Abstract
More than any other technology, nucleic acid sequencing has enabled microbial ecology studies to be complemented with the data volumes necessary to capture the extent of microbial diversity and dynamics in a wide range of environments. In order to truly understand and predict environmental processes, however, the distinction between active, inactive and dead microbial cells is critical. Also, experimental designs need to be sensitive toward varying population complexity and activity, and temporal as well as spatial scales of process rates. There are a number of approaches, including single-cell techniques, which were designed to study in situ microbial activity and that have been successively coupled to nucleic acid sequencing. The exciting new discoveries regarding in situ microbial activity provide evidence that future microbial ecology studies will indispensably rely on techniques that specifically capture members of the microbiome active in the environment. Herein, we review those currently used activity-based approaches that can be directly linked to shotgun nucleic acid sequencing, evaluate their relevance to ecology studies, and discuss future directions.
Similar content being viewed by others
Introduction
Microbial communities perform essential roles in natural and engineered ecosystems. They contribute to the dynamics and complexity of global biogeochemical cycles and are the main drivers for important biotechnological processes. One of the main challenges in evaluating the microbial impact in a given system is that organism abundance, diversity and (re-)action potential can vary across systems and across different temporal and spatial scales and may be correlated with a combination of biological, chemical and physical stimuli. The identification of important microbial factors in a given habitat using 16S ribosomal RNA (rRNA) gene surveys (Tringe and Hugenholtz, 2008), shotgun metagenomics (Handelsman, 2004) and single-cell genomics (Lasken, 2007) has provided important insights into the potential functions of particular microbes. However, using these techniques in their traditional sense does not offer the possibility to fully capture the plasticity of microbial communities and its effect on local and global processes. Instead, it is becoming essential to complement sequencing with methods that distinguish cells that are growing and metabolically active from those that are dormant or dead (or even extracellular DNA), and to understand under what circumstances microbial dormancy could be followed by stochastic awakening (Buerger et al., 2012). Meaningful ecological units, such as microbial guilds (Simberloff and Dayan, 1991), could be identified by monitoring the activity of individual microbes in relevant time series experiments coupled with spatially comprehensive study designs. Several of the discussed tools offer single-cell resolution and thus provide spatial resolution in addition to information on phylogeny and metabolism (Table 1, Figure 1). This review is aimed at highlighting the different ecological and physiological insights into active community members of a given microbiome that can be gained with currently used approaches (in chronological order of published studies) in conjunction with shotgun nucleic acid sequencing. We note that methods that yield activity-labeled samples in situ cannot provide answers to all ecological challenges, however, when applicable, these tools present valuable assets in microbial ecology research. We further discuss prospects of exciting emerging technologies in this field.
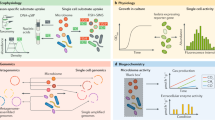
Methods that yield activity-labeled samples and are targeting cell processes in an ‘active’ microbial cell that can be coupled with shotgun sequencing. Colors denote resources (green), cell components (blue) and cell processes (orange). Raman: Raman microspectroscopy. For DNA- and RNA-SIP, total nucleic acids are extracted from the samples, and labeled and unlabeled DNA/RNA is separated by density gradient centrifugation. The ‘heavier’ labeled nucleic acid fractions can be used for construction of metagenomic libraries (Neufeld et al., 2007; Whiteley et al., 2007), whereas PLFAs are analyzed on a mass spectrometer and cannot be combined with nucleic acid sequencing (Jehmlich et al., 2010). Many 13C-, 18O-, 15N-labeled fine chemicals are available (for example, phenol, methanol, ammonia, methane, carbonate, etc.), but the wide-ranging application of SIP is limited by the commercial availability of complex labeled compounds that require expensive custom synthesis. Furthermore, sensitivity of the SIP technique is a function of substrate concentration and the duration of substrate incorporation. Successful SIP is dependent on optimization of substrate concentration to guarantee a significant signal-to-noise ratio and incubation length and avoid enrichment bias (Neufeld et al., 2006) (Table 1). ‘Cross-feeding’, that is, the flow of the isotope label from primary metabolizers to secondary consumers has also been documented (Hutchens et al., 2004; Dumont et al., 2006). RSG is a fluorogenic redox indicator dye available from Molecular Probes, Invitrogen (Carlsbad, CA, USA). RSG yields green fluorescence (488 nm excitation) when modified by bacterial reductases, many of which are parts of electron transport systems. SIP-Raman microspectroscopy has been performed using 13C-, 15N-labeled compounds, as well as with D2O. The addition of D2O (up to a certain concentration and for limited time) is expected to have negligible effects on the microbial community composition and activity patterns, for example, compared with nutrient substrates (Lester et al., 1960; Berry et al., 2015; Kopf et al., 2015) that are traditionally used for SIP experiments. Incorporation of D2O-derived deuterium into the biomass of autotrophic and heterotrophic bacteria and archaea can be unambiguously detected via C-D signature peaks in single-cell Raman spectra (Ashkin, 1970; Berry et al., 2015). However, for comparative studies between active taxa it should be kept in mind that microbes with different physiologies will incorporate different amounts of deuterium at similar growth rates.
An array of methods available for studying microbial in situ activity linked to shotgun sequencing
Methods detecting microbial in situ activity target a variety of physiological processes to determine the activity state of a cell. The earliest developed microbial method that yields activity-labeled samples in situ and can be combined with shotgun sequencing takes advantage of the incorporation of bromodeoxyuridine (BrdU) into newly synthesized DNA (and RNA) of a viable cell and subsequent density gradient centrifugation or immunocapture of BrdU-labeled DNA (Urbach et al., 1999). BrdU labeling has been successfully used in a variety of environments to identify DNA-synthesizing populations. Studies have investigated phosphate stimulation in soil (Borneman, 1999), the active population of lake water (Urbach et al., 1999), bacterial functional redundancy with respect to various carbon substrates in soil (Yin et al., 2000), microbial association with arbuscular mycorrhizal hyphae (Artursson and Jansson, 2003), microbial response to inoculation of various plants with arbuscular mycorrhizal (Artursson et al., 2005), nitrogen addition in Alaskan boreal forest (Handelsman, 2004; Allison et al., 2008), vertical redox gradients in Baltic Sea sediment (Lasken, 2007; Edlund et al., 2008), fungal response to different carbon sources in forest soils (Hanson et al., 2008; Buerger et al., 2012; McGuire et al., 2012) and microbial response to carbon sources of varying recalcitrance in soil (Urbach et al., 1999; Goldfarb et al., 2011). In addition, studies have targeted bacterial and archaeal (Borneman, 1999; Yin et al., 2000; Artursson and Jansson, 2003; Artursson et al., 2005; Goldfarb et al., 2011) as well as fungal (Urbach et al., 1999; Allison and Treseder, 2008; Allison et al., 2008; Hanson et al., 2008) DNA synthesis activity in response to various environmental stimuli in soils and sediments (Yin et al., 2000; Edlund et al., 2008). Although BrdU-labeling experiments can help identify replicating microorganisms and determine average activity in response to various changing environmental conditions, organisms vary in their ability to incorporate the relatively large BrdU molecule into their DNA, and the rate at which incorporation occurs can deviate up to 10-fold between cells (Urbach et al., 1999; Artursson and Jansson, 2003; Hellman et al., 2011). Such variability may lead to incomplete and skewed representation of the active microbial population in a given environment. In addition, owing to low labeling efficiency, BrdU labeling linked to sequencing requires a relatively large number of sample replicates to obtain the amount of DNA needed for 16S rRNA gene and metagenome sequencing, which renders this technique relatively costly and labor intensive (Tables 1 and 2). The aforementioned studies have yielded important insights into the functioning of microbial communities, yet combining BrdU labeling with either fluorescence-activated cell sorting (FACS) and subsequent high-throughput sequencing or solely with high-throughput sequencing has found limited application to date (Mou et al., 2008; David et al., 2014; Hamasaki et al., 2016). Published studies have predominantly included PCR-amplified rRNA gene surveys and only a few metagenome studies are available (Figure 2, Table 3). However, as DNA requirements for shotgun sequencing decrease, sequencing of BrdU-labeled environmental DNA may become more prevalent.
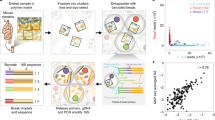
Project statistics by method over the last decade. Counts displayed exclusively include projects using high-throughput sequencing. Metagenome (MetaG) and metatranscriptome (MetaT) projects are depicted by lines (primary y axis), whereas BrdU, DNA-SIP, RNA-SIP and PTR projects are displayed as bars (secondary y axis). Project abundances are cumulative. Number of MetaG and MetaT projects include public sequencing projects as recorded in the Genomes OnLine Database (GOLD) (Pagani et al., 2011) retrieved 15 January 2016. Number of all other activity-based projects include published records that feature high-throughput (next-generation) shotgun sequence data.
Stable isotope probing (SIP) labeling technology can be used to selectively enrich the DNA or RNA of active microbial populations (Radajewski et al., 2003). For SIP analysis, stable isotope (for example, 2H, 13C, 15N or 18O) labeled substrates are added to the environment and as these substrates are consumed by active organisms, labeled atoms are incorporated in their DNA or RNA. Radajewski et al. (2000) first demonstrated this approach by adding various 13C-enriched carbon sources (13CH3OH, 13CH4) to soil and subsequently identifying active methylotrophs. The authors found that methylotrophy was confined to Alphaproteobacteria lineages and identified various new methanol dehydrogenase alpha subunit (mxaF) sequences. DNA-SIP is currently the most applied method, having been used by many research groups to study in situ microbial activity (Figure 2). RNA-SIP provided insights that led to the isolation and subsequent genome sequencing of a Thauera sp., which was identified as key phenol degrading organisms in an industrial wastewater treatment plant (Manefield et al., 2002). Recently, Dumont et al. (2013) obtained a 13CH4 labeled-RNA metatranscriptome from methanotrophs in lake sediment. The labeled metatranscriptome was predominantly enriched in sequences from Methylococcaceae and transcripts of methane monooxygenase (pmoCAB) genes. The authors suggested that SIP and metatranscriptomics can be broadly applicable, but this approach has only been used one other time to our knowledge (Fortunato and Huber, 2015). Although limitations of DNA- and RNA-SIP include availability of labeled compounds, cross-feeding, sensitivity and potential for enrichment bias (Figure 1, Table 1), continuous efforts to optimize SIP, applying it to wider ranges of environments, and combining it with other techniques, such as Raman microspectroscopy and fluorescence in situ hybridization (FISH; Huang et al., 2007) make DNA-SIP the best current method for sequence-based characterization of key populations synthesizing DNA in the presence of isotope-labeled compounds.
Metatranscriptome studies enable the analysis of expressed genes in environmental communities, portraying the near real-time condition of microorganisms because of the relatively short half-life of mRNA (Moran et al., 2012). Studies of microbial communities in soil and marine environments were the first to use high-throughput sequencing technology for mRNA analysis (Leininger et al., 2006; Frias-Lopez et al., 2008; Urich et al., 2008). Since then, several studies in various environments have shown that only a subset of the total microbial community contributes to the sequenced RNA pool, and can therefore be considered transcriptionally active. For example, Lesniewski et al. (2012) were able show that transcriptional microbial activity of hydrothermal plume communities was largely attributed to water column microorganisms rather than members from the seafloor. mRNA is regularly used as an indicator of microbial activity and is logistically advantageous because samples can be frozen instantaneously, thereby enabling delayed processing after freezing, which is not an option for other methods that yield activity-labeled samples in situ (Table 1). This is relevant when in situ conditions cannot easily be maintained during sampling or downstream sample processing. A critical issue is how deeply a community transcriptome is ‘covered’ by the sequence data. If too shallow, libraries will be dominated by transcripts from metabolic pathways shared by most cells and lacking in those representing specialized biogeochemical pathways (Hewson et al., 2009; Gifford et al., 2010; Poretsky et al., 2010). As a consequence, unique expression patterns within a community may be missed, and comparative analyses between communities can become insensitive. Also, depletion of rRNA before metatranscriptome sequencing is not always trivial. Nevertheless, the near-real-time response of mRNA levels to environmental changes experienced by a cell renders assessing inventories of mRNA pools highly informative for identifying ecologically relevant processes while also determining the identities of their microbial drivers. Declining costs, greater sequencing depth and more comprehensive genomic databases render metatranscriptomics an increasingly powerful and popular tool.
With the advent of fluorescence microscopy activity stains have found an application in microbial ecology. These include stains that rely on cell membrane integrity, enzyme activity or the presence of biomolecules (lipids/nucleic acid). For instance, redox sensor green (RSG) and denaturing gradient gel electrophoresis (DGGE) profiling were used to detect and enumerate microbes actively metabolizing the C1 substrates methane, methanol, methylamine and formaldehyde in Lake Washington sediment (Kalyuzhnaya et al., 2008). According to Kalyuzhnaya et al., RSG does not suppress cellular metabolism as was previously found for tetrazolium salts, such as CTC (5-cyano-2,3-ditolyl tetrazolium chloride) (Ullrich et al., 1996). Using FACS, RSG could likely be scaled to separate bacterial and archaeal cells with an electron membrane potential in the presence of environmental stimuli, therefore studying population activity levels.
One of the most recently developed methods that yield activity-labeled samples in situ is bioorthogonal non-canonical amino-acid tagging (BONCAT). This technique is based on the in vivo incorporation of artificial amino acids that carry modifiable chemical tags into newly synthesized proteins (Dieterich et al., 2006). These chemical tags can then be fluorescently labeled via click chemistry (Hatzenpichler et al., 2014). BONCAT has been demonstrated to be effective in labeling the proteomes of a wide range of taxonomically and physiologically distinct archaea and bacteria (Hatzenpichler et al., 2014). For example, dominant cells in an aerobic methanotrophic enrichment culture from deep-sea sediments were identified as Methylococcaceae-related gammaproteobacterium based on FISH analyses. The translational activity of these Methylococcaceae cells was found to be dependent on the addition of methane to the enrichment culture. Genome information from this Methylococcaceae strain confirmed the methanotrophic nature of this bacterium. Rather than studying the bulk proteome, BONCAT is able to specifically target proteins that have been expressed in response to an experimental condition. The method can also be used in conjunction with rRNA-targeted FISH and thereby allow the simultaneous identification of a microbial cell and its translational activity. In a very recent effort to describe anaerobic methane oxidizers from methane seep sediments, BONCAT was combined with FACS to sort translationally active methane-oxidizing consortia. Whole-genome amplification and 16S rRNA gene sequencing of the aggregates revealed a novel archaeal–bacterial association (Hatzenpichler et al., 2016). This technique presents great prospects for future high-throughput sequencing coupled to BONCAT and is expected to help answer questions regarding cellular translational activities in response to environmental cues at micrometer resolution, as well as in a high-throughput manner across larger scales (Figure 3). Limitations of BONCAT include the diversity of amino-acid uptake mechanisms in the environment, the potential for artificial growth stimulation following addition of amino acids and suppression of signal when environmental concentrations of amino acids are high. Aside from these potential limitations, we expect BONCAT to be combined with shotgun metagenomics in the near future.
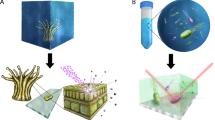
High-throughput workflows of current and emerging in situ microbial activity approaches linked to sequencing. Metatranscriptomics and stable isotope labeling are the most commonly used techniques coupled with next-generation shotgun sequencing technology. Emerging methods that are currently still subject to development and/or optimization involve the incubation of cells and cell clusters with, for example, fluorescent compounds or D2O before selective sorting of active cells using FACS or Raman OT.
Raman microspectroscopy is an established vibrational spectroscopic technique with emerging prospects in microbial ecology. This non-destructive technique enables chemical fingerprinting of individual eukaryotic cells, organelles, or bacterial and archaeal cells. Chemical information derived from a Raman spectrum provides comprehensive and intrinsic information on the chemical composition of the analyzed cell without the need of external labeling, which, for example, enables microbiologists to identify storage compounds, cytochromes and pigments on a single-cell level within a few seconds (Gruber-Vodicka et al., 2011; Majed et al., 2012; Milucka et al., 2012). Interestingly, the incorporation of the stable isotopes 13C, deuterium and 15N into microbial biomass can also be detected via characteristic band shifts in the Raman spectra of the labeled microbes (Wang et al., 2016), but it should be noted that the 15N-induced shifts are difficult to interpret in complex samples. As Raman microspectroscopy can be combined with FISH (Huang et al., 2007) and optical tweezers (OTs), it offers a culture-independent approach to study the physiology of uncultivated microorganisms in their natural ecosystem. For example, Raman-FISH was applied to identify and measure in situ 13C-naphthalene degraders within a microbial community from a complex groundwater system and proved that an uncultured species (Acidovorax sp.) had the key role in naphthalene biodegradation, rather than the cultivated naphthalene biodegrading Pseudomonas sp. obtained from the same groundwater (Huang et al., 2008). OT-Raman (Raman microspectroscopy coupled with OTs) has been applied to trap and measure single cells of blood (Xie et al., 2002) or Escherichia coli (Xie and Li, 2003) and can also be used to sort isotope-labeled microbial cells for subsequent whole genome amplification and sequence-based 16S rRNA gene identification (Huang et al., 2009; Berry et al., 2015). Alternatively, Raman-activated cell ejection using laser-induced forward transfer can be exploited as a low-throughput method for physically removing individual cells from a sample based on their Raman spectrum (Wang et al., 2013; Song et al., 2016).
In a recent study, active microbial cells were identified under near-natural conditions in complex systems via combined use of heavy water (D2O) and Raman microspectroscopy (Berry et al., 2015). D2O-Raman spectroscopy can be combined with FISH for parallel identification of the metabolically active microbes. Furthermore, unlabeled substrates can be added and changes in activities of microbial groups of interest can be quantified (Berry et al., 2015; Eichorst et al., 2015). For example, it was revealed that in the cecal microbiota of mice Akkermansia muciniphila and Bacteroides acidifaciens, two host-compound foragers, exhibited distinctive response patterns to amendments of mucin and sugars (Berry et al., 2015). By applying D2O-labeling and OT-Raman, cecal microbial cells stimulated by mucin or glucosamine were obtained for multiple displacement amplification and subsequent identification through 16S rRNA gene sequencing (Berry et al., 2015). Although offering direct access to genomic information from microbes with a defined activity, sorting via OTs is slow. In this context, it is interesting to note that recently a fast Raman-activated cell sorting system (RACS), was designed and used to collect carotenoid-producing yeast cells (Zhang et al., 2015a). Other recent advances include microfluidics-based cell manipulation techniques, such as flow-based OT-RACS and Dielectrophoresis (DEP-) RACS, and advanced Raman techniques, for example, Surface-enhanced Raman spectroscopy (SERS) flow cytometry, Resonance Raman spectroscopy (RRS) cell counting and Coherent anti-Stokes Raman spectroscopy (CARS) flow cytometry, which are discussed elsewhere (Zhang et al., 2015b). Microfluidic device-based Raman sorting can be readily integrated with downstream processes, such as cell incubation, chemical analysis and PCR (Wang et al., 2005). Unlike FACS, Raman sorting can use more criteria, including the incorporation of stable isotopes, to identify and sort individual cells (Table 1).
Finally, two very recently developed non-invasive bioinformatic approaches for linking metagenomics and bacterial activity use sequence read depth across microbial genomes. DNA replication typically begins at a single origin of replication in bacterial and archaeal chromosomes (Mott and Berger, 2007). Thus, during DNA replication, regions that have already been passed by the replication fork will have two copies while the yet unreplicated regions will have a single copy. Korem et al. (2015) show that metagenome read coverage patterns for different microbial genomes contain a single trough and a single peak, the latter coinciding with the bacterial origin of replication. The peak-to-trough ratio (PTR) was also demonstrated to provide a quantitative measure of a species growth rate in vitro and in vivo, under different growth conditions and in complex bacterial communities. For instance, it was shown that the proliferative behavior of virulent and non-virulent strains of C. rodentium could be accurately predicted by their PTRs. The application of PTRs is promising, not least, because it considers genomic variance between strains, copy number variation of different genomic regions and variable coverage resulting from sequencing depth. However, it requires information on origin of replication sites of the investigated bacteria and archaea and, on a case-by-case basis, manual correction for relative abundance levels of taxa (Table 1). In an effort to expand this method to metagenome data with draft genomes, Brown et al. (2016) developed the index of replication (iRep), which is based on mapping metagenome sequencing reads to the collection of assembled sequences that represent a draft genome. Excluding extremely high and low coverage regions, the overall slope of coverage across a genome is used to calculate iRep (Brown et al., 2016). Applied to metagenomic data, iRep showed, for instance, that Candidate Phyla Radiation bacteria are generally characterized by slower growth rates compared with various human microbiome organisms. iRep requires a ⩾75% complete genome in order to produce accurate measurements and was shown to be then correlated to PTRs and experimentally determined growth rates (Brown et al., 2016).
Choosing an appropriate method for assessing microbial activity in situ
Our view of microbial activity is strongly shaped by the method and parameters chosen. Microbial contribution to processes can neither always be definitively assessed in every environment nor at a scale that allows interpretation at the systems level. But when experiments are designed carefully to address specific ecological questions at relevant and yet tangible temporal or spatial scales and under consideration of the limitations of the chosen tool (as discussed in the following), methods that yield activity-labeled samples in situ, are a great resource. The choice of an appropriate tool for targeting the in situ active microbial population will depend on various factors. The physicochemical characteristics of the environment of interest, for example, water saturation, pH, amino acid and sulfide concentration, determine whether a water- or bioorthogonal amino-acid-based labeling method is practical. Although microbial activity is generally correlated with the availability of water in a system, the addition of water may lead to artificially increased microbial response, for example, in systems that are heterogeneously wetted, such as soil aggregates. On the other hand, adding isotope-labeled substrates other than water as typically done in conventional SIP studies changes the nutrient composition of the system and often leads to biases in microbial activity patterns (Table 1). The presence of a reductase activity, DNA synthesis, transcription and translation may occur in cell maintenance mode, as well as during growth and division, which is problematic for the correlation of these processes to quantitative rates of specific functions (Blazewicz et al., 2013). Estimates for maintenance requirements between cultures and microbes living, for example, in low activity subsurface environments vary widely (Hoehler and Jørgensen, 2013). Similarly, ranges of metabolic rates for the same reaction should be modeled for their specific environmental parameters to optimize incubation durations (Hoehler and Jørgensen, 2013). As defining meaningful scales for microbiology research is a recurring challenge (Paerl and Steppe, 2003), the scale of the microbial impact radius determines the choice of method. For example, in a soil environment techniques based on single-cell resolution rarely provide adequate insights for systems-level mass balance and flux calculations. Each method should be tested for the particular type of microbial community studied before results are interpreted, because discrimination against various cell types or taxa may occur (Berney et al., 2007; Teske and Sørensen, 2007; Narasingarao et al., 2011; Franklin et al., 2013; Lloyd et al., 2013; Eloe-Fadrosh et al., 2016).
As the numbers of bulk metagenome and metatranscriptome studies have greatly increased over the last few years (Figure 2), we expect other in situ activity targeted approaches coupled to shotgun sequencing to follow this trend, as in part already observed by the recent increase in DNA-SIP projects. Although DNA- and RNA-SIP methods are still hampered by throughput, high-throughput pipelines will likely become available for broader community use. FACS-based enrichment of active microbial cells may contribute sequence data in the near future because of its accessibility to users and compatibility with a variety of staining or tagging assays (Figure 3). Similarly, SIP in combination with Raman microspectroscopy and sorting is a promising approach, which may soon become available as high-throughput RACS (Zhang et al., 2015b) (Figure 3). A future venture will be determined by the amenability of methods that yield activity-labeled samples in situ to unicellular eukaryotic microbes, including protists and fungi. Although many nucleic acid sequence-based studies encompass well-characterized microbial communities providing a level of understanding of their role in a particular habitat, allowing community modeling (Ruff et al., 2015), we often lack an understanding of eukaryotic microbes. This group of organisms is extremely heterogeneous and there is evidence for its significance in various systems, for example, in airways of cystic fibrosis (Eickmeier et al., 2014), soil (Taylor et al., 2014) and contaminated aquifers (Holmes et al., 2013). On the other hand, inactive eukaryotic microbes may significantly add to the pool of relic DNA. Hence, methods that yield activity-labeled samples in situ should include these organisms. Availability and cost-effectiveness of the necessary equipment, as well as the scalability of data throughput will likely guide the future of methods that yield activity-labeled samples in situ linked to genomics.
Integrating genomic and functional information from the active microbiome
Capturing the genetic makeup of the active microbiome in situ holds great promise to substantially advance our understanding of microbial community functioning. It enables a shift from cataloging the mere presence of taxa and encoded (potential) functions to understanding their ecological roles, including contributions to ecosystem processes. Some of the intriguing and timely research foci in microbial ecology revolve around expanding our knowledge of microbial guilds, groups of species that exploit the same resource, and of reactions that are thermodynamically feasible enough to sustain microbial growth that are lacking associated microorganisms to date. In these cases, nucleic acid sequencing combined with specific labeled substrate assays and coupled to continuous geochemical profiling would, for instance, allow targeting novel organisms that perform redox reactions of interest and enable comprehensive flux modeling of systems (LaRowe and Amend, 2016). Another timely ecological topic is the significance of so-called ‘strongly interacting species’ (SISs) and temporarily dormant cells. SISs are species that have strong positive or negative impacts on the species they directly interact with (Gibson et al., 2016). SISs are not necessarily keystone species as the removal of SISs from a community does not coercively result in mass extinction (Paine, 1995), but they may include organisms of relatively low abundance (Gibson et al., 2016). SISs can have an important role in shaping the steady states of microbial ecosystems regardless of their relative abundance (Gibson et al., 2016). Hence, identifying SISs and studying the processes in their ‘interaction hub’ in a comparative analysis can help reveal metabolic strategies of ecologically relevant microorganisms and interaction strength heterogeneity, for example, across geochemical gradients. SISs may be periodically dormant and hence require time series sampling and/or stimulation experiments (Epstein, 2013). Dormancy represents a bet-hedging strategy, in which an organism can enter a state of low metabolic activity when environmental conditions are disadvantageous and resuscitate when conditions become favorable (Lennon and Jones, 2011). Dormant cells thus generate seed banks, which are likely to impact diversity dynamics and maintenance in a given ecosystem (Jorgensen, 2011). To better understand the contribution of key population size vs organism state to variable environmental conditions, data on total community (for example, targeted using total DNA), population growth (for example, targeted using BrdU, metagenomics) and specific (for example, targeted using SIP, metatranscriptomics, BONCAT and D2O-Raman microspectroscopy with substrate amendments), as well as general activity (for example, targeted using BONCAT, metatranscriptomics, RSG, D2O-Raman microspectroscopy, iRep, PTR) should each be collected in time series experiments. Similarly important is the characterization of the ecological niche of SISs, which may be difficult because of micro-environmental heterogeneity, for instance in soil (Pester et al., 2010) and biofilms (Augspurger et al., 2010). Owing to intricate structures, including channels and voids, a biofilm can host multiple functional groups of cells in various spatial relationships of each other (Gieseke et al., 2003). These spatial arrangements affect nutrient accessibility and therefore influence levels of competition and mutualism between microbial factors (Santegoeds and Damgaard, 1999). The opportunity to resolve the spatial distribution of active microbial cells will disclose the relevance of proximity for microbial interactions with each other, higher organisms or substrate surfaces. Relevant scaling of experimental designs can be achieved using geochemical profiling and flux analysis before and paralleling the use of community analysis. Retrieval of shotgun sequence information remains essential, for instance, when studying genomic variation and gene regulation within species, resulting in increased ecosystem functioning complexity (Darch et al., 2015). Culture-based approaches, metaproteomics and metabolomics studies will effectively complement these studies. Finally, novel single-molecule sequencing methods, such as nanopore technologies (Oxford Nanopore Technologies (ONT; Schneider, 2012), may soon enable the direct measurement and sequencing of chemically modified bases incorporated into the nucleic acids of transcriptionally active or replicating cells. ONT has recently demonstrated the detection of 5-methylcytosine in bacterial and human DNA (Simpson et al.), highlighting its potential to detect other non-canonical bases in the near-term future (Woyke and Rubin, 2014).
Exciting technological advances have shaped the trajectory of microbial community studies, resulting in fascinating insights into our microbial world. We expect this trend to continue as sequence quality improves at reduced costs with a shift from bulk microbial community analyses to more refined studies that target active populations of interest. These improved and novel technologies will assist in exploring the surging scientific questions and applications that have been engaging the scientific community in the field of microbial ecology.
References
Abu Laban N, Tan B, Dao A, Foght J . (2015). Draft genome sequence of uncultivated desulfosporosinus sp. Strain Tol-M, obtained by stable isotope probing using [13C6]toluene. Gen Announc 3: e01422–14.
Alexander H, Jenkins BD, Rynearson TA, Dyhrman ST . (2015a). Metatranscriptome analyses indicate resource partitioning between diatoms in the field. PNAS 112: E2182–E2190.
Alexander H, Rouco M, Haley ST, Wilson ST, Karl DM, Dyhrman ST . (2015b). Functional group-specific traits drive phytoplankton dynamics in the oligotrophic ocean. PNAS 112: E5972–E5979.
Allison SD, Czimczik CI, Treseder KK . (2008). Microbial activity and soil respiration under nitrogen addition in Alaskan boreal forest. Global Change Biol 14: 1156–1168.
Allison SD, Treseder KK . (2008). Warming and drying suppress microbial activity and carbon cycling in boreal forest soils. Global Change Biol 14: 2898–2909.
Artursson V, Finlay RD, Jansson JK . (2005). Combined bromodeoxyuridine immunocapture and terminal-restriction fragment length polymorphism analysis highlights differences in the active soil bacterial metagenome due to Glomus mosseae inoculation or plant species. Environmental Microbiology 7: 1952–1966.
Artursson V, Jansson JK . (2003). Use of Bromodeoxyuridine Immunocapture To Identify Active Bacteria Associated with Arbuscular Mycorrhizal Hyphae. Appl Environ Microbiol 69: 6208–6215.
Ashkin A . (1970). Acceleration and trapping of particles by radiation pressure. Physical review letters 24: 156–159.
Augspurger C, Karwautz C, Mußmann M, Daims H, Battin TJ . (2010). Drivers of bacterial colonization patterns in stream biofilms. FEMS Microbiology Ecology 72: 47–57.
Baker BJ, Sheik CS, Taylor CA, Jain S, Bhasi A, Cavalcoli JD et al. (2013). Community transcriptomic assembly reveals microbes that contribute to deep-sea carbonand nitrogen cycling. ISME J 7: 1962–1973.
Bello-Ortí B, Howell KJ, Tucker AW, Maskell DJ, Aragon V . (2015). Metatranscriptomics reveals metabolic adaptation and induction of virulence factorsby Haemophilus parasuis during lung infection. Vet Res, 1–13.
Berney M, Hammes F, Bosshard F, Weilenmann HU, Egli T . (2007). Assessment and interpretation of bacterial viability by using the live/dead baclight kit in combination with flow cytometry. Appl Environ Microbiol 73: 3283–3290.
Berry D, Mader E, Lee TK, Woebken D, Wang Y, Zhu D et al. (2015). Tracking heavy water (D2O) incorporation for identifying and sorting active microbial cells. PNAS 112: E194–E203.
Bikel S, Valdez-Lara A, Cornejo-Granados F, Rico K, Canizales-Quinteros S, Soberón X et al. (2015). Combining metagenomics, metatranscriptomics and viromics to explore novel microbial interactions: towards a systems-level understanding of human microbiome. CSBJ 13: 390–401.
Blazewicz SJ, Barnard RL, Daly RA, Firestone MK . (2013). Evaluating rRNA as an indicator of microbial activity in environmental communities: limitations and uses. ISME J. 7: 2061–2068.
Bomar L, Maltz M, Colston S, Graf J . (2011). Directed culturing of microorganisms using metatranscriptomics. mBio 2: e00012–11–e00012–11.
Borneman J . (1999). Culture-independent identification of microorganisms that respond to specified stimuli. Appl Environ Microbiol 65: 3398–3400.
Brown CT, Olm MR, Thomas BC, Banfield JF . (2016). in situ replication rates for uncultivated bacteria in microbial communities. bioRxiv e-pub ahead of print, doi:10.1101/057992.
Buerger S, Spoering A, Gavrish E, Leslin C, Ling L, Epstein SS . (2012). Microbial scout hypothesis, stochastic exit from dormancy, and the nature of slow growers. Appl Environ Microbiol 78: 3221–3228.
Burow LC, Woebken D, Marshall IP, Lindquist EA, Bebout BM, Prufert-Bebout L et al. (2012). Anoxic carbon flux in photosynthetic microbial mats as revealed by metatranscriptomics. ISME J 7: 817–829.
Carvalhais LC, Muzzi F, Tan C-H, Hsien-Choo J, Schenk PM . (2013). Plant growth in Arabidopsis is assisted by compost soil-derived microbial communities. Front Plant Sci 4: 235.
Chaparro JM, Badri DV, Vivanco JM . (2013). Rhizosphere microbiome assemblage is affected by plant development. ISME J 8: 790–803.
Chapelle E, Mendes R, Bakker PAH, Raaijmakers JM . (2015). Fungal invasion of the rhizosphere microbiome. ISME J 10: 265–268.
Chemerys A, Pelletier E, Cruaud C, Martin F, Violet F, Jouanneau Y . (2014). Characterization of novel polycyclic aromatic hydrocarbon dioxygenases from the bacterial metagenomic DNA of a contaminated soil. Appl Environ Microbiol 80: 6591–6600.
Chen Y, Dumont MG, Neufeld JD, Bodrossy L, Stralis-Pavese N, McNamara NP et al. (2008). Revealing the uncultivated majority: combining DNA stable-isotope probing, multiple displacement amplification and metagenomic analyses of uncultivated Methylocystis in acidic peatlands. Environ Microbiol 10: 2609–2622.
Damon C, Vallon L, Zimmermann S, Haider MZ, Galeote V, Dequin S et al. (2011). A novel fungal family of oligopeptide transporters identified by functional metatranscriptomics of soil eukaryotes. ISME J 5: 1871–1880.
Darch SE, McNally A, Harrison F, Corander J, Barr HL, Paszkiewicz K et al. (2015). Recombination is a key driver of genomic and phenotypic diversity in a Pseudomonas aeruginosa population during cystic fibrosis infection. Scientific Reports 5: 7649.
David MM, Cecillon S, Warne BM, Prestat E, Jansson JK, Vogel TM . (2014). Microbial ecology of chlorinated solvent biodegradation. Environ Microbiol 17: 4835–4850.
De Filippis F, Genovese A, Ferranti P, Gilbert JA, Ercolini D . (2016). Metatranscriptomics reveals temperature-driven functional changes in microbiome impacting cheese maturation rate. Sci Rep, 1–11.
Dieterich DC, Link AJ, Graumann J, Tirrell DA, Schuman EM . (2006). Selective identification of newly synthesized proteins in mammalian cells using bioorthogonal noncanonical amino acid tagging (BONCAT). Proceedings of the National Academy of Sciences 103: 9482–9487.
Dumont MG, Pommerenke B, Casper P . (2013). Using stable isotope probing to obtain a targeted metatranscriptome of aerobic methanotrophs in lake sediment. Environmental Microbiology Reports 5: 757–764.
Dumont MG, Radajewski SM, Miguez CB, McDonald IR, Murrell JC . (2006). Identification of a complete methane monooxygenase operon from soil by combining stable isotope probing and metagenomic analysis. Environ Microbiol 8: 1240–1250.
Edlund A, Hårdeman F, Jansson JK, Sjöling S . (2008). Active bacterial community structure along vertical redox gradients in Baltic Sea sediment. Environmental Microbiology 10: 2051–2063.
Eichorst SA, Strasser F, Woyke T, Schintlmeister A, Wagner M, Woebken D . (2015). Advancements in the application of NanoSIMS and Raman microspectroscopy to investigate the activity of microbial cells in soils Häggblom M (ed). FEMS Microbiology Ecology 91: 1–14.
Eickmeier O, Hector A, Singh A, Chotirmall SH, Hartl D . (2014). Fungi in cystic fibrosis: recent findings and unresolved questions. Curr Fungal Infect Rep 9: 1–5.
Eloe-Fadrosh EA, Ivanova NN, Woyke T, Kyrpides NC . (2016). Metagenomics uncovers gaps in amplicon-based detection of microbial diversity. Nature Microbiology 1: 15032.
Embree M, Nagarajan H, Movahedi N, Chitsaz H, Zengler K . (2013). Single-cell genome and metatranscriptome sequencing reveal metabolic interactions ofan alkane-degrading methanogenic community. ISME J 8: 757–767.
Epstein SS . (2013). ScienceDirectThe phenomenon of microbial uncultivability. Current Opinion in Microbiology 16: 636–642.
Eren AM, Esen ÖC, Quince C, Vineis JH, Morrison HG, Sogin ML et al. (2015). Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ 3: e1319.
Eyice OZ, Namura M, Chen Y, Mead A, Samavedam S . (2015). SIP metagenomics identifies uncultivated Methylophilaceae as dimethylsulphide degrading bacteria in soil and lake sediment. ISME J 9: 2336–2348.
Feike J, Jürgens K, Hollibaugh JT, Krüger S, Jost G, Labrenz M . (2011). Measuring unbiased metatranscriptomics in suboxic waters of the central Baltic Sea using a new in situ fixation system. ISME J 6: 461–470.
Fortunato CS, Huber JA . (2015). Coupled RNA-SIP and metatranscriptomics of active chemolithoautotrophic communities at a deep-sea hydrothermal vent. ISME J 10: 1925–1938.
Franklin RB, Luria C, Shozo Ozaki L, Bukaveckas PA . (2013). Community composition and activity state of estuarine bacterioplankton assessed using differential staining and metagenomic analysis of 16 S rDNA and rRNA. Aquat Microb Ecol 69: 247–261.
Franzosa EA, Morgan XC, Segata N, Waldron L, Reyes J, Earl AM et al. (2014). Relating the metatranscriptome and metagenome of the human gut. PNAS 111: E2329–E2338.
Frias-Lopez J, Shi Y, Tyson GW, Coleman ML, Schuster SC, Chisholm SW et al. (2008). Microbial community gene expression in ocean surface waters. PNAS 105: 3805–3810.
Gibson TE, Bashan A, Cao H-T, Weiss ST, Liu Y-Y . (2016). On the Origins and Control of Community Types in the Human Microbiome. In: Gore J (ed). PLoS Comput Biol 12: e1004688.
Gieseke A, Bjerrum L, Wagner M, Amann R . (2003). Structure and activity of multiple nitrifying bacterial populations co‐existing in a biofilm. Environmental Microbiology 5: 355–369.
Gifford SM, Sharma S, Rinta-Kanto JM, Moran MA . (2010). Quantitative analysis of a deeply sequenced marine microbial metatranscriptome. The ISME Journal 5: 461–472.
Goldfarb KC, Karaoz U, Hanson CA, Santee CA, Bradford MA, Treseder KK et al. (2011). Differential growth responses of soil bacterial taxa to carbon substrates of varying chemical recalcitrance. Frontiers in Microbiology 2: 94.
Gruber-Vodicka HR, Dirks U, Leisch N, Baranyi C, Stoecker K, Bulgheresi S et al. (2011). Paracatenula, an ancient symbiosis between thiotrophic Alphaproteobacteria and catenulid flatworms. Proceedings of the National Academy of Sciences 108: 12078–12083.
Gutierrez T, Singleton DR, Berry D, Yang T, Aitken MD, Teske A . (2013). Hydrocarbon-degrading bacteria enriched by the Deepwater Horizon oil spill identified by cultivation and DNA-SIP. ISME J 7: 2091–2104.
Hamasaki K, Taniguchi A, Tada Y, Kaneko R, Miki T . (2016). Active populations of rare microbes in oceanic environments as revealed by bromodeoxyuridine incorporation and 454 tag sequencing. Gene 576: 650–656.
Handelsman J . (2004). Metagenomics: application of genomics to uncultured microorganisms. Microbiology and Molecular Biology Reviews 68: 669–685.
Hanson CA, Allison SD, Bradford MA, Wallenstein MD, Treseder KK . (2008). Fungal Taxa Target Different Carbon Sources in Forest Soil. Ecosystems 11: 1157–1167.
Haroon MF, Hu S, Shi Y, Imelfort M, Keller J, Hugenholtz P et al. (2013). Anaerobic oxidation of methane coupled to nitrate reduction in a novel archaeal lineage. Nature, 1–7.
Hatzenpichler R, Connon SA, Goudeau D, Malmstrom RR, Woyke T, Orphan VJ . (2016). Visualizing in situ translational activity for identifying and sorting slow-growing archaeal−bacterial consortia. PNAS 113: E4069–E4078.
Hatzenpichler R, Scheller S, Tavormina PL, Babin BM, Tirrell DA, Orphan VJ . (2014). in situ visualization of newly synthesized proteins in environmental microbes using amino acid tagging and click chemistry. Environ Microbiol 16: 2568–2590.
He S, Wurtzel O, Singh K, Froula JL, Yilmaz S, Tringe SG et al. (2010). Validation of two ribosomal RNA removal methods for microbial metatranscriptomics. Nat Methods 7: 807–812.
Helbling DE, Ackermann M, Fenner K, Kohler H-PE, Johnson DR . (2011). The activity level of a microbial community function can be predicted from its metatranscriptome. ISME J 6: 902–904.
Hellman M, Berg J, Brandt KK, Hallin S . (2011). Journal of Microbiological Methods. Journal of microbiological methods 86: 376–378.
Hewson I, Poretsky RS, Dyhrman ST, Zielinski B, White AE, Tripp HJ et al. (2009). Microbial community gene expression within colonies of the diazotroph, Trichodesmium, from the Southwest Pacific Ocean. The ISME Journal 3: 1286–1300.
Hilton JA, Satinsky BM, Doherty M, Zielinski B, Zehr JP . (2014). Metatranscriptomics of N2-fixing czanobacteria in the Amayon River plume. ISME J 9: 1557–1569.
Hoehler TM, Jørgensen BB . (2013). Microbial life under extreme energy limitation. Nat Rev Micro 11: 83–94.
Holmes DE, Giloteaux L, Williams KH, Wrighton KC, Wilkins MJ, Thompson CA et al. (2013). Enrichment of specific protozoan populations during in situ bioremediation of uranium- contaminated groundwater. The ISME Journal 7: 1286–1298.
Hua Z-S, Han Y-J, Chen L-X, Liu J, Hu M, Li S-J et al. (2014). Ecological roles of dominant and rare prokaryotes in acid mine drainage revealed by metagenomics and metatranscriptomics. ISME J 9: 1280–1294.
Huang WE, Ferguson A, Singer AC, Lawson K, Thompson IP, Kalin RM et al. (2008). Resolving genetic functions within microbial populations: in situ analyses using rRNA and mrna stable isotope probing coupled with single-cell Raman-fluorescence in situ hybridization. Appl Environ Microbiol 75: 234–241.
Huang WE, Stoecker K, Griffiths R, Newbold L, Daims H, Whiteley AS et al. (2007). Raman-FISH: combining stable-isotope Raman spectroscopy and fluorescence in situ hybridization for the single cell analysis of identity and function. Environmental Microbiology 9: 1878–1889.
Huang WE, Ward AD, Whiteley AS . (2009). Raman tweezers sorting of single microbial cells. Environmental Microbiology Reports 1: 44–49.
Hutchens E, Radajewski S, Dumont MG, McDonald IR, Murrell JC . (2004). Analysis of methanotrophic bacteria in Movile Cave by stable isotope probing. Environmental Microbiology 6: 111–120.
Ishii S, Suzuki S, Tenney A, Norden-Krichmar TM, Nealson KH, Bretschger O . (2015). Microbial metabolic networks in a complex electrogenic bio lm recovered from a stimulus-induced metatranscriptomics approach. Sci Rep, 1–14.
Jehmlich N, Schmidt F, Taubert M, Seifert J, Bastida F, Bergen Von M et al. (2010). Protein-based stable isotope probing. Nat Protoc 5: 1957–1966.
Jiang Y, Xiong X, Danska J, Parkinson J . (2016). Metatranscriptomic analysis of diverse microbial communities reveals core metabolic pathways and microbiome- specific functionality. Microbiome, 1–18.
Jones DS, Flood BE, Bailey JV . (2015). Metatranscriptomic insights into polyphosphate metabolism in marine sediments. ISME J 10: 1015–1019.
Jorgensen BB . (2011). Deep subseafloor microbial cells on physiological standby. PNAS 108: 18193–18194.
Jorth P, Turner KH, Gumus P, Nizam N, Buduneli N, Whiteley M . (2014). Metatranscriptomics of the human oral microbiome during health and disease. mBio 5: e01012–14–e01012–14.
Kalyuzhnaya MG, Hristova KR, Lidstrom ME, Chistoserdova L . (2008a). characterization of a novel methanol dehydrogenase in representatives of burkholderiales: implications for environmental detection of methylotrophy and evidence for convergent evolution. J Bacteriol 190: 3817–3823.
Kalyuzhnaya MG, Lapidus A, Ivanova N, Copeland AC, Mchardy AC, Szeto E et al. (2008b). High-resolution metagenomics targets specific functional types in complex microbial communities. Nat Biotechnol 26: 1029–1034.
Kalyuzhnaya MG, Lidstrom ME, Chistoserdova L . (2008). Real-time detection of actively metabolizing microbes by redox sensing as applied to methylotroph populations in Lake Washington. The ISME Journal 2: 696–706.
Kalyuzhnaya MG, Lidstrom ME, Chistoserdova L . (2008c). Real-time detection of actively metabolizing microbes by redox sensing as applied to methylotroph populations in Lake Washington. ISME J 2: 696–706.
Kim SJ, Park S-J, Jung MY, Kim JG, Madsen EL, Rhee SK . (2014). An uncultivated nitrate-reducing member of the genus herminiimonas degrades toluene. Appl Environ Microbiol 80: 3233–3243.
Klindworth A, Mann AJ, Huang S, Wichels A, Quast C, Waldmann J et al. (2014). Diversity and activity of marine bacterioplankton during a diatom bloom in the North Sea assessed by total RNA and pyrotag sequencing. Mar Genom 18: 185–192.
Kodzius R, Gojobori T . (2016). Single-cell technologies in environmental omics. Gene 576: 701–707.
Konopka MC, Strovas TJ, Ojala DS, Chistoserdova L, Lidstrom ME, Kalyuzhnaya MG . (2010). Respiration response imaging for real-time detection of microbial function at the single-cell level. Appl Environ Microbiol 77: 67–72.
Kopf SH, McGlynn SE, Green-Saxena A, Guan Y, Newman DK, Orphan VJ . (2015). Heavy water and 15 N labelling with NanoSIMS analysis reveals growth rate-dependent metabolic heterogeneity in chemostats. Environmental Microbiology 17: 2542–2556.
Korem T, Zeevi D, Suez J, Weinberger A, Avnit-Sagi T, Pompan-Lotan M et al. (2015). Growth dynamics of gut microbiota in health and disease inferred from single metagenomic samples. Science 349: 1101–1106.
Lamendella R, Strutt S, Borglin S, Chakraborty R, Tas N, Mason OU et al. (2014). Assessment of the Deepwater Horizon oil spill impact on Gulf coast microbial communities. Front Microbiol 5. e-pub ahead of print, doi: 10.3389/fmicb.2014.00130/abstract.
LaRowe DE, Amend JP . (2016). The energetics of anabolism in natural settings. ISME J. 10: 1285–1295.
Lasken RS . (2007). Single-cell genomic sequencing using Multiple Displacement Amplification. Current Opinion in Microbiology 10: 510–516.
Lehembre F, Doillon D, David E, Perrotto S, Baude J, Foulon J et al. (2013). Soil metatranscriptomics for mining eukaryotic heavy metal resistance genes. Environ Microbiol 15: 2829–2840.
Leininger S, Urich T, Schloter M, Schwark L, Qi J, Nicol GW et al. (2006). Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature 442: 806–809.
Lennon JT, Jones SE . (2011). Microbial seed banks: the ecological and evolutionary implications of dormancy. Nat Rev Micro 9: 119–130.
Lesniewski RA, Jain S, Anantharaman K, Schloss PD, Dick GJ . (2012). The metatranscriptome of a deep-sea hydrothermal plume is dominated by water column methanotrophs and lithotrophs. The ISME Journal 6: 2257–2268.
Lester W, Sun SH, Seber A . (1960). Observations on the influence of deuterium on bacterial growth. Annals of the New York Academy of Sciences 84: 667–677.
Lim YW, Schmieder R, Haynes M, Willner D, Furlan M, Youle M et al. (2013). Metagenomics and metatranscriptomics: windows on CF-associated viral and microbial communities. J Cyst Fibros 12: 154–164.
Lin S, Zhang H, Zhuang Y, Tran B . (2010). Spliced leader–based metatranscriptomic analyses lead to recognition of hidden genomic features in dinoflagellates. Proc Natl Acad Sci 107: 20033–20038.
Lloyd KG, May MK, Kevorkian RT, Steen AD . (2013). Meta-analysis of quantification methods shows that archaea and bacteria have similar abundances in the subseafloor. Appl Environ Microbiol 79: 7790–7799.
Majed N, Chernenko T, Diem M, Gu AZ . (2012). Identification of functionally relevant populations in enhanced biological phosphorus removal processes based on intracellular polymers profiles and insights into the metabolic diversity and heterogeneity. Environmental Science & Technology 46: 5010–5017.
Manefield M, Whiteley AS, Griffiths RI, Bailey MJ . (2002). RNA Stable Isotope Probing, a Novel Means of Linking Microbial Community Function to Phylogeny. Appl Environ Microbiol 68: 5367–5373.
Marzano S-YL, Domier LL . (2016). Novel mycoviruses discovered from metatranscriptomics survey of soybean phyllosphere phytobiomes. Virus Res 213: 332–342.
McGuire KL, Bent E, Borneman J, Majumder A, Allison SD, Tresederi KK . (2012). Functional diversity in resource use by fungi. Ecology 91: 2324–2332.
Milucka J, Ferdelman TG, Polerecky L, Franzke D, Wegener G, Schmid M et al. (2012). Zero-valent sulphur is a key intermediate in marine methane oxidation. Nature 491: 541–546.
Mondot S, Lepage P . (2016). The human gut microbiome and its dysfunctions through the meta-omics prism. Ann N Y Acad Sci, 1–11; http://doi.org/10.1111/nyas.13033.
Moran MA, Satinsky B, Gifford SM, Luo H, Rivers A, Chan L-K et al. (2012). Sizing up metatranscriptomics. ISME J. 7: 237–243.
Mott ML, Berger JM . (2007). DNA replication initiation: mechanisms and regulation in bacteria. Nat Rev Micro 5: 343–354.
Mou X, Sun S, Edwards RA, Hodson RE, Moran MA . (2008). Bacterial carbon processing by generalist species in the coastal ocean. Nature 451: 708–711.
Narasingarao P, Podell S, Ugalde JA, Brochier-Armanet CEL, Emerson JB, Brocks JJ et al. (2011). De novo metagenomic assembly reveals abundant novel major lineage of Archaea in hypersaline microbial communities. The ISME Journal 6: 81–93.
Neufeld JD, Chen Y, Dumont MG, Murrell JC . (2008). Marine methylotrophs revealed by stable-isotope probing, multiple displacement amplification and metagenomics. Environ Microbiol 10: 1526–1535.
Neufeld JD, Dumont MG, Vohra J, Murrell JC . (2006). Methodological Considerations for the Use of Stable Isotope Probing in Microbial Ecology. Microb Ecol 53: 435–442.
Neufeld JD, Vohra J, Dumont MG, Lueders T, Manefield M, Friedrich MW et al. (2007). DNA stable-isotope probing. Nat Protoc 2: 860–866.
Orsi WD, Edgcomb VP, Christman GD, Biddle JF . (2013). Gene expression in the deep biosphere. Nature 499: 205–208.
Ottman N, Smidt H, De Vos WM, Belzer C . (2012). The function of our microbiota: who is out there and what do they do? Front Cell Inf Microbio 2: 104.
Paerl HW, Steppe TF . (2003). Scaling up: the next challenge in environmental microbiology. Environmental Microbiology 5: 1025–1038.
Pagani I, Liolios K, Jansson J, Chen I-MA, Smirnova T, Nosrat B et al. (2011). The Genomes OnLine Database (GOLD) v.4: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Research 40: D571–D579.
Paine RT . (1995), A conversation on refining the concept of keystone species.
Pester M, Bittner N, Deevong P, Wagner M, Loy A . (2010). A ‘rare biosphere’ microorganism contributes to sulfate reduction in a peatland. The ISME Journal, 1–12.
Pinnell LJ, Dunford E, Ronan P, Hausner M, Neufeld JD . (2014). Recovering glycoside hydrolase genes from active tundra cellulolytic bacteria. Can J Microbiol 60: 469–476.
Poretsky RS, Sun S, Mou X, Moran MA . (2010). Transporter genes expressed by coastal bacterioplankton in response to dissolved organic carbon. Environmental Microbiology 12: 616–627.
Radajewski S, Ineson P, Parekh NR, Murrell JC . (2000). Stable-isotope probing as a tool in microbial ecology. Nature 403: 646–649.
Radajewski S, McDonald IR, Murrell JC . (2003). Stable-isotope probing of nucleic acids: a window to the function of uncultured microorganisms. Current Opinion in Biotechnology 14: 296–302.
Ruff SE, Biddle JF, Teske AP, Knittel K, Boetius A, Ramette A . (2015). Global dispersion and local diversification of the methane seep microbiome. PNAS 112: 4015–4020 201421865.
Sanders JG, Beinart RA, Stewart FJ, DeLong EF, Girguis PR . (2013). Metatranscriptomics reveal differences in in situ energy and nitrogen metabolism among hydrothermal vent snail symbionts. ISME J 7: 1556–1567.
Santegoeds CM, Damgaard LR . (1999). Distribution of sulfate-reducing and methanogenic bacteria in anaerobic aggregates determined by microsensor and molecular analyses. Appl Environ Microbiol 65: 4618–4629.
Satinsky BM, Crump BC, Smith CB, Sharma S, Zielinski BL, Doherty M et al. (2014). Microspatial gene expression patterns in the Amazon River Plume. PNAS 111: 11085–11090.
Schneider G. F, Dekker C . (2012) News and Views. Nature Publishing Group, 30 (4): 326–328; http://doi.org/10.1038/nbt.2181.
Schwab C, Tveit AT, Schleper C, Urich T . (2014). Gene expression of lactobacilli in murine forestomach biofilms. Microbial Biotechnol 7: 347–359.
Schwartz E, Hayer M, Hungate BA, Koch BJ, McHugh TA, Mercurio W et al. (2016). Stable isotope probing with 18O-water to investigate microbial growth and death in environmental samples. Curr Opin Biotechnol 41: 14–18.
Shi Y, Tyson GW, Delong EF . (2009). Metatranscriptomics reveals unique microbial small RNAs in the ocean’s water column. Nature 459: 266–269.
Simberloff D, Dayan T . (1991). The guild concept and the structure of ecological communities. Annu Rev Ecol Syst 22: 115–143.
Simpson JT, Workman R, Zuzarte PC, David M, Dursi LJ, Timp W Detecting DNA Methylation using the Oxford Nanopore Technologies MinION sequencer. e-pub ahead of print doi:10.1101/047142.
Simón-Soro A, Guillen-Navarro M, Mira A . (2014). Metatranscriptomics reveals overall active bacterial composition in caries lesions. J Oral Microbiol 6: 1252.
Song Y, Yin H, Huang WE . (2016). ScienceDirectRaman activated cell sorting. Current opinion in chemical biology 33: 1–8.
Steen IH, Dahle H, Stokke R, Roalkvam I, Daae F-L, Rapp HT et al. (2016). Novel Barite Chimneys at the Loki's Castle Vent Field Shed Light on Key Factors Shaping Microbial Communities and Functions in Hydrothermal Systems. Fron Microbiol 6: 4248.
Sul WJ, Park J, Quensen JF, Rodrigues JLM, Seliger L, Tsoi TV et al. (2009). DNA-stable isotope probing integrated with metagenomics for retrieval of biphenyl dioxygenase genes from polychlorinated biphenyl-contaminated river sediment. Appl Environ Microbiol 75: 5501–5506.
Taylor DL, Hollingsworth TN, McFarland JW . (2014). A first comprehensive census of fungi in soil reveals both hyperdiversity and fine-scale niche partitioning. Ecological Monographs 84: 3–20.
Teske A, Sørensen KB . (2007). Uncultured archaea in deep marine subsurface sediments: have we caught them all? The ISME Journal 2: 3–18.
Tringe SG, Hugenholtz P . (2008). A renaissance for the pioneering 16 S rRNA gene. Current Opinion in Microbiology 11: 442–446.
Ullrich S, Karrasch B, Hoppe H, Jeskulke K, Mehrens M . (1996). Toxic effects on bacterial metabolism of the redox dye 5-cyano-2,3-ditolyl tetrazolium chloride. Appl Environ Microbiol 62: 4587–4593.
Urbach E, Vergin KL, Giovannoni SJ . (1999). Immunochemical detection and isolation of DNA from metabolically active bacteria. Appl Environ Microbiol 65: 1207–1213.
Urich T, Lanzén A, Qi J, Huson DH, Schleper C, Schuster SC . (2008). Simultaneous assessment of soil microbial community structure and function through analysis of the meta-transcriptome Ward N (ed). PLoS One 3: e2527.
Ursell LK, Knight R . (2013). Xenobiotics and the human gut microbiome: metatranscriptomics reveal the active players. Cell Metab 17: 317–318.
Verastegui Y, Cheng J, Engel K, Kolczynski D, Mortimer S, Lavigne J et al. (2014). Multisubstrate isotope labeling and metagenomic analysis of active soil bacterial communities. mBio 5: e01157–14–e01157–14.
Vila-Costa M, Sharma S, Moran MA, Casamayor EO . (2012). Diel gene expression profiles of a phosphorus limited mountain lake using metatranscriptomics. Environ Microbiol 15: 1190–1203.
Wang T-H, Peng Y, Zhang C, Wong PK, Ho C-M . (2005). Single-molecule tracing on a fluidic microchip for quantitative detection of low-abundance nucleic acids. Journal of the American Chemical Society 127: 5354–5359.
Wang Y, Huang WE, Cui L, Wagner M . (2016). Single cell stable isotope probing in microbiology using Raman microspectroscopy. Current Opinion in Biotechnology 41: 34–42.
Wang Y, Ji Y, Wharfe ES, Meadows RS, March P, Goodacre R et al. (2013). Raman Activated Cell Ejection for Isolation of Single Cells. Anal Chem 85: 10697–10701.
Werner JJ, Garcia ML, Perkins SD, Yarasheski KE, Smith SR, Muegge BD et al. (2014). Microbial community dynamics and stability during an ammonia-induced shift to syntrophic acetate oxidation. Appl Environ Microbiol 80: 3375–3383.
Whiteley AS, Thomson B, Lueders T, Manefield M . (2007). RNA stable-isotope probing. Nat Protoc 2: 838–844.
Woyke T, Rubin EM . (2014). Searching for new branches on the tree of life. Science 346: 698–699.
Xie C, Dinno MA, Li Y-Q . (2002). Near-infrared Raman spectroscopy of single optically trapped biological cells. Opt Lett 27: 249–251.
Xie C, Li Y-Q . (2003). Confocal micro-Raman spectroscopy of single biological cells using optical trapping and shifted excitation difference techniques. J Appl Phys 93: 2982.
Yin B, Crowley D, Sparovek G, De Melo WJ, Borneman J . (2000). Bacterial functional redundancy along a soil reclamation gradient. Appl Environ Microbiol 66: 4361–4365.
Zhang P, Ren L, Zhang X, Shan Y, Wang Y . (2015a). Raman-activated cell sorting based on dielectrophoretic single-cell trap and release. Anal Chem 87: 2282–2289.
Zhang Q, Zhang P, Gou H, Mou C, Huang WE, Yang M et al. (2015b). Towards high-throughput microfluidic Raman-activated cell sorting. Analyst 140: 6163–6174.
Zhang T, Tremblay PL, Chaurasia AK, Smith JA, Bain TS, Lovley DR . (2013). Anaerobic benzene oxidation via phenol in geobacter metallireducens. Appl Environ Microbiol 79: 7800–7806.
Acknowledgements
This work was conducted by the US Department of Energy Joint Genome Institute, a DOE Office of Science User Facility, and was supported under contract no. DE-AC02-05CH11231. Research in the Wagner lab on single-cell activity assays was supported by an ERC Advanced Grant (NITRICARE, 294343), as well as by the Emerging Technologies Opportunity Program, which is supported by the Office of Science of the US Department of Energy under Contract DE-AC02-05CH11231.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Singer, E., Wagner, M. & Woyke, T. Capturing the genetic makeup of the active microbiome in situ. ISME J 11, 1949–1963 (2017). https://doi.org/10.1038/ismej.2017.59
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.59
This article is cited by
-
Advances in metaproteomic profiling of molecular microbiology and environmental responses
Systems Microbiology and Biomanufacturing (2024)
-
Organelle 16S rRNA amplicon sequencing enables profiling of active gut microbiota in murine model
Applied Microbiology and Biotechnology (2022)
-
Microbial ecology of biofiltration used for producing safe drinking water
Applied Microbiology and Biotechnology (2022)
-
Nitrospira in wastewater treatment: applications, opportunities and research gaps
Reviews in Environmental Science and Bio/Technology (2022)
-
Raman microspectroscopy for microbiology
Nature Reviews Methods Primers (2021)