Article
|
Open Access
Featured
-
-
Article
| Open Access Emergent properties as by-products of prebiotic evolution of aminoacylation ribozymes
Emergent properties as by-products of prebiotic evolution of aminoacylation ribozymes
Complex biochemical systems exhibit traits that appear to be highly adapted. Studies of catalytic RNA demonstrate that adaptive traits, such as increased specificity and error tolerance, could originate as evolutionary by-products.
- Evan Janzen
- , Yuning Shen
- & Irene A. Chen
-
Article
| Open Access Probing TDP-43 condensation using an in silico designed aptamer
Probing TDP-43 condensation using an in silico designed aptamer
Here, the authors generated an artificial RNA molecule, or aptamer, specific for the Amyotrophic Lateral Sclerosis protein TDP-43. By interacting avidly with its target, the aptamer can be exploited to track TDP-43 phase transition in vitro and in cells.
- Elsa Zacco
- , Owen Kantelberg
- & Gian Gaetano Tartaglia
-
Article
| Open Access RNase III CLASH in MRSA uncovers sRNA regulatory networks coupling metabolism to toxin expression
RNase III CLASH in MRSA uncovers sRNA regulatory networks coupling metabolism to toxin expression
Regulatory small RNA (sRNA) interact with mRNAs to regulate their stability, transcription, and translation via diverse mechanisms. Here, McKellar et al. apply RNase IIICLASH of multi-drug resistant Staphylococcus aureus under different culture conditions to link the network of RNA-RNA interactions to environmental conditions and find that the production of small membrane-permeabilizing toxins is strongly regulated by sRNAs.
- Stuart W. McKellar
- , Ivayla Ivanova
- & Sander Granneman
-
Article
| Open Access Hydrophobic-cationic peptides modulate RNA polymerase ribozyme activity by accretion
Hydrophobic-cationic peptides modulate RNA polymerase ribozyme activity by accretion
Macromolecular aggregates may have played an important role in the origin of life. Here, the authors report hydrophobic-cationic peptides that form insoluble aggregates, which reversibly accrete RNA on their surfaces, and enhance RNA polymerization by a ribozyme.
- Peiying Li
- , Philipp Holliger
- & Shunsuke Tagami
-
Article
| Open Access The RNA-bound proteome of MRSA reveals post-transcriptional roles for helix-turn-helix DNA-binding and Rossmann-fold proteins
The RNA-bound proteome of MRSA reveals post-transcriptional roles for helix-turn-helix DNA-binding and Rossmann-fold proteins
RNA-binding proteins play key roles in controlling gene expression in many organisms. Here, Chu et al. identify hundreds of RNA-binding proteins in methicillin-resistant Staphylococcus aureus, and show that a major transcription factor uses its helix-turn-helix domain to bind RNAs near intrinsic transcription terminators.
- Liang-Cui Chu
- , Pedro Arede
- & Sander Granneman
-
Article
| Open Access RNA supply drives physiological granule assembly in neurons
RNA supply drives physiological granule assembly in neurons
RNA granules are important regulators of RNA metabolism. Here the authors report that RNA granules containing RNA helicase DDX6 disassemble during neuronal maturation.
- Karl E. Bauer
- , Niklas Bargenda
- & Michael A. Kiebler
-
Article
| Open Access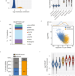 HDLBP binds ER-targeted mRNAs by multivalent interactions to promote protein synthesis of transmembrane and secreted proteins
HDLBP binds ER-targeted mRNAs by multivalent interactions to promote protein synthesis of transmembrane and secreted proteins
RNA binding protein HDLBP (or Vigilin) localizes in the endoplasmic reticulum (ER) membrane. Here the authors show that HDLBP contributes to translation of ER-targeted mRNAs.
- Ulrike Zinnall
- , Miha Milek
- & Markus Landthaler
-
Article
| Open Access Nucleotide-amino acid π-stacking interactions initiate photo cross-linking in RNA-protein complexes
Nucleotide-amino acid π-stacking interactions initiate photo cross-linking in RNA-protein complexes
Although UV-induced cross-linking is a widely used method to study RNA-protein complexes, the cross-linking reactions are poorly understood. Here, the authors show that π-stacking interactions between nucleobases and aromatic amino acids play a key role in the cross-linking process.
- Anna Knörlein
- , Chris P. Sarnowski
- & Jonathan Hall
-
Article
| Open Access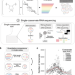 Characterization of RNA content in individual phase-separated coacervate microdroplets
Characterization of RNA content in individual phase-separated coacervate microdroplets
Here, the authors demonstrate that single cell RNA sequencing technology can be leveraged to characterize RNA content of individual membrane-free condensates formed by liquid-liquid phase separation processes such as coacervation.
- Damian Wollny
- , Benjamin Vernot
- & Barbara Treutlein
-
Article
| Open Access Regulated IRE1α-dependent decay (RIDD)-mediated reprograming of lipid metabolism in cancer
Regulated IRE1α-dependent decay (RIDD)-mediated reprograming of lipid metabolism in cancer
IRE1α cleaves several mRNAs upon accumulation of misfolded proteins. Here the authors show that active IRE1α cleaves DGAT2 mRNA encoding the rate-limiting enzyme in the synthesis of triacylglycerols, suggesting a role of IRE1α in reprogramming lipid metabolism in cancer cells.
- Aitor Almanza
- , Katarzyna Mnich
- & Afshin Samali
-
Article
| Open Access Enhancer RNAs stimulate Pol II pause release by harnessing multivalent interactions to NELF
Enhancer RNAs stimulate Pol II pause release by harnessing multivalent interactions to NELF
Enhancer RNAs (eRNAs) can stimulate gene transcription through various mechanisms. Here, the authors identify the molecular features within eRNAs that are critical for their action in facilitating RNA Polymerase II release from the paused state.
- Vladyslava Gorbovytska
- , Seung-Kyoon Kim
- & Claus-D. Kuhn
-
Article
| Open Access Pytheas: a software package for the automated analysis of RNA sequences and modifications via tandem mass spectrometry
Pytheas: a software package for the automated analysis of RNA sequences and modifications via tandem mass spectrometry
RNA modifications represent a critical aspect of RNA biology that is not well suited to sequencing methods. Here, the authors provide a software tool for automated analysis of RNA tandem mass spectra with full support of modifications, isotope labelling, and control of false discovery rate.
- Luigi D’Ascenzo
- , Anna M. Popova
- & James R. Williamson
-
Article
| Open Access CFTR mRNAs with nonsense codons are degraded by the SMG6-mediated endonucleolytic decay pathway
CFTR mRNAs with nonsense codons are degraded by the SMG6-mediated endonucleolytic decay pathway
Currently, there is no therapy for patients with cystic fibrosis caused by nonsense mutations. Here the authors show that CFTR mRNAs with nonsense codons are predominantly degraded by the SMG6-mediated branch of the NMD pathway, providing potential therapeutic strategies for the devastating disease.
- Edward J. Sanderlin
- , Melissa M. Keenan
- & Lulu Huang
-
Article
| Open Access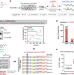 TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer
TRMT6/61A-dependent base methylation of tRNA-derived fragments regulates gene-silencing activity and the unfolded protein response in bladder cancer
RNA modifications are important regulators of RNA biology. Here we report N1-methyladenosine (m1A) enrichment on 22-nucleotide tRNA fragments and its effect on gene-silencing. Higher level of m1A in bladder cancer is accompanied by gene dysregulation in unfolded protein response.
- Zhangli Su
- , Ida Monshaugen
- & Anindya Dutta
-
Article
| Open Access DENR controls JAK2 translation to induce PD-L1 expression for tumor immune evasion
DENR controls JAK2 translation to induce PD-L1 expression for tumor immune evasion
Several mechanisms have been associated with transcriptional and post-transcriptional regulation of PD-L1 in cancer. Here the authors show that a RNA binding protein, DENR, positively regulates the translation of JAK2 and induces PD-L1 expression in tumor cells, associated with immune evasion.
- Baiwen Chen
- , Jiajia Hu
- & Hua-Bing Li
-
Article
| Open Access Low complexity RGG-motif sequence is required for Processing body (P-body) disassembly
Low complexity RGG-motif sequence is required for Processing body (P-body) disassembly
Processing bodies (PBs) are higher order RNA-protein complexes that determine mRNA fate. Here, the authors show Sbp1 is responsible for PB disassembly and pinpoint a repeats of arginine and glycine (RGG) motif in Sbp1 that play a key role.
- Raju Roy
- , Gitartha Das
- & Purusharth I. Rajyaguru
-
Article
| Open Access Reprogrammed tracrRNAs enable repurposing of RNAs as crRNAs and sequence-specific RNA biosensors
Reprogrammed tracrRNAs enable repurposing of RNAs as crRNAs and sequence-specific RNA biosensors
In type II CRISPR systems, the guide RNA (gRNA) comprises a CRISPR RNA (crRNA) and a hybridized trans-acting CRISPR RNA (tracrRNA), both being essential in guided DNA targeting functions. Here the authors investigate the programmability of crRNA-tracrRNA hybridization for Cas9 and apply this to biosensing.
- Yang Liu
- , Filipe Pinto
- & Baojun Wang
-
Article
| Open Access SHAPE-guided RNA structure homology search and motif discovery
SHAPE-guided RNA structure homology search and motif discovery
SHAPEwarp is a method that allows identifying structurally-similar RNAs by direct comparison of reactivity profiles derived from chemical probing experiments. Its application to viral genomes identified conserved RNA structure elements.
- Edoardo Morandi
- , Martijn J. van Hemert
- & Danny Incarnato
-
Article
| Open Access Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
Combinatorial optimization of mRNA structure, stability, and translation for RNA-based therapeutics
The authors develop an RNA sequencing-based platform, PERSIST-seq, to simultaneously delineate in-cell mRNA stability, ribosome load, and in-solution stability of a diverse mRNA library to derive design principles for improved mRNA therapeutics.
- Kathrin Leppek
- , Gun Woo Byeon
- & Rhiju Das
-
Article
| Open Access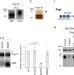 Siwi cooperates with Par-1 kinase to resolve the autoinhibitory effect of Papi for Siwi-piRISC biogenesis
Siwi cooperates with Par-1 kinase to resolve the autoinhibitory effect of Papi for Siwi-piRISC biogenesis
Siwi-piRISC protects the germline genome from DNA damage caused by selfish movement of transposons by suppressing their expression. Here, the authors show how molecularly Papi, which plays an important role in the production of Siwi-piRISC, cooperates with Par-1 kinase to ensure the accumulation of Siwi-piRISC in germ cells.
- Hiromi Yamada
- , Kazumichi M. Nishida
- & Mikiko C. Siomi
-
Article
| Open Access Rational design of hairpin RNA excited states reveals multi-step transitions
Rational design of hairpin RNA excited states reveals multi-step transitions
RNA molecules exhibit conformational fluctuations between ground states and excited states. Here the authors designed and verified small hairpin RNAs with predefined secondary structure reshufflings. In light of Van’t Hoff analysis and accelerated molecular dynamics simulation, a mechanism of multistep sequential transition has been revealed.
- Ge Han
- & Yi Xue
-
Article
| Open Access An RNA-binding protein acts as a major post-transcriptional modulator in Bacillus anthracis
An RNA-binding protein acts as a major post-transcriptional modulator in Bacillus anthracis
HitRS is a two-component system that responds to cell envelope damage in the bacterium Bacillus anthracis. Here, the authors identify an RNA-binding protein that regulates HitRS function by modulating the stability of the hitRS mRNA. In addition, the protein binds to over 70 RNAs and affects the expression of genes involved in multiple cellular processes.
- Hualiang Pi
- , Andy Weiss
- & Eric P. Skaar
-
Article
| Open Access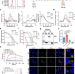 RNA G-quadruplex in TMPRSS2 reduces SARS-CoV-2 infection
RNA G-quadruplex in TMPRSS2 reduces SARS-CoV-2 infection
Understanding the mechanisms of SARS-CoV-2 infection is important to control the pandemic. Here the authors show the biological and pathological role of RNA G-quadruplex structure in both SARS-CoV-2 genome and host factors, particularly TMPRSS2.
- Geng Liu
- , Wenya Du
- & Xianghui Fu
-
Article
| Open Access The methyl phosphate capping enzyme Bmc1/Bin3 is a stable component of the fission yeast telomerase holoenzyme
The methyl phosphate capping enzyme Bmc1/Bin3 is a stable component of the fission yeast telomerase holoenzyme
There exists a remarkable evolutionary breadth of telomerase holoenzyme components among eukaryotes. Here the authors report that the fission yeast methyl phosphate capping enzyme Bin3/Bmc1 regulates telomerase activity by promoting holoenzyme formation.
- Jennifer Porat
- , Moaine El Baidouri
- & Mark A. Bayfield
-
Article
| Open Access Co-translational assembly orchestrates competing biogenesis pathways
Co-translational assembly orchestrates competing biogenesis pathways
The biogenesis of nuclear pores imposes a logistic challenge for cells. Here, the authors investigate structural motifs for co-translational interactions in nucleoporins and find that co-translational assembly events differ between paralogous assembly pathways thus contributing to faithful assembly.
- Maximilian Seidel
- , Anja Becker
- & Martin Beck
-
Article
| Open Access Landscape of adenosine-to-inosine RNA recoding across human tissues
Landscape of adenosine-to-inosine RNA recoding across human tissues
Gabay et al. provide a highly-accurate atlas of recoding by A-to-I RNA editing in human, profiled across tissues and cell subpopulations. Most highly edited sites are evolutionary conserved in non-primate mammals, attesting for adaptation.
- Orshay Gabay
- , Yoav Shoshan
- & Eli Eisenberg
-
Article
| Open Access Secondary structural ensembles of the SARS-CoV-2 RNA genome in infected cells
Secondary structural ensembles of the SARS-CoV-2 RNA genome in infected cells
Lan et al. report RNA structure ensembles across the entire SARSCoV-2 genome in infected human cells at single nucleotide resolution. They find alternative RNA conformations critical for promoting near-native frameshifting rates in ORF1ab.
- Tammy C. T. Lan
- , Matty F. Allan
- & Silvi Rouskin
-
Article
| Open Access Crosstalk between CRISPR-Cas9 and the human transcriptome
Crosstalk between CRISPR-Cas9 and the human transcriptome
The off-target effects of CRISPR-Cas9 are thought to be mediated by its cognate guide RNA. Here the authors show that Cas9 independently interacts with the human transcriptome, correlating with elevated RNA editing even under guide RNA co-expression.
- Aaron A. Smargon
- , Assael A. Madrigal
- & Gene W. Yeo
-
Article
| Open Access Small Cajal body-associated RNA 2 (scaRNA2) regulates DNA repair pathway choice by inhibiting DNA-PK
Small Cajal body-associated RNA 2 (scaRNA2) regulates DNA repair pathway choice by inhibiting DNA-PK
Proper repair of DNA double-strand breaks is essential for genomic stability. Here, the authors report that a long non-coding RNA, scaRNA2, inhibits DNA-PK and thereby regulates the choice between error-prone NHEJ and error-free HR DNA repair.
- Sofie Bergstrand
- , Eleanor M. O’Brien
- & Marianne Farnebo
-
Article
| Open Access Specific length and structure rather than high thermodynamic stability enable regulatory mRNA stem-loops to pause translation
Specific length and structure rather than high thermodynamic stability enable regulatory mRNA stem-loops to pause translation
This study shows that rather than high thermodynamic stability, specific length and structure enable regulatory mRNA stem-loops to pause translation. These findings aid identification of new regulatory mRNA stem-loops.
- Chen Bao
- , Mingyi Zhu
- & Dmitri N. Ermolenko
-
Article
| Open Access Chemical reversible crosslinking enables measurement of RNA 3D distances and alternative conformations in cells
Chemical reversible crosslinking enables measurement of RNA 3D distances and alternative conformations in cells
Determination of RNA 3D structures in vivo is a challenging problem. Here, the authors describe a chemical crosslinking method (SHARC) that they use in combination with RNA sequencing to measure distances between nucleotides in RNA 3D structures.
- Ryan Van Damme
- , Kongpan Li
- & Willem A. Velema
-
Article
| Open Access A late-stage assembly checkpoint of the human mitochondrial ribosome large subunit
A late-stage assembly checkpoint of the human mitochondrial ribosome large subunit
Rebelo-Guiomar et al. unveil late stage assembly intermediates of the human mitochondrial ribosome by inactivating the methyltransferase MRM2 in cells. Absence of MRM2 impairs organismal homeostasis, while its catalytic activity is dispensable for mitoribosomal biogenesis.
- Pedro Rebelo-Guiomar
- , Simone Pellegrino
- & Michal Minczuk
-
Article
| Open Access The RNA methyltransferase METTL8 installs m3C32 in mitochondrial tRNAsThr/Ser(UCN) to optimise tRNA structure and mitochondrial translation
The RNA methyltransferase METTL8 installs m3C32 in mitochondrial tRNAsThr/Ser(UCN) to optimise tRNA structure and mitochondrial translation
RNA modifications are key regulators of RNA functions. Here, the authors identify METTL8 as the enzyme installing m3C32 in mitochondrial tRNAThr/Ser(UCN). Lack of these modifications affects tRNA structure and impairs mitochondrial translation.
- Nicole Kleiber
- , Nicolas Lemus-Diaz
- & Markus T. Bohnsack
-
Article
| Open Access A small RNA that cooperatively senses two stacked metabolites in one pocket for gene control
A small RNA that cooperatively senses two stacked metabolites in one pocket for gene control
Riboswitches contain an aptamer domain that recognizes a metabolite and an expression platform that regulates gene expression. Here the authors report the crystal structure of a preQ1-sensing riboswitch from Carnobacterium antarcticus that shows two metabolites in a single binding pocket.
- Griffin M. Schroeder
- , Chapin E. Cavender
- & Joseph E. Wedekind
-
Article
| Open Access Decoding non-canonical mRNA decay by the endoplasmic-reticulum stress sensor IRE1α
Decoding non-canonical mRNA decay by the endoplasmic-reticulum stress sensor IRE1α
IRE1 helps mitigate endoplasmic-reticulum stress by cleaving specific mRNAs at a conserved sequence endomotif via regulated IRE1-dependent decay (RIDD). Here the authors discover a more promiscuous IRE1 activity dubbed RIDD lacking endomotif (RIDDLE).
- Adrien Le Thomas
- , Elena Ferri
- & Avi Ashkenazi
-
Article
| Open Access The short isoform of the host antiviral protein ZAP acts as an inhibitor of SARS-CoV-2 programmed ribosomal frameshifting
The short isoform of the host antiviral protein ZAP acts as an inhibitor of SARS-CoV-2 programmed ribosomal frameshifting
Programmed ribosomal frameshifting (PRF) occurs in many viruses including SARS-CoV-2 to allow the translation of multiple proteins from a single transcript. Here, the authors identify the human short isoform of the zinc-finger antiviral protein (ZAP-S) as a direct regulator of PRF in SARS-CoV-2 that severely impairs SARS-CoV-2 frameshifting in cells and directly interacts with the SARS-CoV-2 RNA; interfering with the folding of the frameshift RNA element.
- Matthias M. Zimmer
- , Anuja Kibe
- & Neva Caliskan
-
Article
| Open Access RNF219 attenuates global mRNA decay through inhibition of CCR4-NOT complex-mediated deadenylation
RNF219 attenuates global mRNA decay through inhibition of CCR4-NOT complex-mediated deadenylation
The CCR4-NOT complex is a key regulator of eukaryotic mRNA deadenylation. Here the authors identified the E3 ubiquitin ligase RNF219 as an acetylation-regulated cofactor of the complex, which inhibits mRNA decay through a direct interaction with NOT9.
- Fabian Poetz
- , Joshua Corbo
- & Georg Stoecklin
-
Article
| Open Access Structural and molecular basis for Cardiovirus 2A protein as a viral gene expression switch
Structural and molecular basis for Cardiovirus 2A protein as a viral gene expression switch
Many RNA viruses employ programmed –1 ribosomal frameshifting (PRF) to expand their coding capacity and optimize production of viral proteins. Here, the authors report structural and biophysical analysis of protein 2A from a cardiovirus, with insights into the mechanism of its PRF-stimulatory function.
- Chris H. Hill
- , Lukas Pekarek
- & Ian Brierley
-
Article
| Open Access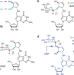 Synthesis and structure elucidation of the human tRNA nucleoside mannosyl-queuosine
Synthesis and structure elucidation of the human tRNA nucleoside mannosyl-queuosine
Mannosyl-queuosine (manQ) is a non-canonical RNA nucleoside present in the anticodon loop of certain tRNAs. Here, the authors use a combination of total synthesis and mass spectrometry to contradict the literature-reported structure and show that manQ features an alpha-allyl connectivity of its mannose moiety.
- Markus Hillmeier
- , Mirko Wagner
- & Thomas Carell
-
Article
| Open Access Atlas of RNA editing events affecting protein expression in aged and Alzheimer’s disease human brain tissue
Atlas of RNA editing events affecting protein expression in aged and Alzheimer’s disease human brain tissue
Resources reporting RNA editing sites from brain tissue have been published. Here, the authors provide an atlas of RNA editing events found in the aged and Alzheimer’s disease human brain tissue resulting in changes at protein level.
- Yiyi Ma
- , Eric B. Dammer
- & Philip L. De Jager
-
Article
| Open Access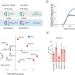 NUDT2 initiates viral RNA degradation by removal of 5′-phosphates
NUDT2 initiates viral RNA degradation by removal of 5′-phosphates
RNA of some viruses is protected from degradation by a 5′ triphosphate group. Here the authors identify nudix hydrolase 2 (NUDT2) as novel antiviral defense protein that dephosphorylates viral RNA and thereby enables its degradation.
- Beatrice T. Laudenbach
- , Karsten Krey
- & Andreas Pichlmair
-
Article
| Open Access MDA5 disease variant M854K prevents ATP-dependent structural discrimination of viral and cellular RNA
MDA5 disease variant M854K prevents ATP-dependent structural discrimination of viral and cellular RNA
MDA5 is the primary immune sensor for SARS-CoV-2 and many other viruses. Mutations in MDA5 can cause disease. Here the authors employ CryoEM and biochemical methods to show how steric constraints cause MDA5 to misrecognize endogenous RNA as viral RNA.
- Qin Yu
- , Alba Herrero del Valle
- & Yorgo Modis
-
Article
| Open Access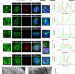 DHX15-independent roles for TFIP11 in U6 snRNA modification, U4/U6.U5 tri-snRNP assembly and pre-mRNA splicing fidelity
DHX15-independent roles for TFIP11 in U6 snRNA modification, U4/U6.U5 tri-snRNP assembly and pre-mRNA splicing fidelity
In yeast, TFIP11 and DHX15 promote the disassembly of spliceosome complex after splicing is completed. Here the authors show that human TFIP11 functions independently of DHX15 and is required for U6 snRNA 2’-O-methylation and U4/U6.U5 tri-snRNP assembly.
- Amandine Duchemin
- , Tina O’Grady
- & Denis Mottet
-
Article
| Open Access Bi-directional ribosome scanning controls the stringency of start codon selection
Bi-directional ribosome scanning controls the stringency of start codon selection
Start codon selection is commonly thought to occur through the unidirectional scanning of the mRNA by the 40 S ribosome. Here the authors provide evidence that the pre-initiation complex can backslide on the mRNA to initiate translation at upstream AUG codons.
- Yifei Gu
- , Yuanhui Mao
- & Shu-Bing Qian
-
Article
| Open Access Pseudoknot length modulates the folding, conformational dynamics, and robustness of Xrn1 resistance of flaviviral xrRNAs
Pseudoknot length modulates the folding, conformational dynamics, and robustness of Xrn1 resistance of flaviviral xrRNAs
Exoribonuclease-resistant RNAs (xrRNAs) are RNA elements that block the exoribonucleolytic degradation of RNA. Here the authors show how a long-range pseudoknot length modulates the Mg2+-dependence of flaviviral xrRNA’s folding, conformational dynamics and Xrn1 resistance.
- Xiaolin Niu
- , Ruirui Sun
- & Xianyang Fang
-
Article
| Open Access Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
Association of snR190 snoRNA chaperone with early pre-60S particles is regulated by the RNA helicase Dbp7 in yeast
The molecular events underlying the assembly and maturation of the early pre-60S particles during eukaryotic ribosome synthesis are not well understood. Here, the authors combine yeast genetics and biochemical experiments to characterise the functions of two important players of eukaryotic ribosome biogenesis, the box C/D snoRNP snR190 and the helicase Dbp7, which both interact. They show that the snR190 snoRNA acts as a RNA chaperone that assists the structuring of the 25S rRNA during the maturation of early pre-60S particles and that Dbp7 is important for facilitating remodeling events in the peptidyl transferase center region of the 25S rRNAs during the maturation of early pre-60S particles.
- Mariam Jaafar
- , Julia Contreras
- & Anthony K. Henras
-
Article
| Open Access Photoactivatable ribonucleosides mark base-specific RNA-binding sites
Photoactivatable ribonucleosides mark base-specific RNA-binding sites
RNA-protein interactions play critical roles in post-transcriptional gene regulation. Here the authors demonstrate pRBS-ID, an updated MS/MS-based method that combines the benefits of photoactivatable ribonucleosides and the chemical cleavage of RNA.
- Jong Woo Bae
- , Sangtae Kim
- & Jong-Seo Kim
-
Article
| Open Access Coupled protein synthesis and ribosome-guided piRNA processing on mRNAs
Coupled protein synthesis and ribosome-guided piRNA processing on mRNAs
Ribosome can mediate piRNA biogenesis from long non-coding RNAs following translation of the short open reading frames. Here the authors show that 80S ribosome also guides piRNA production from 3’ UTR of protein-coding genes after translation of long open reading frames, indicating a general piRNA biogenesis mechanism regardless of their precursor ORF length.
- Yu H. Sun
- , Ruoqiao Huiyi Wang
- & Xin Zhiguo Li
-
Article
| Open Access RN7SK small nuclear RNA controls bidirectional transcription of highly expressed gene pairs in skin
RN7SK small nuclear RNA controls bidirectional transcription of highly expressed gene pairs in skin
The noncoding RNA RN7SK regulates RNA polymerase II pausing and splicing. Here the authors deplete RN7SK in mouse and human during epidermal stem cell differentiation and reveal a novel role in orchestrating bidirectional transcription of highly expressed gene pairs.
- Roberto Bandiera
- , Rebecca E. Wagner
- & Michaela Frye

 Limited effects of m6A modification on mRNA partitioning into stress granules
Limited effects of m6A modification on mRNA partitioning into stress granules