierarchy in drosophila melanogaster
BMC Systems Biology (2015)
-
Candidate gene association studies: a comprehensive guide to useful in silicotools
BMC Genetics (2013)