Article
|
Open Access
Featured
-
-
Article
| Open Access LARP7 family proteins have conserved function in telomerase assembly
LARP7 family proteins have conserved function in telomerase assembly
The telomerase holoenzyme is minimally composed of the reverse transcriptase and the RNA template. Here the authors identify Lar7 as a member of the full complex that helps to stabilise it and protect telomerase RNA from degradation.
- Laura C. Collopy
- , Tracy L. Ware
- & Kazunori Tomita
-
Article
| Open Access Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1
Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1
RNA-binding protein hnRNP A2/B1 is suggested to promote miRNA processing as a m6A 'reader'. Here, the authors determine crystal structures of RRM domains of hnRNP A2/B1 in complex with various RNA substrates and determine that hnRNP A2/B1 may function as an auxiliary factor in 'm6A switch' instead.
- Baixing Wu
- , Shichen Su
- & Jinbiao Ma
-
Article
| Open Access TDRD5 binds piRNA precursors and selectively enhances pachytene piRNA processing in mice
TDRD5 binds piRNA precursors and selectively enhances pachytene piRNA processing in mice
Pachytene piRNAs are abundant piRNAs in mammalian adult testes but their biogenesis pathway is not fully understood. Here, the authors identify TDRD5 as a piRNA biogenesis factor in mice, showing that it binds piRNA precursors and promotes pachytene piRNA production from specific transcript regions.
- Deqiang Ding
- , Jiali Liu
- & Chen Chen
-
Article
| Open Access CryoEM structure of Saccharomyces cerevisiae U1 snRNP offers insight into alternative splicing
CryoEM structure of Saccharomyces cerevisiae U1 snRNP offers insight into alternative splicing
U1 snRNP is critical for 5′ splicing site recognition in pre-mRNA splicing. Here the authors describe the cryo-EM structure of the yeast U1 snRNP and suggest that PrpF39 is an alternative splicing factor essential for the successful recruitment of U1 snRNP by other alternative splicing factors.
- Xueni Li
- , Shiheng Liu
- & Rui Zhao
-
Article
| Open Access Aromatic side-chain conformational switch on the surface of the RNA Recognition Motif enables RNA discrimination
Aromatic side-chain conformational switch on the surface of the RNA Recognition Motif enables RNA discrimination
The RNA Recognition Motif (RRM) is the most ubiquitous RNA binding domain. Here the authors combined NMR and molecular dynamics simulations and show that the RRM RNA binding surface exists in different states and that a conformational switch of aromatic side-chains fine-tunes sequence specific binding affinities.
- Nana Diarra dit Konté
- , Miroslav Krepl
- & Frédéric H.-T. Allain
-
Article
| Open Access The E3 ubiquitin ligase and RNA-binding protein ZNF598 orchestrates ribosome quality control of premature polyadenylated mRNAs
The E3 ubiquitin ligase and RNA-binding protein ZNF598 orchestrates ribosome quality control of premature polyadenylated mRNAs
Translation of aberrant mRNAs causes ribosome stalling and translation arrest, followed by recycling of the stalled ribosome complex. Here the authors show that the Zinc Finger Protein 598 (ZNF598/Hel2) is implicated in sensing faulty translation of prematurely polyadenylated mRNAs through the recognition of AAA codons.
- Aitor Garzia
- , Seyed Mehdi Jafarnejad
- & Nahum Sonenberg
-
Article
| Open Access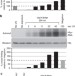 Kinetic CRAC uncovers a role for Nab3 in determining gene expression profiles during stress
Kinetic CRAC uncovers a role for Nab3 in determining gene expression profiles during stress
Protein RNA interactions are dynamic and regulated in response to environmental changes. Here the authors describe ‘kinetic CRAC’, an approach that allows time resolved analyses of protein RNA interactions with minute time point resolution and apply it to gain insight into the function of the RNA-binding protein Nab3.
- Rob van Nues
- , Gabriele Schweikert
- & Sander Granneman
-
Article
| Open Access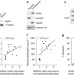 The RNA-binding protein vigilin regulates VLDL secretion through modulation of Apob mRNA translation
The RNA-binding protein vigilin regulates VLDL secretion through modulation of Apob mRNA translation
RNA-binding proteins (RBP) are an emerging group of post-translational regulators. Here the authors show that the RBP vigilin regulates translation of mRNA encoding for proatherogenic proteins—apoB, apoC-III and fibronectin—representing a potential therapeutic target in cardiovascular diseases.
- Mehrpouya B. Mobin
- , Stefanie Gerstberger
- & Markus Stoffel
-
Article
| Open Access YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4–NOT deadenylase complex
YTHDF2 destabilizes m6A-containing RNA through direct recruitment of the CCR4–NOT deadenylase complex
The YTHDF family of proteins are able to bind and regulate the stability of methylated N6 RNA. Here the authors show that this decreased m6A RNA stability is mediated by direct recruitment of the CCR4–NOT deadenylase complex through YTHDF proteins.
- Hao Du
- , Ya Zhao
- & Ligang Wu
-
Article
| Open Access RNA polymerase I–Rrn3 complex at 4.8 Å resolution
RNA polymerase I–Rrn3 complex at 4.8 Å resolution
RNA polymerase I is the central enzyme that synthesizes ribosomal RNA in eukaryotic cells, and its regulation underlies cell growth. Here the authors present a high-resolution cryo-EM structure of the Pol I-Rrn3 complex that explains how Rrn3 specifically recognizes Pol I to form an initiation competent complex.
- Christoph Engel
- , Jürgen Plitzko
- & Patrick Cramer
-
Article
| Open Access A conserved abundant cytoplasmic long noncoding RNA modulates repression by Pumilio proteins in human cells
A conserved abundant cytoplasmic long noncoding RNA modulates repression by Pumilio proteins in human cells
The human genome contains thousands of long noncoding RNAs which have been preserved by evolution, through their functions are poorly described. Here the authors show that NORAD binds the Pumilo homologues PUM1 and PUM2 to regulate mRNA levels of genes involved in chromosome segregation.
- Ailone Tichon
- , Noa Gil
- & Igor Ulitsky
-
Article
| Open Access Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses
Distinct and shared functions of ALS-associated proteins TDP-43, FUS and TAF15 revealed by multisystem analyses
Abnormal functions of RNA-binding proteins TAF15, FUS and TDP43 are associated with amyotrophic lateral sclerosis. Here, Kapeli et al. characterize the RNA targets of TAF15 and identify points of convergence and divergence between the targets of TAF15, FUS and TDP43 in several neuronal systems.
- Katannya Kapeli
- , Gabriel A. Pratt
- & Gene W. Yeo
-
Article
| Open Access Global changes of the RNA-bound proteome during the maternal-to-zygotic transition in Drosophila
Global changes of the RNA-bound proteome during the maternal-to-zygotic transition in Drosophila
Early development is controlled by maternally deposited mRNAs and the RNA-binding proteins (RBPs) that regulate them. Here the authors describe the identification of a large number of RBPs bound to polyadenylated RNAs in Drosophilaembryos before and after the maternal-to-zygotic transition, revealing changes in RBPs activity during development.
- Vasiliy O. Sysoev
- , Bernd Fischer
- & Anne Ephrussi
-
Article
| Open Access A spliceosome intermediate with loosely associated tri-snRNP accumulates in the absence of Prp28 ATPase activity
A spliceosome intermediate with loosely associated tri-snRNP accumulates in the absence of Prp28 ATPase activity
The assembly of the splicesome involves several distinct stages that require the sequential action of DExD/H-box RNA helicases. Here, the authors uncover a new intermediate, the pre-B complex, that accumulates in the presence of an inactive form of the DEAD-box protein Prp28.
- Carsten Boesler
- , Norbert Rigo
- & Reinhard Lührmann
-
Article
| Open Access Structural basis for the recognition of guide RNA and target DNA heteroduplex by Argonaute
Structural basis for the recognition of guide RNA and target DNA heteroduplex by Argonaute
Argonaute proteins are important in the silencing machinery with some regulatory RNAs. Here, the authors solve the structure of an argonaute protein in complex with both the guide RNA and target DNA and propose a mechanism for their recognition.
- Tomohiro Miyoshi
- , Kosuke Ito
- & Toshio Uchiumi
-
Article
| Open Access Loss of the RNA-binding protein TACO1 causes late-onset mitochondrial dysfunction in mice
Loss of the RNA-binding protein TACO1 causes late-onset mitochondrial dysfunction in mice
Mutations in the translational activator of cytochrome c oxidase subunit I (TACO1) causes cytochrome c oxidase deficiency and Leigh Syndrome in patients. Here, the authors characterize mice with a mutation that causes lack of TACO1 expression, identifying a mouse model that could be useful for preclinical trials.
- Tara R. Richman
- , Henrik Spåhr
- & Aleksandra Filipovska
-
Article
| Open Access mTORC1 and CK2 coordinate ternary and eIF4F complex assembly
mTORC1 and CK2 coordinate ternary and eIF4F complex assembly
Ternary complex (TC) and eIF4F complex assembly are rate-limiting steps in translation initiation that are regulated by eIF2α phosphorylation and the mTOR/4E-BP pathway. Here the authors show that the protein kinases mTORC1 and CK2 coordinate TC and eIF4F complex assembly through eIF2β to stimulate cell proliferation.
- Valentina Gandin
- , Laia Masvidal
- & Ivan Topisirovic
-
Article
| Open Access Serial interactome capture of the human cell nucleus
Serial interactome capture of the human cell nucleus
RNA-binding proteins are involved in the posttranscriptional regulation of a large number of cellular processes and several recent studies have sought to describe the extent of the RNA-binding proteome. Here, Conrad et al. describe serIC, a stringent approach they apply towards defining the RNA-binding proteome of the mammalian nucleus.
- Thomas Conrad
- , Anne-Susann Albrecht
- & Ulf Andersson Ørom
-
Article
| Open Access Quaking promotes monocyte differentiation into pro-atherogenic macrophages by controlling pre-mRNA splicing and gene expression
Quaking promotes monocyte differentiation into pro-atherogenic macrophages by controlling pre-mRNA splicing and gene expression
Post-transcriptional control of RNA is important in health and disease. Here, the authors show that the RNA-binding protein Quaking guides pre-mRNA splicing and transcript abundance during monocyte to macrophage differentiation, and that Quaking depletion impairs pro-atherogenic foam cell formation.
- Ruben G. de Bruin
- , Lily Shiue
- & Eric P. van der Veer
-
Article
| Open Access Roquin recognizes a non-canonical hexaloop structure in the 3′-UTR of Ox40
Roquin recognizes a non-canonical hexaloop structure in the 3′-UTR of Ox40
Roquin is an RNA-binding protein that prevents autoimmunity by limiting expression of receptors such as Ox40. Here, the authors identify an RNA structure that they describe as an alternative decay element, and they characterise its interaction with Roquin using structural and biochemical techniques.
- Robert Janowski
- , Gitta A. Heinz
- & Michael Sattler
-
Article
| Open Access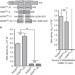 An extended U2AF65–RNA-binding domain recognizes the 3′ splice site signal
An extended U2AF65–RNA-binding domain recognizes the 3′ splice site signal
The pre-mRNA splicing factor U2AF65 recognizes 3′ splice sites in human gene transcripts, but the details are not fully understood. Here, the authors report U2AF65 structures and single molecule FRET that reveal mechanistic insights into splice site recognition.
- Anant A. Agrawal
- , Enea Salsi
- & Clara L. Kielkopf
-
Article
| Open Access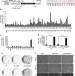 The acetyllysine reader BRD3R promotes human nuclear reprogramming and regulates mitosis
The acetyllysine reader BRD3R promotes human nuclear reprogramming and regulates mitosis
The reprogramming of fibroblasts to pluripotent stem cells has been well documented but there is interest in identifying additional factors involved. Here, the authors perform a screen of human kinases and show that the bromodomain protein, BRD3R, can promote reprogramming and suggest a role for this factor in regulating mitosis.
- Zhicheng Shao
- , Ruowen Zhang
- & Kejin Hu
-
Article
| Open Access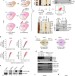 The RNA-binding proteomes from yeast to man harbour conserved enigmRBPs
The RNA-binding proteomes from yeast to man harbour conserved enigmRBPs
RNA-binding proteins (RBPs) are implicated in many biological functions. Here the authors expand the human and yeast RNA interactome identifying new and conserved RBPs, several of which with no prior function assigned to RNA biology or structural motifs known to mediate RNA-binding, and suggesting new roles of RNA as modulators of protein function.
- Benedikt M. Beckmann
- , Rastislav Horos
- & Matthias W. Hentze
-
Article
| Open Access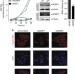 mRNA export through an additional cap-binding complex consisting of NCBP1 and NCBP3
mRNA export through an additional cap-binding complex consisting of NCBP1 and NCBP3
The processing of RNAs transcribed by RNA polymerase II requires a cap-binding complex (CBC), consisting of NCBP1 and NCBP2. Here, the authors report an alternative CBC formed by NCBP1 and a previously uncharacterized protein, NCBP3 that is critical for RNA processing under cellular stress conditions.
- Anna Gebhardt
- , Matthias Habjan
- & Andreas Pichlmair
-
Article
| Open Access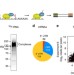 RNA regulatory networks diversified through curvature of the PUF protein scaffold
RNA regulatory networks diversified through curvature of the PUF protein scaffold
Members of the PUF family of RNA-binding proteins bind multiple mRNAs in vivo. Here the authors show that the S. cerevisiaePuf5p binds targets of varying lengths that correlate with biological functions. The RNA-binding sites adopt different structures to adapt to a fixed protein scaffold.
- Daniel Wilinski
- , Chen Qiu
- & Marvin Wickens
-
Article
| Open Access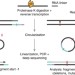 Improved binding site assignment by high-resolution mapping of RNA–protein interactions using iCLIP
Improved binding site assignment by high-resolution mapping of RNA–protein interactions using iCLIP
Individual-nucleotide resolution crosslinking and immunoprecipitation (iCLIP) can map RNA binding sites of RNA-binding proteins (RBPs). Here, the authors report an analysis tool that improves the binding site assignment for some RBPs that have length-dependent broader distribution for their iCLIP fragments.
- Christian Hauer
- , Tomaz Curk
- & Andreas E. Kulozik
-
Article
| Open Access RC3H1 post-transcriptionally regulates A20 mRNA and modulates the activity of the IKK/NF-κB pathway
RC3H1 post-transcriptionally regulates A20 mRNA and modulates the activity of the IKK/NF-κB pathway
The RNA-binding protein RC3H1/ROQUIN1 promotes the degradation of mRNA by binding to a consensus CDE present in the 3′UTR. Here the authors expand the set of consensus sequences through which RCH31 binds and regulates mRNA encoding members of the DNA damage response and IKK/NF-κB pathway.
- Yasuhiro Murakawa
- , Michael Hinz
- & Markus Landthaler
-
Article
| Open Access Telomeric G-quadruplexes are a substrate and site of localization for human telomerase
Telomeric G-quadruplexes are a substrate and site of localization for human telomerase
G-quadruplexes formed by four guanine bases in a square planar arrangement in telomeres may prevent extension of this region by telomerase. Here, the authors show that telomerase can localize to and partially unwind and extend G-quadruplexes, suggesting an important biological role for G-quadruplexes.
- Aaron L. Moye
- , Karina C. Porter
- & Tracy M. Bryan
-
Article |
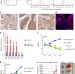 Transformation of the intestinal epithelium by the MSI2 RNA-binding protein
Transformation of the intestinal epithelium by the MSI2 RNA-binding protein
In mammals there are two Musashi proteins, MSI1 and MSI2, orthologues of the Drosophila protein, with roles in asymmetric stem cell division and cell fate determination. Here the authors report new functions for MSI2 in colorectal cancer using in vitro loss of function and in vivoectopic overexpression.
- Shan Wang
- , Ning Li
- & Christopher J. Lengner
-
Article
| Open Access Roquin binds microRNA-146a and Argonaute2 to regulate microRNA homeostasis
Roquin binds microRNA-146a and Argonaute2 to regulate microRNA homeostasis
Roquin is an RNA-binding protein that promotes the degradation of specific mRNAs and is crucial for the maintenance of peripheral immune tolerance. Here the authors show that, in addition to its target mRNAs, Roquin can bind miR-146a and the RISC component Ago2 to control homeostasis of both RNA species.
- Monika Srivastava
- , Guowen Duan
- & Carola G. Vinuesa
-
Article |
 ALS-causative mutations in FUS/TLS confer gain and loss of function by altered association with SMN and U1-snRNP
ALS-causative mutations in FUS/TLS confer gain and loss of function by altered association with SMN and U1-snRNP
Dominant mutations in the RNA-binding protein FUS/TLS cause amyotrophic lateral sclerosis (ALS), an adult-onset motor neuron degenerative disease. Here, the authors show that ALS-causative FUS/TLS mutations directly bind the SMN and U1-snRNP complexes, producing both loss and gain of function effects on RNA processing.
- Shuying Sun
- , Shuo-Chien Ling
- & Don W. Cleveland
-
Article |
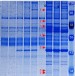 RNA-binding proteins regulate the expression of the immune activating ligand MICB
RNA-binding proteins regulate the expression of the immune activating ligand MICB
The expression of stress-induced ligands and their recognition by the NKG2D-activating receptor is important for the elimination of virally infected and cancerous cells by cytotoxic lymphocytes. Here, the authors provide insights into the post-transcriptional mechanism regulating the expression of the NKG2D ligand, MICB.
- Daphna Nachmani
- , Tony Gutschner
- & Ofer Mandelboim
-
Article
| Open Access Three-dimensional analysis of ribonucleoprotein complexes in influenza A virus
Three-dimensional analysis of ribonucleoprotein complexes in influenza A virus
The influenza A virus genome consists of eight RNA segments, which permits genetic reassortment and contributes to the emergence of novel strains with pandemic potential. Here, electron tomography is used to study the three-dimensional structure of ribonucleoprotein complexes within progeny virions.
- Takeshi Noda
- , Yukihiko Sugita
- & Yoshihiro Kawaoka
-
Article
| Open Access Selective inhibition of microRNA accessibility by RBM38 is required for p53 activity
Selective inhibition of microRNA accessibility by RBM38 is required for p53 activity
MicroRNAs bind to the 3′-untranslated region of genes to regulate expression. In this study, an RNA-binding protein, RMB38, is shown to selectively regulate the access of some microRNAs to their targets, and control the expression of some p53 target genes.
- Nicolas Léveillé
- , Ran Elkon
- & Reuven Agami
-
Article |
 TERRA transcripts are bound by a complex array of RNA-binding proteins
TERRA transcripts are bound by a complex array of RNA-binding proteins
Recent work has revealed that the TTAGGG DNA repeats of telomeres are transcribed to form 'TERRA'. In this study, a set of RNA-binding proteins are shown to bind TERRA transcripts, altering the location of these transcripts at telomeres and regulating telomere abundance and length.
- Isabel López de Silanes
- , Martina Stagno d'Alcontres
- & Maria A Blasco

 Pof8 is a La-related protein and a constitutive component of telomerase in fission yeast
Pof8 is a La-related protein and a constitutive component of telomerase in fission yeast