Featured
-
-
Article
| Open Access DIALib-QC an assessment tool for spectral libraries in data-independent acquisition proteomics
DIALib-QC an assessment tool for spectral libraries in data-independent acquisition proteomics
Most data-independent acquisition (DIA) methods depend on mass spectral libraries for peptide identification but tools to assess library quality are lacking. Here, the authors develop DIALib- QC for the systematic evaluation and correction of spectral libraries.
- Mukul K. Midha
- , David S. Campbell
- & Robert L. Moritz
-
Article
| Open Access Integration of absolute multi-omics reveals dynamic protein-to-RNA ratios and metabolic interplay within mixed-domain microbiomes
Integration of absolute multi-omics reveals dynamic protein-to-RNA ratios and metabolic interplay within mixed-domain microbiomes
Here, the authors perform a temporal multi-omic analysis of a minimalistic cellulose-degrading and methane-producing consortium at the strain level and estimate protein-to-RNA ratios and RNA-protein dynamics of the community simultaneously over time.
- F. Delogu
- , B. J. Kunath
- & P. B. Pope
-
Article
| Open Access Identification of modified peptides using localization-aware open search
Identification of modified peptides using localization-aware open search
Mass spectrometry-based proteomics is the method of choice for the global mapping of post-translational modifications, but matching and scoring peaks with unknown masses remains challenging. Here, the authors present a refined open search strategy to score all peaks with higher sensitivity and accuracy.
- Fengchao Yu
- , Guo Ci Teo
- & Alexey I. Nesvizhskii
-
Article
| Open Access Strategies to enable large-scale proteomics for reproducible research
Strategies to enable large-scale proteomics for reproducible research
Clinical proteomics critically depends on the ability to acquire highly reproducible data over an extended period of time. Here, the authors assess reproducibility over four months across different mass spectrometers and develop a computational approach to mitigate variation among instruments over time.
- Rebecca C. Poulos
- , Peter G. Hains
- & Qing Zhong
-
Article
| Open Access Proteome activity landscapes of tumor cell lines determine drug responses
Proteome activity landscapes of tumor cell lines determine drug responses
Proteome activity has a major role in cancer progression and response to drugs. Here, the authors use comprehensive proteomic and phosphoproteomic data, in conjunction with drug-sensitivity screens, to generate a community resource consisting of landscapes of pathway and kinase activity across different cell lines
- Martin Frejno
- , Chen Meng
- & Bernhard Kuster
-
Article
| Open Access Focus on the spectra that matter by clustering of quantification data in shotgun proteomics
Focus on the spectra that matter by clustering of quantification data in shotgun proteomics
Matching mass spectra to peptide sequences is the usual first step in proteomics data analysis, often followed by peptide quantification. Here, the authors show that clustering and quantifying mass spectral features prior to peptide identification can increase the sensitivity of label-free quantitative proteomics.
- Matthew The
- & Lukas Käll
-
Article
| Open Access The Archaeal Proteome Project advances knowledge about archaeal cell biology through comprehensive proteomics
The Archaeal Proteome Project advances knowledge about archaeal cell biology through comprehensive proteomics
While archaeal proteomics advanced rapidly, a comprehensive proteome database for archaea is lacking. Therefore, the authors here launch the Archaeal Proteome Project, a community-effort providing insights into archaeal cell biology via the combined reanalysis of Haloferax volcanii proteomics data.
- Stefan Schulze
- , Zachary Adams
- & Mechthild Pohlschroder
-
Article
| Open Access Cancer neoantigen prioritization through sensitive and reliable proteogenomics analysis
Cancer neoantigen prioritization through sensitive and reliable proteogenomics analysis
Identifying mutation-derived neoantigens by proteogenomics requires robust strategies for quality control. Here, the authors propose peptide retention time as an evaluation metric for proteogenomics quality control methods, and develop a deep learning algorithm for accurate retention time prediction.
- Bo Wen
- , Kai Li
- & Bing Zhang
-
Article
| Open Access Quantitative proteomic landscape of metaplastic breast carcinoma pathological subtypes and their relationship to triple-negative tumors
Quantitative proteomic landscape of metaplastic breast carcinoma pathological subtypes and their relationship to triple-negative tumors
Metaplastic breast carcinoma (MBC) is among the most aggressive subtypes of triple-negative breast cancer (TNBC) but the underlying proteome profiles are unknown. Here, the authors characterize the protein signatures of human MBC tissue samples and their relationship to TNBC and normal breast tissue.
- Sabra I. Djomehri
- , Maria E. Gonzalez
- & Celina G. Kleer
-
Article
| Open Access Generating high quality libraries for DIA MS with empirically corrected peptide predictions
Generating high quality libraries for DIA MS with empirically corrected peptide predictions
Data-independent acquisition-mass spectrometry (MS) typically requires many preparatory MS runs to produce experiment-specific spectral libraries. Here, the authors show that empirical correction of in silico predicted spectral libraries enables efficient generation of high-quality experiment-specific libraries.
- Brian C. Searle
- , Kristian E. Swearingen
- & Mathias Wilhelm
-
Article
| Open Access Rapid and site-specific deep phosphoproteome profiling by data-independent acquisition without the need for spectral libraries
Rapid and site-specific deep phosphoproteome profiling by data-independent acquisition without the need for spectral libraries
Localizing phosphorylation sites by data-independent acquisition (DIA)-based proteomics is still challenging. Here, the authors develop algorithms for phosphosite localization and stoichiometry determination, and incorporate them into single-shot DIA-phosphoproteomics workflows.
- Dorte B. Bekker-Jensen
- , Oliver M. Bernhardt
- & Jesper V. Olsen
-
Article
| Open Access The epichaperome is a mediator of toxic hippocampal stress and leads to protein connectivity-based dysfunction
The epichaperome is a mediator of toxic hippocampal stress and leads to protein connectivity-based dysfunction
The biology of Alzheimer’s disease (AD) remains unknown. We propose AD is a protein connectivity-based dysfunction disorder whereby a switch of the chaperome into epichaperomes rewires proteome-wide connectivity, leading to brain circuitry malfunction that can be corrected by novel therapeutics.
- Maria Carmen Inda
- , Suhasini Joshi
- & Gabriela Chiosis
-
Article
| Open Access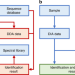 In silico spectral libraries by deep learning facilitate data-independent acquisition proteomics
In silico spectral libraries by deep learning facilitate data-independent acquisition proteomics
Data-independent acquisition (DIA) is an emerging technology in proteomics but it typically relies on spectral libraries built by data-dependent acquisition (DDA). Here, the authors use deep learning to generate in silico spectral libraries directly from protein sequences that enable more comprehensive DIA experiments than DDA-based libraries.
- Yi Yang
- , Xiaohui Liu
- & Liang Qiao
-
Article
| Open Access ProTargetMiner as a proteome signature library of anticancer molecules for functional discovery
ProTargetMiner as a proteome signature library of anticancer molecules for functional discovery
Anticancer drugs often have widespread effects on the cellular proteome. Here, the authors generate a proteome signature library of drug-treated cancer cell lines and develop a software tool to deconvolute drug targets and gain insights into their mechanisms of action.
- Amir Ata Saei
- , Christian Michel Beusch
- & Roman A. Zubarev
-
Article
| Open Access Complex I is bypassed during high intensity exercise
Complex I is bypassed during high intensity exercise
During high-intensity exercise, muscles convert glucose to lactate, in a process that is energetically less efficient than respiration. Here the authors develop a computational model based on muscle proteomic data showing that bypassing mitochondrial complex I increases ATP production rates, and validate these model predictions in an exercise test on 5 subjects.
- Avlant Nilsson
- , Elias Björnson
- & Jens Nielsen
-
Article
| Open Access The distinction of CPR bacteria from other bacteria based on protein family content
The distinction of CPR bacteria from other bacteria based on protein family content
Recent studies have identified a large, phylogenetically distinct clade of bacteria, the candidate phyla radiation (CPR). Here, Méheust and colleagues analyze almost 3600 genomes to characterize the protein family content of CPR versus other bacteria and archaea.
- Raphaël Méheust
- , David Burstein
- & Jillian F. Banfield
-
Article
| Open Access Deep learning extends de novo protein modelling coverage of genomes using iteratively predicted structural constraints
Deep learning extends de novo protein modelling coverage of genomes using iteratively predicted structural constraints
Prediction of protein structures on the scale of genomes remains a challenge. Here the authors introduce a protein structure prediction method that uses deep learning to predict inter-atomic distances, torsion angles and hydrogen bonds, and apply it to predict the structures of 1475 Pfam domains.
- Joe G. Greener
- , Shaun M. Kandathil
- & David T. Jones
-
Article
| Open Access Comparative analysis of mRNA and protein degradation in prostate tissues indicates high stability of proteins
Comparative analysis of mRNA and protein degradation in prostate tissues indicates high stability of proteins
Protein degradation in clinical samples is largely unexplored. Here, the authors analyze the transcriptome and proteome of clinical tissue samples and develop an algorithm to assess protein degradation, showing that protein degradation is negligible in most tissue samples and does not correlate with transcript degradation.
- Wenguang Shao
- , Tiannan Guo
- & Ruedi Aebersold
-
Article
| Open Access An additive Gaussian process regression model for interpretable non-parametric analysis of longitudinal data
An additive Gaussian process regression model for interpretable non-parametric analysis of longitudinal data
Longitudinal data are common in biomedical research, but their analysis is often challenging. Here, the authors present an additive Gaussian process regression model specifically designed for statistical analysis of longitudinal experimental data.
- Lu Cheng
- , Siddharth Ramchandran
- & Harri Lähdesmäki
-
Article
| Open Access Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry
Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry
Data-independent acquisition (DIA)-based proteomics often relies on mass spectrum libraries from data-dependent acquisition experiments. Here, the authors present a method to generate DIA-based chromatogram libraries, enabling DIA-only workflows and detecting more peptides than with spectrum libraries alone.
- Brian C. Searle
- , Lindsay K. Pino
- & Michael J. MacCoss
-
Article
| Open Access Integrative proteomics in prostate cancer uncovers robustness against genomic and transcriptomic aberrations during disease progression
Integrative proteomics in prostate cancer uncovers robustness against genomic and transcriptomic aberrations during disease progression
Understanding of molecular events in cancer requires proteome-level characterisation. Here, proteome profiling of patient samples representing primary and progressed prostate cancer enables the authors to identify pathway alterations that are not reflected at the genomic and transcriptomic levels.
- Leena Latonen
- , Ebrahim Afyounian
- & Tapio Visakorpi
-
Article
| Open Access Discovery of coding regions in the human genome by integrated proteogenomics analysis workflow
Discovery of coding regions in the human genome by integrated proteogenomics analysis workflow
Proteogenomics enables the discovery of protein coding regions and disease-relevant mutations but their verification remains challenging. Here, the authors combine peptide discovery, curation and validation in an integrated proteogenomics workflow, robustly identifying unknown coding regions and mutations.
- Yafeng Zhu
- , Lukas M. Orre
- & Janne Lehtiö
-
Article
| Open Access Noumeavirus replication relies on a transient remote control of the host nucleus
Noumeavirus replication relies on a transient remote control of the host nucleus
Large dsDNA viruses either replicate in or disrupt the nucleus to gain access to host RNA polymerases, or they rely on virus-encoded, packaged RNA polymerases. Here, the authors show that Noumeavirus replicates in the cytoplasm and relies on a transient recruitment of nuclear proteins to initiate replication.
- Elisabeth Fabre
- , Sandra Jeudy
- & Chantal Abergel
-
Article
| Open Access Improving GENCODE reference gene annotation using a high-stringency proteogenomics workflow
Improving GENCODE reference gene annotation using a high-stringency proteogenomics workflow
Identifying and annotating functional elements in the human genome remains a challenging but important task. Here the authors propose a priority annotation score to rank identifications and suggest how proteogenomics evidence can be interpreted and what additional information substantiates protein-coding potential for annotation.
- James C. Wright
- , Jonathan Mudge
- & Jennifer Harrow
-
Article
| Open Access Quantitative maps of protein phosphorylation sites across 14 different rat organs and tissues
Quantitative maps of protein phosphorylation sites across 14 different rat organs and tissues
The function of proteins is often regulated by their phosphorylation at specific amino-acid residues. The authors of this article have catalogued phosphoproteins and their phosphorylation sites in 14 rat organs and tissues, and provide these data as a resource for researchers.
- Alicia Lundby
- , Anna Secher
- & Jesper V. Olsen

 A computational method for detection of ligand-binding proteins from dose range thermal proteome profiles
A computational method for detection of ligand-binding proteins from dose range thermal proteome profiles