Featured
-
-
Brief Communication |
 A multi-purpose, regenerable, proteome-scale, human phosphoserine resource for phosphoproteomics
A multi-purpose, regenerable, proteome-scale, human phosphoserine resource for phosphoproteomics
Iterative Synthetically Phosphorylated Isomers (iSPI) is a proteome-scale library of human-derived phosphoserine-containing phosphopeptides with precisely known positions of phosphorylation. This multi-purpose resource is available for optimization, standardization, and benchmarking of key steps in phosphoproteomics workflows.
- Brandon M. Gassaway
- , Jiaming Li
- & Steven P. Gygi
-
Brief Communication |
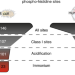 Histidine phosphorylation in human cells; a needle or phantom in the haystack?
Histidine phosphorylation in human cells; a needle or phantom in the haystack?
Extensive analyses of mammalian phosphoproteomics datasets show that protein histidine phosphorylation in human cells may not be as prevalent as previously thought.
- Niels M. Leijten
- , Albert J. R. Heck
- & Simone Lemeer
-
Article |
 Global profiling of phosphorylation-dependent changes in cysteine reactivity
Global profiling of phosphorylation-dependent changes in cysteine reactivity
This article describes a chemical proteomic approach to quantitatively relate serine/threonine phosphorylation to changes in the reactivity of cysteine residues, thereby affecting their potential to be post-translationally modified and/or targeted by electrophilic small molecules.
- Esther K. Kemper
- , Yuanjin Zhang
- & Benjamin F. Cravatt
-
Research Highlight |
Engineered protein–protein toggle network
Sensitive synthetic protein-phosphorylation networks manifest fast bistable dynamics.
- Lin Tang
-
Matters Arising |
 Impact of phosphorylation on thermal stability of proteins
Impact of phosphorylation on thermal stability of proteins
- Clément M. Potel
- , Nils Kurzawa
- & Mikhail M. Savitski
-
Matters Arising |
 Identification of phosphosites that alter protein thermal stability
Identification of phosphosites that alter protein thermal stability
- Ian R. Smith
- , Kyle N. Hess
- & Judit Villén
-
-
-
Article |
 Cell-type-specific signaling networks in heterocellular organoids
Cell-type-specific signaling networks in heterocellular organoids
Mass cytometry in combination with a thiol-reactive barcoding strategy allows analysis and comparison of cell-type-specific signaling networks in organoids.
- Xiao Qin
- , Jahangir Sufi
- & Christopher J. Tape
-
Article |
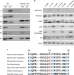 An antibody for analysis of autophagy induction
An antibody for analysis of autophagy induction
A new method of autophagy measurement is based on the detection of phospho-ATG16L1, a conserved early marker of autophagy. Sensitive detection can be achieved in multiple biological systems and assays with advantages over standard methods.
- Wensheng Tian
- , Reham Alsaadi
- & Ryan C. Russell
-
Article |
 High throughput discovery of functional protein modifications by Hotspot Thermal Profiling
High throughput discovery of functional protein modifications by Hotspot Thermal Profiling
A massively parallel biophysical approach, Hotspot Thermal Profiling, analyzes the effects of phosphorylation on protein stability across the proteome in live cells.
- Jun X. Huang
- , Gihoon Lee
- & Raymond E. Moellering
-
Brief Communication |
 Thesaurus: quantifying phosphopeptide positional isomers
Thesaurus: quantifying phosphopeptide positional isomers
The search engine Thesaurus detects and quantifies phosphopeptide positional isomers from data-independent acquisition and parallel reaction monitoring mass spectrometry data, enabling studies of how neighboring phosphosites are regulated.
- Brian C. Searle
- , Robert T. Lawrence
- & Judit Villén
-
Brief Communication |
 Efficient and precise editing of endogenous transcripts with SNAP-tagged ADARs
Efficient and precise editing of endogenous transcripts with SNAP-tagged ADARs
Fusion of a guide RNA directly to an adenosine deaminase via a SNAP-tag allows for precise and efficient RNA editing.
- Paul Vogel
- , Matin Moschref
- & Thorsten Stafforst
-
Brief Communication |
 Widespread bacterial protein histidine phosphorylation revealed by mass spectrometry-based proteomics
Widespread bacterial protein histidine phosphorylation revealed by mass spectrometry-based proteomics
A twist on a common method used for enriching phosphorylated peptides for mass spectrometry-based proteomics analysis now reveals previously undetected and widespread histidine phosphorylation in Escherichia coli.
- Clement M Potel
- , Miao-Hsia Lin
- & Simone Lemeer
-
Article |
 Genetically encoded biosensors for visualizing live-cell biochemical activity at super-resolution
Genetically encoded biosensors for visualizing live-cell biochemical activity at super-resolution
New fluorescent biosensors enable the first super-resolution imaging of enzyme activity in live cells via fluorescence fluctuation increase by contact (FLINC).
- Gary C H Mo
- , Brian Ross
- & Jin Zhang
-
Brief Communication |
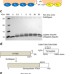 Enzyme-catalyzed expressed protein ligation
Enzyme-catalyzed expressed protein ligation
The enzyme subtiligase can be used to catalyze expressed protein ligation of proteins of interest with peptides lacking an N-terminal cysteine. This enables the analysis of protein modifications in the context of the native primary sequence.
- Samuel H Henager
- , Nam Chu
- & Philip A Cole
-
This Month |
 Judit Villén
Judit Villén
Measuring phosphoproteins reproducibly and taking time to bike.
- Vivien Marx
-
Brief Communication |
 Plug-and-play analysis of the human phosphoproteome by targeted high-resolution mass spectrometry
Plug-and-play analysis of the human phosphoproteome by targeted high-resolution mass spectrometry
A method (and resource) demonstrating the mining of information from large-scale phosphoproteomics data sets is presented, allowing users to build targeted parallel reaction monitoring mass spectrometry assays to study phosphosites of interest.
- Robert T Lawrence
- , Brian C Searle
- & Judit Villén
-
Research Highlights |
Efficient generation of proteins with site-specific phosphoserines
An optimized strategy allows genetic encoding of phosphoserine and its nonhydrolyzable analog.
- Rita Strack
-
Methods in Brief |
Phosphohistidine proteomics
-
Research Highlights |
Nanopores sense protein modifications
Protein nanopores can detect post-translational modifications in proteins.
- Tal Nawy
-
Research Highlights |
Antibodies, made to order
Informed by existing molecular recognition motifs, researchers design effective scaffolds for directed evolution of post-translational modification–specific antibodies.
- Michael Eisenstein
-
Article |
 Global analysis of phosphorylation and ubiquitylation cross-talk in protein degradation
Global analysis of phosphorylation and ubiquitylation cross-talk in protein degradation
Two methods for identifying protein isoforms that are concurrently phosphorylated and ubiquitylated are applied in yeast to identify phosphorylation sites that regulate ubiquitin proteasome–mediated proteome degradation.
- Danielle L Swaney
- , Pedro Beltrao
- & Judit Villén
-
Tools in Brief |
Measuring phosphorylation-site stoichiometry
-
Article |
 In situ analysis of tyrosine phosphorylation networks by FLIM on cell arrays
In situ analysis of tyrosine phosphorylation networks by FLIM on cell arrays
Combining reverse transfection of protein tyrosine kinase substrates on cell arrays with fluorescence resonance energy transfer (FRET) measurements by fluorescence lifetime imaging microscopy (FLIM) allows quantitative assessment of phosphorylation patterns and identification of feedback loops at single-cell resolution.
- Hernán E Grecco
- , Pedro Roda-Navarro
- & Philippe I H Bastiaens

 Post-translational modification-centric base editor screens to assess phosphorylation site functionality in high throughput
Post-translational modification-centric base editor screens to assess phosphorylation site functionality in high throughput Huang et al. reply
Huang et al. reply