Article
|
Open Access
Featured
-
-
Article
| Open Access Emergence of fractal geometries in the evolution of a metabolic enzyme
Emergence of fractal geometries in the evolution of a metabolic enzyme
Citrate synthase from the cyanobacterium Synechococcus elongatus is shown to self-assemble into Sierpiński triangles, a finding that opens up the possibility that other naturally occurring molecular-scale fractals exist.
- Franziska L. Sendker
- , Yat Kei Lo
- & Georg K. A. Hochberg
-
Article
| Open Access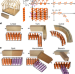 Blueprinting extendable nanomaterials with standardized protein blocks
Blueprinting extendable nanomaterials with standardized protein blocks
A study describes an approach using designed building blocks that are far more regular in geometry than natural proteins to construct modular multicomponent protein assemblies.
- Timothy F. Huddy
- , Yang Hsia
- & David Baker
-
Article
| Open Access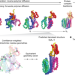 Illuminating protein space with a programmable generative model
Illuminating protein space with a programmable generative model
Evolution has produced a range of diverse proteins, and now a generative model called Chroma can expand that set by allowing the user to design new proteins and protein complexes with desired properties and functions.
- John B. Ingraham
- , Max Baranov
- & Gevorg Grigoryan
-
Article
| Open Access De novo design of protein structure and function with RFdiffusion
De novo design of protein structure and function with RFdiffusion
Fine-tuning the RoseTTAFold structure prediction network on protein structure denoising tasks yields a generative model for protein design that achieves outstanding performance on a wide range of protein structure and function design challenges.
- Joseph L. Watson
- , David Juergens
- & David Baker
-
Article
| Open Access De novo design of protein interactions with learned surface fingerprints
De novo design of protein interactions with learned surface fingerprints
A surface-centric approach captures the physical and chemical determinants of molecular recognition, enabling the de novo design of protein interactions and of artificial proteins with function.
- Pablo Gainza
- , Sarah Wehrle
- & Bruno E. Correia
-
Article
| Open Access De novo design of modular peptide-binding proteins by superhelical matching
De novo design of modular peptide-binding proteins by superhelical matching
A computational approach is used to design modular proteins that bind to synthetic peptides and disordered regions of human proteins with high affinity and specificity.
- Kejia Wu
- , Hua Bai
- & David Baker
-
Article
| Open Access De novo design of luciferases using deep learning
De novo design of luciferases using deep learning
A deep-learning-based strategy is used to design artificial luciferases that catalyse the oxidative chemiluminescence of diphenylterazine with high substrate specificity and catalytic efficiency.
- Andy Hsien-Wei Yeh
- , Christoffer Norn
- & David Baker
-
Article |
 NMR-guided directed evolution
NMR-guided directed evolution
NMR spectroscopy has been used to guide the directed evolution of myoglobin to a Kemp eliminase with high catalytic efficiency, outlining an approach that is likely to be generally applicable to other enzyme activities.
- Sagar Bhattacharya
- , Eleonora G. Margheritis
- & Ivan V. Korendovych
-
Article |
 De novo design of discrete, stable 310-helix peptide assemblies
De novo design of discrete, stable 310-helix peptide assemblies
A study demonstrates the rational de novo design of water-soluble assemblies constructed from long 310-helical peptides, and details their characterization by circular dichroism spectroscopy, analytical ultracentrifugation and X-ray crystallography.
- Prasun Kumar
- , Neil G. Paterson
- & Derek N. Woolfson
-
Article
| Open Access Design of protein-binding proteins from the target structure alone
Design of protein-binding proteins from the target structure alone
A design pipeline is presented whereby binding proteins can be designed de novo without the need for prior information on binding hotspots or fragments from structures of complexes with binding partners.
- Longxing Cao
- , Brian Coventry
- & David Baker
-
Article |
 Overcoming universal restrictions on metal selectivity by protein design
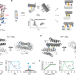
Overcoming universal restrictions on metal selectivity by protein design
An alternative approach to metalloprotein design shows that it is possible to overcome the restrictions of the Irving–Williams series and achieve both flexibility and specificity in the binding of metal ions.
- Tae Su Choi
- & F. Akif Tezcan
-
Article |
 A backbone-centred energy function of neural networks for protein design
A backbone-centred energy function of neural networks for protein design
Modelling by SCUBA of the backbone-centred energy surface extends the diversity of designable proteins.
- Bin Huang
- , Yang Xu
- & Haiyan Liu
-
Article |
 De novo protein design by deep network hallucination
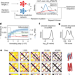
De novo protein design by deep network hallucination
The trRosetta neural network was used to iteratively optimise model proteins from random 100-amino-acid sequences, resulting in ‘hallucinated’ proteins, which when expressed in bacteria closely resembled the model structures.
- Ivan Anishchenko
- , Samuel J. Pellock
- & David Baker
-
Article |
 Bispecific IgG neutralizes SARS-CoV-2 variants and prevents escape in mice
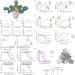
Bispecific IgG neutralizes SARS-CoV-2 variants and prevents escape in mice
The bispecific IgG1-like CoV-X2 prevents SARS-CoV-2 spike binding to ACE2, neutralizes SARS-CoV-2 and its variants of concern, protects against disease in a mouse model, whereas the parental monoclonal antibodies generate viral escape.
- Raoul De Gasparo
- , Mattia Pedotti
- & Luca Varani
-
Article |
 De novo design of modular and tunable protein biosensors
De novo design of modular and tunable protein biosensors
A modular de novo designed biosensor platform consisting of a cage and key molecule is developed, and used to create sensors for seven distinct proteins including the spike protein from SARS-CoV-2 and anti-SARS antibodies.
- Alfredo Quijano-Rubio
- , Hsien-Wei Yeh
- & David Baker
-
Article |
 Computational design of transmembrane pores
Computational design of transmembrane pores
An approach for the design of protein pores is demonstrated by the computational design and subsequent experimental expression of both an ion-selective and a large transmembrane pore.
- Chunfu Xu
- , Peilong Lu
- & David Baker
-
Article |
 Structure of a D2 dopamine receptor–G-protein complex in a lipid membrane
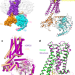
Structure of a D2 dopamine receptor–G-protein complex in a lipid membrane
The structure of the D2 dopamine receptor in complex with its G protein reveals how dopamine receptors are activated and, importantly, how a G-protein-coupled receptor can interact with its G protein in a phospholipid membrane.
- Jie Yin
- , Kuang-Yui M. Chen
- & Daniel M. Rosenbaum
-
Article |
 Constructing protein polyhedra via orthogonal chemical interactions
Constructing protein polyhedra via orthogonal chemical interactions
An inorganic chemical approach to biomolecular design is used to generate ‘cages’ that can simultaneously promote symmetry and multiple modes of protein interactions.
- Eyal Golub
- , Rohit H. Subramanian
- & F. Akif Tezcan
-
Article |
 De novo design of bioactive protein switches
De novo design of bioactive protein switches
A technique for the de novo design of switchable protein systems controlled by induced conformational change is demonstrated for three functional motifs, in vitro and in yeast and mammalian cells.
- Robert A. Langan
- , Scott E. Boyken
- & David Baker
-
Letter |
 Modular and tunable biological feedback control using a de novo protein switch
Modular and tunable biological feedback control using a de novo protein switch
DegronLOCKR designer-protein technology is used to implement synthetic positive- and negative-feedback systems in the yeast mating pathway as well as feedback control of a synthetic gene circuit.
- Andrew H. Ng
- , Taylor H. Nguyen
- & Hana El-Samad
-
Letter |
 De novo protein design by citizen scientists
De novo protein design by citizen scientists
Proteins designed de novo by players of the online protein-folding game Foldit can be expressed in Escherichia coli and adopt the designed structure in solution.
- Brian Koepnick
- , Jeff Flatten
- & David Baker
-
Letter |
 Design and evolution of an enzyme with a non-canonical organocatalytic mechanism
Design and evolution of an enzyme with a non-canonical organocatalytic mechanism
A hydrolytic enzyme with a non-canonical organocatalytic mechanism was generated by introducing Nδ-methylhistidine into a designed active site using engineered translation components, allowing optimization of enzyme performance using laboratory evolution.
- Ashleigh J. Burke
- , Sarah L. Lovelock
- & Anthony P. Green
-
Letter |
 An ultra-stable gold-coordinated protein cage displaying reversible assembly
An ultra-stable gold-coordinated protein cage displaying reversible assembly
An artificial protein cage is readily assembled by metal ion coordination and disassembled by reducing agents, and displays excellent chemical and thermal stability.
- Ali D. Malay
- , Naoyuki Miyazaki
- & Jonathan G. Heddle
-
Article |
 Time-resolved protein activation by proximal decaging in living systems
Time-resolved protein activation by proximal decaging in living systems
The authors report a general strategy for in vivo protein activation using light-controlled proximal decaging directed by in silico screening.
- Jie Wang
- , Yuan Liu
- & Peng R. Chen
-
Article |
 De novo design of potent and selective mimics of IL-2 and IL-15
De novo design of potent and selective mimics of IL-2 and IL-15
A hyper-stable de novo protein mimic of interleukin-2 computationally designed to not interact with a regulatory T-cell specific receptor subunit has improved therapeutic activity in mouse models of melanoma and colon cancer.
- Daniel-Adriano Silva
- , Shawn Yu
- & David Baker
-
Letter |
 Programmable design of orthogonal protein heterodimers
Programmable design of orthogonal protein heterodimers
Computational design incorporating modular buried hydrogen networks produces highly orthogonal protein heterodimers.
- Zibo Chen
- , Scott E. Boyken
- & David Baker
-
Article |
 De novo design of a fluorescence-activating β-barrel
De novo design of a fluorescence-activating β-barrel
The elucidation of general principles for designing β-barrels enables the de novo creation of fluorescent proteins.
- Jiayi Dou
- , Anastassia A. Vorobieva
- & David Baker
-
Letter |
 Acoustic reporter genes for noninvasive imaging of microorganisms in mammalian hosts
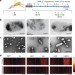
Acoustic reporter genes for noninvasive imaging of microorganisms in mammalian hosts
Heterologous expression of engineered gas vesicles allows noninvasive, deep-tissue ultrasound visualization of engineered bacteria in vivo in mouse tumour models and in the gastrointestinal tract.
- Raymond W. Bourdeau
- , Audrey Lee-Gosselin
- & Mikhail G. Shapiro
-
Article |
 Massively parallel de novo protein design for targeted therapeutics
Massively parallel de novo protein design for targeted therapeutics
A massively parallel computational and experimental approach for de novo designing and screening small hyperstable proteins targeting influenza haemagglutinin and botulinum neurotoxin B identifies new therapeutic candidates more robust than traditional antibody therapies.
- Aaron Chevalier
- , Daniel-Adriano Silva
- & David Baker
-
Letter |
 Proteins evolve on the edge of supramolecular self-assembly
Proteins evolve on the edge of supramolecular self-assembly
Introducing a single ‘sticky’ (hydrophobic) amino acid by point mutation into symmetric protein complexes frequently triggers their association into higher-order assemblies, without affecting their native fold and structure.
- Hector Garcia-Seisdedos
- , Charly Empereur-Mot
- & Emmanuel D. Levy
-
Letter |
 Surrogate Wnt agonists that phenocopy canonical Wnt and β-catenin signalling
Surrogate Wnt agonists that phenocopy canonical Wnt and β-catenin signalling
The authors describe water-soluble surrogate Wnt agonists, with specificity towards some frizzled (FZD) receptors, which can maintain human intestinal organoid cultures and have effects on the mouse liver in vivo.
- Claudia Y. Janda
- , Luke T. Dang
- & K. Christopher Garcia
-
Letter |
 Designed proteins induce the formation of nanocage-containing extracellular vesicles
Designed proteins induce the formation of nanocage-containing extracellular vesicles
Autonomously produced hybrid biological nanomaterials termed ‘enveloped protein nanocages’ incorporate features for membrane binding, self-assembly, and ESCRT recruitment for cellular release.
- Jörg Votteler
- , Cassandra Ogohara
- & Wesley I. Sundquist
-
Article |
 Accurate de novo design of hyperstable constrained peptides
Accurate de novo design of hyperstable constrained peptides
Computational methods for the de novo design of conformationally restricted peptides produce exceptionally stable short peptides stabilized by backbone cyclization and/or internal disulfide bonds that are promising starting points for a new generation of peptide-based drugs.
- Gaurav Bhardwaj
- , Vikram Khipple Mulligan
- & David Baker
-
Letter |
 Design of a hyperstable 60-subunit protein icosahedron
Design of a hyperstable 60-subunit protein icosahedron
The computational design of an extremely stable icosahedral self-assembling protein nanocage is presented; the icosahedron should be useful for applications ranging from calibrating fluorescence microscopy to drug delivery.
- Yang Hsia
- , Jacob B. Bale
- & David Baker
-
Letter |
 Exploring the repeat protein universe through computational protein design
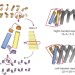
Exploring the repeat protein universe through computational protein design
In this study, 83 proteins containing helix–loop–helix–loop repeats were designed—with sequences unrelated to known repeat proteins—and experimentally characterized; 43 solution X-ray scattering spectra and 15 structures of the designed proteins show that these non-natural repeat proteins have a broad range of curvatures and that their overall structures are in close agreement with design models.
- TJ Brunette
- , Fabio Parmeggiani
- & David Baker
-
Letter |
 Rational design of α-helical tandem repeat proteins with closed architectures
Rational design of α-helical tandem repeat proteins with closed architectures
The development and validation of computational methods for geometry-guided de novo design of tandem repeat protein architectures.
- Lindsey Doyle
- , Jazmine Hallinan
- & Philip Bradley
-
Letter |
 Computational design of co-assembling protein–DNA nanowires
Computational design of co-assembling protein–DNA nanowires
Computational protein design is used to create a protein–DNA co-assembling nanomaterial; by varying the arrangement of protein-binding sites on the double-stranded DNA, a ‘nanowire’ with single-molecule width can be spontaneously formed by mixing the protein and double-stranded DNA building blocks.
- Yun Mou
- , Jiun-Yann Yu
- & Stephen L. Mayo
-
Article |
 Accurate design of co-assembling multi-component protein nanomaterials
Accurate design of co-assembling multi-component protein nanomaterials
A computational method is reported that can be used to design protein nanomaterials in which two distinct subunits co-assemble into a specific architecture; five 24-subunit cage-like protein nanomaterials are designed, and experiments show that their structures are in close agreement with the computational design models.
- Neil P. King
- , Jacob B. Bale
- & David Baker
-
Article |
 Proof of principle for epitope-focused vaccine design
Proof of principle for epitope-focused vaccine design
Computational protein design methods are used to generate new candidates for a human respiratory syncytial virus (RSV) vaccine; artificial protein scaffolds that mimic the structure of a RSV epitope are shown to induce RSV-specific neutralizing antibodies in macaques.
- Bruno E. Correia
- , John T. Bates
- & William R. Schief
-
Letter |
 Precision is essential for efficient catalysis in an evolved Kemp eliminase
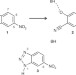
Precision is essential for efficient catalysis in an evolved Kemp eliminase
A computationally designed enzyme that was evolved to accelerate a chemical reaction 6 × 108-fold approaches the exceptional efficiency of highly optimized natural enzymes and provides valuable lessons for the creation of more sophisticated artificial catalysts.
- Rebecca Blomberg
- , Hajo Kries
- & Donald Hilvert
-
Letter |
 Computational design of ligand-binding proteins with high affinity and selectivity
Computational design of ligand-binding proteins with high affinity and selectivity
Computational protein design is used to create a protein that binds the steroid digoxigenin (DIG) with high affinity and selectivity; the computational design methods described here should help to enable the development of a new generation of small molecule receptors for synthetic biology, diagnostics and therapeutics.
- Christine E. Tinberg
- , Sagar D. Khare
- & David Baker
-
News & Views |
 A toolbox for protein design
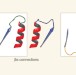
A toolbox for protein design
Some of the principles underlying how amino-acid sequences determine the three-dimensional structures of proteins have been defined. This has enabled a successful approach to designing protein folds from scratch. See Article p.222
- Birte Höcker
-
News |
 Proteins made to order
Proteins made to order
Researchers design proteins from scratch with predictable structures.
- Jessica Marshall
-
Article |
 Principles for designing ideal protein structures
Principles for designing ideal protein structures
Rules that allow the design of strongly funnelled protein folding energy landscapes by relating secondary structure patterns to protein tertiary motifs are used to produce ideal protein structures stabilized by completely consistent local and non-local interactions.
- Nobuyasu Koga
- , Rie Tatsumi-Koga
- & David Baker

 Computationally restoring the potency of a clinical antibody against Omicron
Computationally restoring the potency of a clinical antibody against Omicron