Featured
-
-
Article
| Open Access Structure-based prediction and characterization of photo-crosslinking in native protein–RNA complexes
Structure-based prediction and characterization of photo-crosslinking in native protein–RNA complexes
Feng et al. developed a computational method PxR3D-map to jointly analyze crosslinked nucleotides and amino acids in protein-RNA complexes, which revealed key structural features underlying photocrosslinking of protein and RNA in cells.
- Huijuan Feng
- , Xiang-Jun Lu
- & Chaolin Zhang
-
Article
| Open Access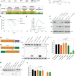 A TRIM21-based bioPROTAC highlights the therapeutic benefit of HuR degradation
A TRIM21-based bioPROTAC highlights the therapeutic benefit of HuR degradation
Overexpression of human antigen R (HuR) correlates with high grade tumours and poor patient prognosis. Here, the authors engineer a TRIM21 biological PROTAC to demonstrate the benefit of a targeted protein degradation approach to deplete HuR, resulting in tumour growth inhibition in pre-clinical cancer models by altering the HuR-regulated proteome.
- Alice Fletcher
- , Dean Clift
- & James Hunt
-
Article
| Open Access Dynamic characterization and interpretation for protein-RNA interactions across diverse cellular conditions using HDRNet
Dynamic characterization and interpretation for protein-RNA interactions across diverse cellular conditions using HDRNet
Predicting dynamic RNA-RBP interactions in diverse cell lines is an important challenge in unravelling RNA function and post-transcriptional regulatory mechanisms. Here, authors develop HDRNet, an end-to-end deep-learning-based framework for accurately predicting dynamic RBP binding events across various cellular conditions.
- Haoran Zhu
- , Yuning Yang
- & Xiangtao Li
-
Article
| Open Access Integrative solution structure of PTBP1-IRES complex reveals strong compaction and ordering with residual conformational flexibility
Integrative solution structure of PTBP1-IRES complex reveals strong compaction and ordering with residual conformational flexibility
An integrated structural biology approach is utilized to elucidate the solution structure of the polypyrimidine-tract binding protein 1 (PTBP1/hnRNP I) complexed with an internal ribosome entry site (IRES) RNA fragment from encephalomyocarditis virus (EMCV).
- Georg Dorn
- , Christoph Gmeiner
- & Frédéric H.-T. Allain
-
Article
| Open Access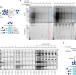 Signal-noise metrics for RNA binding protein identification reveal broad spectrum protein-RNA interaction frequencies and dynamics
Signal-noise metrics for RNA binding protein identification reveal broad spectrum protein-RNA interaction frequencies and dynamics
The identification of RNA-bound proteomes is hampered by a lack of quantitative metrics for evaluating RNA binding function. Here, the authors report LEAP-RBP as a method for purification of RNA-bound proteins and introduce signal-based metrics for robust profiling of RNA-bound proteomes.
- JohnCarlo Kristofich
- & Christopher V. Nicchitta
-
Article
| Open Access Characterising the RNA-binding protein atlas of the mammalian brain uncovers RBM5 misregulation in mouse models of Huntington’s disease
Characterising the RNA-binding protein atlas of the mammalian brain uncovers RBM5 misregulation in mouse models of Huntington’s disease
RNA-Binding Proteins (RBPs) are critical regulators of RNA biology. Here, the authors describe the Brain-pCLAP methodology, uncover the RBP atlas of the mouse brain and demonstrate the differential binding of the splicing factor RBM5 to Huntington’s disease relevant transcripts in R6/2 mice.
- Meeli Mullari
- , Nicolas Fossat
- & Michael L. Nielsen
-
Article
| Open Access Cytosolic Ptbp2 modulates axon growth in motoneurons through axonal localization and translation of Hnrnpr
Cytosolic Ptbp2 modulates axon growth in motoneurons through axonal localization and translation of Hnrnpr
The neuronal RNA-binding protein Ptbp2 is known to regulate neuronal differentiation by modulating alternative splicing. Here, the authors reveal an additional role of cytosolic Ptbp2, which regulates axon growth by fine-tuning the mRNA transport and local synthesis of an RNA-binding protein hnRNP R.
- Saeede Salehi
- , Abdolhossein Zare
- & Michael Sendtner
-
Article
| Open Access The preference signature of the SARS-CoV-2 Nucleocapsid NTD for its 5’-genomic RNA elements
The preference signature of the SARS-CoV-2 Nucleocapsid NTD for its 5’-genomic RNA elements
SARS-CoV-2 genome turnover is mediated by its N protein, but precise parameters driving the necessary RNA specificity have remained enigmatic. Here, Korn et al. reveal N’s N-terminal domain to distinguish regulatory viral RNA motifs with a preference for transiently folded elements of functional impact.
- Sophie Marianne Korn
- , Karthikeyan Dhamotharan
- & Andreas Schlundt
-
Article
| Open Access PIE-seq: identifying RNA-binding protein targets by dual RNA-deaminase editing and sequencing
PIE-seq: identifying RNA-binding protein targets by dual RNA-deaminase editing and sequencing
Tracking protein-RNA interaction across cell types is challenging. Here, Ruan et al develop a dual-deaminase method called PIE-Seq, where protein targets are marked by both C-to-U and A-to-I RNA base editors, and apply it to 25 human RNA-binding proteins.
- Xiangbin Ruan
- , Kaining Hu
- & Xiaochang Zhang
-
Article
| Open Access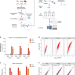 The RNA-binding protein landscapes differ between mammalian organs and cultured cells
The RNA-binding protein landscapes differ between mammalian organs and cultured cells
Characterization of RNA-binding proteins (RBPs) in tissues has been hampered by technical constraints. Here, the authors describe ex vivo eRIC, a method for global profiling of RBPs active in mammalian organs, and report comprehensive RBP atlases from mouse brain, kidney and liver.
- Joel I. Perez-Perri
- , Dunja Ferring-Appel
- & Matthias W. Hentze
-
Article
| Open Access Structural basis of sequence-specific RNA recognition by the antiviral factor APOBEC3G
Structural basis of sequence-specific RNA recognition by the antiviral factor APOBEC3G
Interaction between APOBEC3G and RNA is critical for its antiviral function. Here, the authors report four co-crystal structures of rhesus macaque APOBEC3G and RNA, demonstrating unpaired AA and GA dinucleotide motifs are preferentially recognized.
- Hanjing Yang
- , Kyumin Kim
- & Xiaojiang S. Chen
-
Article
| Open Access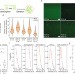 Formation of synthetic RNA protein granules using engineered phage-coat-protein -RNA complexes
Formation of synthetic RNA protein granules using engineered phage-coat-protein -RNA complexes
Condensates composed of RNA and proteins are biologically vital but generally poorly understood. Here, the authors engineer synthetic RNA-protein condensates and show that gel-like condensates form as a result of liquid-gel phase separation in a specific and selective fashion.
- Naor Granik
- , Noa Katz
- & Roee Amit
-
Article
| Open Access The solution structure of Dead End bound to AU-rich RNA reveals an unusual mode of tandem RRM-RNA recognition required for mRNA regulation
The solution structure of Dead End bound to AU-rich RNA reveals an unusual mode of tandem RRM-RNA recognition required for mRNA regulation
The authors report an unusual mode of AU-rich RNA recognition by the RNA recognition motifs of DND1, a protein essential for germline development, in a 27.5 kDa NMR structure and provide additional insight on DND1 function from cell-based experiments.
- Malgorzata M. Duszczyk
- , Harry Wischnewski
- & Frédéric H.-T. Allain
-
Article
| Open Access The splicing factor RBM17 drives leukemic stem cell maintenance by evading nonsense-mediated decay of pro-leukemic factors
The splicing factor RBM17 drives leukemic stem cell maintenance by evading nonsense-mediated decay of pro-leukemic factors
Leukemic stem cells (LSCs) drive chemoresistance and relapse in acute myeloid leukemia. Here, the authors show that the splicing factor RBM17 supports LSCs through avoiding nonsense-mediated decay of pro-leukaemic factors such as the translation initiation factor EIF4A2.
- Lina Liu
- , Ana Vujovic
- & Yu Lu
-
Article
| Open Access Probing TDP-43 condensation using an in silico designed aptamer
Probing TDP-43 condensation using an in silico designed aptamer
Here, the authors generated an artificial RNA molecule, or aptamer, specific for the Amyotrophic Lateral Sclerosis protein TDP-43. By interacting avidly with its target, the aptamer can be exploited to track TDP-43 phase transition in vitro and in cells.
- Elsa Zacco
- , Owen Kantelberg
- & Gian Gaetano Tartaglia
-
Article
| Open Access Structural basis of template strand deoxyuridine promoter recognition by a viral RNA polymerase
Structural basis of template strand deoxyuridine promoter recognition by a viral RNA polymerase
Promoter recognition is a critical step in the initiation of transcription of DNA to RNA. Here, the authors describe a novel mechanism by which a phage-encoded RNA polymerase recognizes viral promoters containing deoxyuridines instead of thymidines.
- Alec Fraser
- , Maria L. Sokolova
- & Petr G. Leiman
-
Article
| Open Access RNA supply drives physiological granule assembly in neurons
RNA supply drives physiological granule assembly in neurons
RNA granules are important regulators of RNA metabolism. Here the authors report that RNA granules containing RNA helicase DDX6 disassemble during neuronal maturation.
- Karl E. Bauer
- , Niklas Bargenda
- & Michael A. Kiebler
-
Article
| Open Access Nucleotide-amino acid π-stacking interactions initiate photo cross-linking in RNA-protein complexes
Nucleotide-amino acid π-stacking interactions initiate photo cross-linking in RNA-protein complexes
Although UV-induced cross-linking is a widely used method to study RNA-protein complexes, the cross-linking reactions are poorly understood. Here, the authors show that π-stacking interactions between nucleobases and aromatic amino acids play a key role in the cross-linking process.
- Anna Knörlein
- , Chris P. Sarnowski
- & Jonathan Hall
-
Article
| Open Access A cryptic pocket in Ebola VP35 allosterically controls RNA binding
A cryptic pocket in Ebola VP35 allosterically controls RNA binding
Many viral proteins are thought to be unlikely candidates for drug discovery as they lack obvious drug binding sites. Here, the authors use computational approaches followed by experimental validation to identify a cryptic pocket within the Ebola virus protein VP35.
- Matthew A. Cruz
- , Thomas E. Frederick
- & Gregory R. Bowman
-
Article
| Open Access Photoactivatable ribonucleosides mark base-specific RNA-binding sites
Photoactivatable ribonucleosides mark base-specific RNA-binding sites
RNA-protein interactions play critical roles in post-transcriptional gene regulation. Here the authors demonstrate pRBS-ID, an updated MS/MS-based method that combines the benefits of photoactivatable ribonucleosides and the chemical cleavage of RNA.
- Jong Woo Bae
- , Sangtae Kim
- & Jong-Seo Kim
-
Article
| Open Access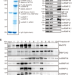 Rett syndrome linked to defects in forming the MeCP2/Rbfox/LASR complex in mouse models
Rett syndrome linked to defects in forming the MeCP2/Rbfox/LASR complex in mouse models
MeCP2 mutations can cause Rett syndrome, a severe childhood neurological disorder. Here the authors show that MeCP2 mediates the higher-order assembly of a large splicing complex Rbfox/LASR, which is disrupted in the mouse models of Rett syndrome.
- Yan Jiang
- , Xing Fu
- & Jingyi Hui
-
Article
| Open Access Assembly defects of human tRNA splicing endonuclease contribute to impaired pre-tRNA processing in pontocerebellar hypoplasia
Assembly defects of human tRNA splicing endonuclease contribute to impaired pre-tRNA processing in pontocerebellar hypoplasia
Mutations within subunits of the tRNA splicing endonuclease complex (TSEN) are associated with pontocerebellar hypoplasia (PCH). Here the authors show that tRNA intron excision is catalyzed by tetrameric TSEN assembled from inactive heterodimers, and provide evidence that modulation of TSEN stability may contribute to PCH phenotypes.
- Samoil Sekulovski
- , Pascal Devant
- & Simon Trowitzsch
-
Article
| Open Access Defining the RBPome of primary T helper cells to elucidate higher-order Roquin-mediated mRNA regulation
Defining the RBPome of primary T helper cells to elucidate higher-order Roquin-mediated mRNA regulation
An extensive RNA binding protein atlas (RBPome) for primary T cells would be a useful resource. Here the authors use two different methods to characterise the mouse and human T cell RBPome and show regulation of Roquin-1/2 dependent and independent pathways.
- Kai P. Hoefig
- , Alexander Reim
- & Vigo Heissmeyer
-
Article
| Open Access Profiling PRMT methylome reveals roles of hnRNPA1 arginine methylation in RNA splicing and cell growth
Profiling PRMT methylome reveals roles of hnRNPA1 arginine methylation in RNA splicing and cell growth
Arginine methyltransferases (PRMTs) are involved in the regulation of various physiological and pathological conditions. Using proteomics, the authors here profile the methylation substrates of PRMTs 4, 5 and 7 and characterize the roles of these enzymes in cancer-associated splicing regulation.
- Wen-juan Li
- , Yao-hui He
- & Wen Liu
-
Article
| Open Access Towards plant resistance to viruses using protein-only RNase P
Towards plant resistance to viruses using protein-only RNase P
New approaches to plant disease control are important for pathogens that are difficult to control by existing methods. Here, the authors report a potential strategy to combat plant viruses by cytosolic expressed protein-only RNase P and show its ability for in vitro cleavage of tRNA-like structures existing in many plant viruses.
- Anthony Gobert
- , Yifat Quan
- & Philippe Giegé
-
Article
| Open Access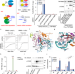 A scaffold lncRNA shapes the mitosis to meiosis switch
A scaffold lncRNA shapes the mitosis to meiosis switch
In fission yeast, the antagonistic RNA-binding proteins Mmi1 and Mei2 respectively promote and inhibit meiotic mRNA degradation during mitotic growth. Here the authors show that the lncRNA mamRNA scaffolds Mmi1 and Mei2 proteins to enable their mutual controls.
- Vedrana Andric
- , Alicia Nevers
- & Mathieu Rougemaille
-
Article
| Open Access Native mass spectrometry reveals the initial binding events of HIV-1 rev to RRE stem II RNA
Native mass spectrometry reveals the initial binding events of HIV-1 rev to RRE stem II RNA
The HIV-1 RNA-binding protein rev facilitates nuclear export of viral RNA. Here, the authors use native mass spectrometry to study the interactions between rev-derived peptides and rev response elements of HIV-1 RNA, providing mechanistic insights into rev recognition and recruitment.
- Eva-Maria Schneeberger
- , Matthias Halper
- & Kathrin Breuker
-
Article
| Open Access Identification of phenothiazine derivatives as UHM-binding inhibitors of early spliceosome assembly
Identification of phenothiazine derivatives as UHM-binding inhibitors of early spliceosome assembly
So far only a few compounds have been reported as splicing modulators. Here, the authors combine high-throughput screening, chemical synthesis, NMR, X-ray crystallography with functional studies and develop phenothiazines as inhibitors for the U2AF Homology Motif (UHM) domains of proteins that regulate splicing and show that they inhibit early spliceosome assembly on pre-mRNA substrates in vitro.
- Pravin Kumar Ankush Jagtap
- , Tomáš Kubelka
- & Michael Sattler
-
Article
| Open Access High-throughput mutagenesis reveals unique structural features of human ADAR1
High-throughput mutagenesis reveals unique structural features of human ADAR1
Human ADAR proteins are responsible for RNA editing, conversion of adenosine to inosine in double-stranded RNA. Here the authors report a previously unknown zinc ion-binding site in the catalytic domain of human ADAR1 using high throughput mutagenesis, biochemical assay and Rosetta-based protein structure modeling.
- SeHee Park
- , Erin E. Doherty
- & Peter A. Beal
-
Article
| Open Access CryoEM structure of the low-complexity domain of hnRNPA2 and its conversion to pathogenic amyloid
CryoEM structure of the low-complexity domain of hnRNPA2 and its conversion to pathogenic amyloid
hnRNPA2 is involved in RNA metabolism and can form both functional amyloid-like fibrils in membraneless organelles, and pathogenic fibrils in neurodegenerative conditions. Here, the authors present the cryo-EM fibril structure of the wild-type hnRNPA2 low-complexity domain (LCD) and the crystal structure of a LCD segment with the disease causing D290V variant and discuss how mutations can transform fibril structure from a functional to a pathogenic form.
- Jiahui Lu
- , Qin Cao
- & David S. Eisenberg
-
Article
| Open Access hnRNP H/F drive RNA G-quadruplex-mediated translation linked to genomic instability and therapy resistance in glioblastoma
hnRNP H/F drive RNA G-quadruplex-mediated translation linked to genomic instability and therapy resistance in glioblastoma
RNA G-quadruplexes (RG4s) have been functionally linked to cancer gene expression. Here, Herviou, Le Bras et al. have identified the protein machinery modulating RG4s and reveal the role and mechanism of hnRNP H/F and DHX36 in RG4-mediated translational regulation affecting cancer treatment in glioblastoma.
- Pauline Herviou
- , Morgane Le Bras
- & Stefania Millevoi
-
Article
| Open Access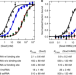 The Sox2 transcription factor binds RNA
The Sox2 transcription factor binds RNA
Some transcription factors have been proposed to functionally interact with RNA to facilitate proper regulation of gene expression. Here the authors demonstrate that human Sox2 interact directly and with high affinity to RNAs through its HMG DNA-binding domain.
- Zachariah E. Holmes
- , Desmond J. Hamilton
- & Robert T. Batey
-
Article
| Open Access Insights into the assembly and architecture of a Staufen-mediated mRNA decay (SMD)-competent mRNP
Insights into the assembly and architecture of a Staufen-mediated mRNA decay (SMD)-competent mRNP
The Staufen proteins recognize secondary structures in 3’-untranslated regions in mRNA transcripts and induce degradation of these mRNAs with the help of the RNA helicase UPF1. Here the authors report that the nonsense-mediated mRNA decay factor UPF2 mediates the interaction between Stau1 and UPF1 in Staufen-mediated mRNA decay.
- Manjeera Gowravaram
- , Juliane Schwarz
- & Sutapa Chakrabarti
-
Article
| Open Access The structural basis for RNA selectivity by the IMP family of RNA-binding proteins
The structural basis for RNA selectivity by the IMP family of RNA-binding proteins
ZBP1 and IMP2 belong to the IGF2BP family of RNA-binding proteins. Here the authors employed SELEX, NMR spectroscopy and mutagenesis to characterize the RNA-binding preference of ZBP1 and IMP2.
- Jeetayu Biswas
- , Vivek L. Patel
- & Carolina Eliscovich
-
Article
| Open Access Lysine/RNA-interactions drive and regulate biomolecular condensation
Lysine/RNA-interactions drive and regulate biomolecular condensation
Processing bodies (P-bodies) are non-membrane-bound protein/RNA granules in the cytosol. Here the authors combine bioinformatics, NMR and cell based assays and find that lysine is enriched in the disordered regions of P-body-associated proteins and show that lysine-rich polypeptides form highly dynamic lysine/RNA-coacervates and lysine acetylation reverses liquid-liquid phase separation.
- Tina Ukmar-Godec
- , Saskia Hutten
- & Markus Zweckstetter
-
Article
| Open Access CAPRI enables comparison of evolutionarily conserved RNA interacting regions
CAPRI enables comparison of evolutionarily conserved RNA interacting regions
Comprehensive characterisation of RNA-protein interactions requires different levels of resolution. Here, the authors present an integrated mass spectrometry-based approach that allows them to define the Drosophila RNA-protein interactome from the level of multisubunit complexes down to the RNA-binding amino acid.
- Amol Panhale
- , Florian M. Richter
- & Asifa Akhtar
-
Article
| Open Access Structural basis for reversible amyloids of hnRNPA1 elucidates their role in stress granule assembly
Structural basis for reversible amyloids of hnRNPA1 elucidates their role in stress granule assembly
Low complexity (LC) domains can drive the formation of both amyloid fibrils and protein droplets. Here, the authors identify reversible amyloid cores from the LC of hnRNPA1, based on which they elucidate the structural basis of reversible fibrillation and its interplay with hnRNPA1 droplet formation.
- Xinrui Gui
- , Feng Luo
- & Dan Li
-
Article
| Open Access Staufen2-mediated RNA recognition and localization requires combinatorial action of multiple domains
Staufen2-mediated RNA recognition and localization requires combinatorial action of multiple domains
The Staufen family of RNA-binding proteins are conserved microtubule dependent mRNA transporter factors. Here the authors use biochemical and functional approaches to characterize the RNA-binding properties of mouse Staufen 2 and study the mRNA binding capacity of its two domains dsRBDs 1 and 2.
- Simone Heber
- , Imre Gáspár
- & Dierk Niessing
-
Article
| Open Access Purification of cross-linked RNA-protein complexes by phenol-toluol extraction
Purification of cross-linked RNA-protein complexes by phenol-toluol extraction
RNA binding proteins are important regulators of RNA function. Here the authors describe a method for isolation of RNA-protein complexes that does not rely on a specific RNA sequence or motif, and demonstrate the approach by providing the global RNA-bound proteomes of human HEK293 cells and Salmonella Typhimurium.
- Erika C. Urdaneta
- , Carlos H. Vieira-Vieira
- & Benedikt M. Beckmann
-
Article
| Open Access Molecular interactions between Hel2 and RNA supporting ribosome-associated quality control
Molecular interactions between Hel2 and RNA supporting ribosome-associated quality control
Ribosome-associated quality control (RQC) pathways monitor and respond to stalling of the translating ribosome. Here the authors show that the ribosome associated RQC factor Hel2/ZNF598, an E3 ubiquitin ligase, generally interacts with mRNAs in the vicinity of stop codons.
- Marie-Luise Winz
- , Lauri Peil
- & David Tollervey
-
Article
| Open Access LOTUS domain protein MARF1 binds CCR4-NOT deadenylase complex to post-transcriptionally regulate gene expression in oocytes
LOTUS domain protein MARF1 binds CCR4-NOT deadenylase complex to post-transcriptionally regulate gene expression in oocytes
The RNA-binding protein MARF1 is required for post-transcriptional regulation of mRNAs during mouse oogenesis. Here, by analyzing a Drosophila MARF1 mutant, the authors show that MARF1 recruits CCR4-NOT deadenylase to shorten the poly-A tails of target mRNAs such as cyclin A and suppress their translation during Drosophila oogenesis.
- Li Zhu
- , Suresh K. Kandasamy
- & Ryuya Fukunaga
-
Article
| Open Access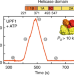 UPF1-like helicase grip on nucleic acids dictates processivity
UPF1-like helicase grip on nucleic acids dictates processivity
UPF1 is a highly processive helicase that plays an essential role in nonsense-mediated mRNA decay. Here the authors use single molecule binding assays to establish a functionally important relationship between helicase grip to nucleic acids, binding lifetime and the duration of translocation.
- Joanne Kanaan
- , Saurabh Raj
- & Hervé Le Hir
-
Article
| Open Access Molecular basis for disassembly of an importin:ribosomal protein complex by the escortin Tsr2
Molecular basis for disassembly of an importin:ribosomal protein complex by the escortin Tsr2
Ribosomal proteins are transported to the nucleus with the help of importins, from which they are released prior to incorporation into the nascent ribosome. Here the authors report the NMR structure of the ribosomal protein eS26 in complex with the escortin Tsr2 and shed light on the mechanism of eS26 release from importin.
- Sabina Schütz
- , Erich Michel
- & Vikram Govind Panse
-
Article
| Open Access Global pairwise RNA interaction landscapes reveal core features of protein recognition
Global pairwise RNA interaction landscapes reveal core features of protein recognition
RNA–protein interactions often depend on the recognition of extended RNA elements but the identification of these motifs is challenging. Here, the authors present a global integrated approach to analyze RNA–protein binding landscapes, mapping extended RNA interaction motifs for four RNA-binding proteins.
- Qin Zhou
- , Nikesh Kunder
- & Zachary T. Campbell
-
Article
| Open Access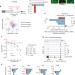 Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
The function and consequences of alternative polyadenylation (APA) in stressed cells are largely unclear. Here, the authors show that stress-induced mRNA degradation depends on 3′UTR length and that APA-mediated 3′UTR shortening is an adaptive stress response mechanism for selective transcript stabilization.
- Dinghai Zheng
- , Ruijia Wang
- & Bin Tian
-
Article
| Open Access HuR regulates telomerase activity through TERC methylation
HuR regulates telomerase activity through TERC methylation
Mutations in the RNA component TERC can cause telomerase dysfunction but the underlying mechanisms are largely unknown. Here, the authors show that RNA-binding protein HuR regulates telomerase function by enhancing the methylation of TERC, which is impaired by several disease-relevant TERC mutations.
- Hao Tang
- , Hu Wang
- & Wengong Wang
-
Article
| Open Access Molecular principles underlying dual RNA specificity in the Drosophila SNF protein
Molecular principles underlying dual RNA specificity in the Drosophila SNF protein
It remains poorly understood how a single RNA-binding protein recognizes diverse RNA targets. Here the authors use an integrative approach to study the binding of spliceosomal SNF protein to U1 and U2 small nuclear RNAs in the presence or absence of auxiliary protein U2A’ and show how SNF’s conformational dynamics are tuned to recognize different stem-loop structures.
- Gert Weber
- , Gregory T. DeKoster
- & Markus C. Wahl
-
Article
| Open Access Structural analysis of human ARS2 as a platform for co-transcriptional RNA sorting
Structural analysis of human ARS2 as a platform for co-transcriptional RNA sorting
Arsenic resistance protein 2 (ARS2) plays an important role in nuclear RNA metabolism and interacts with the nuclear cap-binding complex (CBC). Here the authors present the human ARS2 structure and identify regions important for its interactions with binding partners supporting that mutually exclusive higher order CBC-ARS2 complexes are formed.
- Wiebke Manuela Schulze
- , Frank Stein
- & Stephen Cusack
-
Article
| Open Access LARP7-like protein Pof8 regulates telomerase assembly and poly(A)+TERRA expression in fission yeast
LARP7-like protein Pof8 regulates telomerase assembly and poly(A)+TERRA expression in fission yeast
A functional telomerase complex requires that the catalytic TERT subunit be assembled with the template RNA TER1. Here the authors show that Pof8, a possible LARP7 family protein, is required for assembly of the telomerase complex, and repression of lncRNA transcripts at telomeres in S. pombe.
- Amanda K. Mennie
- , Bettina A. Moser
- & Toru M. Nakamura

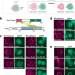 ReLo is a simple and rapid colocalization assay to identify and characterize direct protein–protein interactions
ReLo is a simple and rapid colocalization assay to identify and characterize direct protein–protein interactions