Featured
-
-
Article
| Open Access Amino acid auxotrophies in human gut bacteria are linked to higher microbiome diversity and long-term stability
Amino acid auxotrophies in human gut bacteria are linked to higher microbiome diversity and long-term stability
- Svenja Starke
- , Danielle M. M. Harris
- & Silvio Waschina
-
Article
| Open Access Genomic and transcriptomic insights into complex virus–prokaryote interactions in marine biofilms
Genomic and transcriptomic insights into complex virus–prokaryote interactions in marine biofilms
- Kun Zhou
- , Tin Yan Wong
- & Pei-Yuan Qian
-
Article |
 Oxytetracycline and heavy metals promote the migration of resistance genes in the intestinal microbiome by plasmid transfer
Oxytetracycline and heavy metals promote the migration of resistance genes in the intestinal microbiome by plasmid transfer
- Xiaojun Lin
- , Chaonan Zhang
- & Yanbin Xu
-
Article
| Open Access Host and microbiome jointly contribute to environmental adaptation
Host and microbiome jointly contribute to environmental adaptation
- Carola Petersen
- , Inga K. Hamerich
- & Hinrich Schulenburg
-
Article
| Open Access Starvation responses impact interaction dynamics of human gut bacteria Bacteroides thetaiotaomicron and Roseburia intestinalis
Starvation responses impact interaction dynamics of human gut bacteria Bacteroides thetaiotaomicron and Roseburia intestinalis
- Bin Liu
- , Daniel Rios Garza
- & Karoline Faust
-
Comment
| Open Access Novel T4ASS effector with quorum quenching activity
Novel T4ASS effector with quorum quenching activity
- Vittorio Venturi
- & Cristina Bez
-
Article |
 Physiological and evolutionary contexts of a new symbiotic species from the nitrogen-recycling gut community of turtle ants
Physiological and evolutionary contexts of a new symbiotic species from the nitrogen-recycling gut community of turtle ants
- Benoît Béchade
- , Christian S. Cabuslay
- & Jacob A. Russell
-
Review Article
| Open Access Pesticide exposure and the microbiota-gut-brain axis
Pesticide exposure and the microbiota-gut-brain axis
- Rie Matsuzaki
- , Eoin Gunnigle
- & John F. Cryan
-
Article
| Open Access Growth rate is a dominant factor predicting the rhizosphere effect
Growth rate is a dominant factor predicting the rhizosphere effect
- José L. López
- , Arista Fourie
- & Bas E. Dutilh
-
Article
| Open Access Delivery mechanism can enhance probiotic activity against honey bee pathogens
Delivery mechanism can enhance probiotic activity against honey bee pathogens
- Brendan A. Daisley
- , Andrew P. Pitek
- & Elina Niño
-
Article
| Open Access Sulfur cycling connects microbiomes and biogeochemistry in deep-sea hydrothermal plumes
Sulfur cycling connects microbiomes and biogeochemistry in deep-sea hydrothermal plumes
- Zhichao Zhou
- , Patricia Q. Tran
- & Karthik Anantharaman
-
Article
| Open Access Metabolic influence of core ciliates within the rumen microbiome
Metabolic influence of core ciliates within the rumen microbiome
- Thea O. Andersen
- , Ianina Altshuler
- & Phillip B. Pope
-
-
Article |
 Mining chicken ileal microbiota for immunomodulatory microorganisms
Mining chicken ileal microbiota for immunomodulatory microorganisms
- Yan Liu
- , Yuqing Feng
- & Dan Liu
-
Article
| Open Access Co-diversification of an intestinal Mycoplasma and its salmonid host
Co-diversification of an intestinal Mycoplasma and its salmonid host
- Jacob A. Rasmussen
- , Pia Kiilerich
- & Morten T. Limborg
-
Brief Communication
| Open Access Pulsed, continuous or somewhere in between? Resource dynamics matter in the optimisation of microbial communities
Pulsed, continuous or somewhere in between? Resource dynamics matter in the optimisation of microbial communities
- Andrew D. Letten
- & William B. Ludington
-
Article
| Open Access Dissecting microbial communities and resistomes for interconnected humans, soil, and livestock
Dissecting microbial communities and resistomes for interconnected humans, soil, and livestock
- Alexandre Maciel-Guerra
- , Michelle Baker
- & Tania Dottorini
-
Article |
 Suppression of the insect cuticular microbiomes by a fungal defensin to facilitate parasite infection
Suppression of the insect cuticular microbiomes by a fungal defensin to facilitate parasite infection
- Song Hong
- , Yanlei Sun
- & Chengshu Wang
-
Article
| Open Access Genomic diversity and biosynthetic capabilities of sponge-associated chlamydiae
Genomic diversity and biosynthetic capabilities of sponge-associated chlamydiae
- Jennah E. Dharamshi
- , Natalia Gaarslev
- & Thijs J. G. Ettema
-
Article
| Open Access Metaproteome plasticity sheds light on the ecology of the rumen microbiome and its connection to host traits
Metaproteome plasticity sheds light on the ecology of the rumen microbiome and its connection to host traits
- Goor Sasson
- , Sarah Moraïs
- & Itzhak Mizrahi
-
Article
| Open Access Atmospheric chemosynthesis is phylogenetically and geographically widespread and contributes significantly to carbon fixation throughout cold deserts
Atmospheric chemosynthesis is phylogenetically and geographically widespread and contributes significantly to carbon fixation throughout cold deserts
- Angelique E. Ray
- , Julian Zaugg
- & Belinda C. Ferrari
-
Article |
 Microbiome of the freshwater sponge Ephydatia muelleri shares compositional and functional similarities with those of marine sponges
Microbiome of the freshwater sponge Ephydatia muelleri shares compositional and functional similarities with those of marine sponges
- Scott Sugden
- , Johannes Holert
- & Lisa Y. Stein
-
Article |
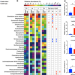 Amino acid utilization allows intestinal dominance of Lactobacillus amylovorus
Amino acid utilization allows intestinal dominance of Lactobacillus amylovorus
- Yujia Jing
- , Chunlong Mu
- & Weiyun Zhu
-
Article |
 Ecological memory of prior nutrient exposure in the human gut microbiome
Ecological memory of prior nutrient exposure in the human gut microbiome
- Jeffrey Letourneau
- , Zachary C. Holmes
- & Lawrence A. David
-
Article
| Open Access Rapid bacterioplankton transcription cascades regulate organic matter utilization during phytoplankton bloom progression in a coastal upwelling system
Rapid bacterioplankton transcription cascades regulate organic matter utilization during phytoplankton bloom progression in a coastal upwelling system
- Benjamin Pontiller
- , Sandra Martínez-García
- & Jarone Pinhassi
-
Article
| Open Access Cross-feeding niches among commensal leaf bacteria are shaped by the interaction of strain-level diversity and resource availability
Cross-feeding niches among commensal leaf bacteria are shaped by the interaction of strain-level diversity and resource availability
- Mariana Murillo-Roos
- , Hafiz Syed M. Abdullah
- & Matthew T. Agler
-
Article
| Open Access Priority effects shape the structure of infant-type Bifidobacterium communities on human milk oligosaccharides
Priority effects shape the structure of infant-type Bifidobacterium communities on human milk oligosaccharides
- Miriam N. Ojima
- , Lin Jiang
- & Takane Katayama
-
Article |
 Mobile genetic elements used by competing coral microbial populations increase genomic plasticity
Mobile genetic elements used by competing coral microbial populations increase genomic plasticity
- Pengxia Wang
- , Yi Zhao
- & Xiaoxue Wang
-
Article
| Open Access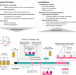 Dynamic metabolic interactions and trophic roles of human gut microbes identified using a minimal microbiome exhibiting ecological properties
Dynamic metabolic interactions and trophic roles of human gut microbes identified using a minimal microbiome exhibiting ecological properties
- Sudarshan A. Shetty
- , Ioannis Kostopoulos
- & Clara Belzer
-
Article
| Open Access Complex and unexpected outcomes of antibiotic therapy against a polymicrobial infection
Complex and unexpected outcomes of antibiotic therapy against a polymicrobial infection
- Lydia-Ann J. Ghuneim
- , Ruma Raghuvanshi
- & Robert A. Quinn
-
Article |
 Ecological dynamics of the gut microbiome in response to dietary fiber
Ecological dynamics of the gut microbiome in response to dietary fiber
- Hongbin Liu
- , Chen Liao
- & Lei Dai
-
Article
| Open Access Identification of beneficial and detrimental bacteria impacting sorghum responses to drought using multi-scale and multi-system microbiome comparisons
Identification of beneficial and detrimental bacteria impacting sorghum responses to drought using multi-scale and multi-system microbiome comparisons
- Mingsheng Qi
- , Jeffrey C. Berry
- & Rebecca S. Bart
-
Article |
 Trophic interactions between predatory protists and pathogen-suppressive bacteria impact plant health
Trophic interactions between predatory protists and pathogen-suppressive bacteria impact plant health
- Sai Guo
- , Chengyuan Tao
- & Stefan Geisen
-
Article
| Open Access Synthetic bacterial community derived from a desert rhizosphere confers salt stress resilience to tomato in the presence of a soil microbiome
Synthetic bacterial community derived from a desert rhizosphere confers salt stress resilience to tomato in the presence of a soil microbiome
- Lucas Schmitz
- , Zhichun Yan
- & Xu Cheng
-
Article
| Open Access Pseudomonas aeruginosa polysaccharide Psl supports airway microbial community development
Pseudomonas aeruginosa polysaccharide Psl supports airway microbial community development
- Sara N. Stoner
- , Joshua J. Baty
- & Jessica A. Scoffield
-
Article
| Open Access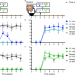 Experimental evaluation of ecological principles to understand and modulate the outcome of bacterial strain competition in gut microbiomes
Experimental evaluation of ecological principles to understand and modulate the outcome of bacterial strain competition in gut microbiomes
- Rafael R. Segura Munoz
- , Sara Mantz
- & Amanda E. Ramer-Tait
-
Article
| Open Access Mammalian gut metabolomes mirror microbiome composition and host phylogeny
Mammalian gut metabolomes mirror microbiome composition and host phylogeny
- Rachel Gregor
- , Maraike Probst
- & Itzhak Mizrahi
-
Article
| Open Access Carbon assimilating fungi from surface ocean to subseafloor revealed by coupled phylogenetic and stable isotope analysis
Carbon assimilating fungi from surface ocean to subseafloor revealed by coupled phylogenetic and stable isotope analysis
- William D. Orsi
- , Aurèle Vuillemin
- & Gonzalo V. Gomez-Saez
-
Article
| Open Access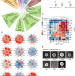 In vitro interaction network of a synthetic gut bacterial community
In vitro interaction network of a synthetic gut bacterial community
- Anna S. Weiss
- , Anna G. Burrichter
- & Bärbel Stecher
-
Article
| Open Access Mechanisms underlying interactions between two abundant oral commensal bacteria
Mechanisms underlying interactions between two abundant oral commensal bacteria
- Dasith Perera
- , Anthony McLean
- & Matthew Ramsey
-
Review Article
| Open Access Plant-microbe interactions in the phyllosphere: facing challenges of the anthropocene
Plant-microbe interactions in the phyllosphere: facing challenges of the anthropocene
- Rosaëlle Perreault
- & Isabelle Laforest-Lapointe
-
Perspective
| Open Access Phages in the infant gut: a framework for virome development during early life
Phages in the infant gut: a framework for virome development during early life
- Michael Shamash
- & Corinne F. Maurice
-
Article
| Open Access Network structure of resource use and niche overlap within the endophytic microbiome
Network structure of resource use and niche overlap within the endophytic microbiome
- Matthew Michalska-Smith
- , Zewei Song
- & Linda L. Kinkel
-
Article
| Open Access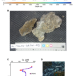 Quantifying the effects of hydrogen on carbon assimilation in a seafloor microbial community associated with ultramafic rocks
Quantifying the effects of hydrogen on carbon assimilation in a seafloor microbial community associated with ultramafic rocks
- Ömer K. Coskun
- , Aurèle Vuillemin
- & William D. Orsi
-
Article |
 Cytokinin drives assembly of the phyllosphere microbiome and promotes disease resistance through structural and chemical cues
Cytokinin drives assembly of the phyllosphere microbiome and promotes disease resistance through structural and chemical cues
- Rupali Gupta
- , Dorin Elkabetz
- & Maya Bar
-
Article
| Open Access Modeling host-associating microbes under selection
Modeling host-associating microbes under selection
- Florence Bansept
- , Nancy Obeng
- & Arne Traulsen
-
Perspective
| Open Access Open challenges for microbial network construction and analysis
Open challenges for microbial network construction and analysis
- Karoline Faust
-
Article
| Open Access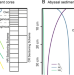 Microbial community structure in hadal sediments: high similarity along trench axes and strong changes along redox gradients
Microbial community structure in hadal sediments: high similarity along trench axes and strong changes along redox gradients
- Clemens Schauberger
- , Ronnie N. Glud
- & Bo Thamdrup
-
Article
| Open Access Compositional and genetic alterations in Graves’ disease gut microbiome reveal specific diagnostic biomarkers
Compositional and genetic alterations in Graves’ disease gut microbiome reveal specific diagnostic biomarkers
- Qiyun Zhu
- , Qiangchuan Hou
- & Jiachao Zhang

 Ginsenoside Rg3 enriches SCFA-producing commensal bacteria to confer protection against enteric viral infection via the cGAS-STING-type I IFN axis
Ginsenoside Rg3 enriches SCFA-producing commensal bacteria to confer protection against enteric viral infection via the cGAS-STING-type I IFN axis