Article
|
Open Access
Featured
-
-
Article
| Open Access Gut microbial carbohydrate metabolism contributes to insulin resistance
Gut microbial carbohydrate metabolism contributes to insulin resistance
Faecal carbohydrates, particularly host-accessible monosaccharides, are increased in individuals with insulin resistance and are associated with microbial carbohydrate metabolisms and host inflammatory cytokines.
- Tadashi Takeuchi
- , Tetsuya Kubota
- & Hiroshi Ohno
-
Article
| Open Access Inference and reconstruction of the heimdallarchaeial ancestry of eukaryotes
Inference and reconstruction of the heimdallarchaeial ancestry of eukaryotes
Analyses of multiple phylogenetic marker datasets of Asgard archaea provide insight into the transition from prokaryotes to eukaryotes, specifically placing eukaryotes within Asgard archaea and as a sister lineage to Hodarchaeales.
- Laura Eme
- , Daniel Tamarit
- & Thijs J. G. Ettema
-
Article
| Open Access Mirusviruses link herpesviruses to giant viruses
Mirusviruses link herpesviruses to giant viruses
A phylogeny-guided genome-resolved metagenomic analysis of DNA viruses in the ocean reveals atypical plankton-infecting relatives of herpesviruses that form a putative new phylum dubbed Mirusviricota.
- Morgan Gaïa
- , Lingjie Meng
- & Tom O. Delmont
-
Article
| Open Access The person-to-person transmission landscape of the gut and oral microbiomes
The person-to-person transmission landscape of the gut and oral microbiomes
Data from more than 9,700 human stool and oral metagenomes has been used to decipher the strain transmission patterns of the human microbiome from mother to infant, within households and within populations.
- Mireia Valles-Colomer
- , Aitor Blanco-Míguez
- & Nicola Segata
-
Article
| Open Access Decoupling of respiration rates and abundance in marine prokaryoplankton
Decoupling of respiration rates and abundance in marine prokaryoplankton
Cell-specific respiration rates differ by more than 1,000× among prokaryoplankton genera, and the majority of respiration was found to be performed by minority members of prokaryoplankton, whereas cells of the most prevalent lineages had extremely low respiration rates.
- Jacob H. Munson-McGee
- , Melody R. Lindsay
- & Ramunas Stepanauskas
-
Article
| Open Access Borgs are giant genetic elements with potential to expand metabolic capacity
Borgs are giant genetic elements with potential to expand metabolic capacity
Borgs are remarkably large, divergent archaeal extrachromosomal elements with metabolic genes linked to the methane cycle.
- Basem Al-Shayeb
- , Marie C. Schoelmerich
- & Jillian F. Banfield
-
Article |
 Discovery of bioactive microbial gene products in inflammatory bowel disease
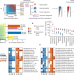
Discovery of bioactive microbial gene products in inflammatory bowel disease
A computational system termed MetaWIBELE (workflow to identify novel bioactive elements in the microbiome) is used to identify microbial gene products that are potentially bioactive and have a functional role in the pathogenesis of inflammatory bowel disease.
- Yancong Zhang
- , Amrisha Bhosle
- & Eric A. Franzosa
-
Article |
 Petabase-scale sequence alignment catalyses viral discovery
Petabase-scale sequence alignment catalyses viral discovery
Serratus, an open-source cloud-computing infrastructure, can be used to screen millions of nucleic acid sequencing libraries at the petabase scale, and has enabled many new RNA viruses to be identified efficiently.
- Robert C. Edgar
- , Brie Taylor
- & Artem Babaian
-
Article
| Open Access Late Quaternary dynamics of Arctic biota from ancient environmental genomics
Late Quaternary dynamics of Arctic biota from ancient environmental genomics
A large-scale metagenomic analysis of plant and mammal environmental DNA reveals complex ecological changes across the circumpolar region over the past 50,000 years, as biota responded to changing climates, culminating in the postglacial extinction of large mammals and emergence of modern ecosystems.
- Yucheng Wang
- , Mikkel Winther Pedersen
- & Eske Willerslev
-
Article
| Open Access Reconstruction of ancient microbial genomes from the human gut
Reconstruction of ancient microbial genomes from the human gut
Ancient microbiomes from palaeofaeces are more similar to non-industrialized than industrialized human gut microbiomes regardless of geography, but 39% of their de novo reconstructed genomes represent previously undescribed microbial species.
- Marsha C. Wibowo
- , Zhen Yang
- & Aleksandar D. Kostic
-
Article |
 Expanded diversity of Asgard archaea and their relationships with eukaryotes
Expanded diversity of Asgard archaea and their relationships with eukaryotes
Comparative analysis of 162 genomes of Asgard archaea results in six newly proposed phyla, including a deep branch that is provisionally named Wukongarchaeota, and sheds light on the evolutionary history of this clade.
- Yang Liu
- , Kira S. Makarova
- & Meng Li
-
Article |
 Statin therapy is associated with lower prevalence of gut microbiota dysbiosis
Statin therapy is associated with lower prevalence of gut microbiota dysbiosis
A cross-sectional analysis of participants in the MetaCardis Body Mass Index Spectrum cohort finds that the higher prevalence of gut microbiota dysbiosis in individuals with obesity is not observed in those who take statin drugs.
- Sara Vieira-Silva
- , Gwen Falony
- & Jeroen Raes
-
Article |
 Recycling and metabolic flexibility dictate life in the lower oceanic crust
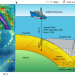
Recycling and metabolic flexibility dictate life in the lower oceanic crust
Analyses of microbial communities that live 10–750 m below the seafloor at Atlantis Bank, Indian Ocean, provide insights into how these microorganisms survive by coupling energy sources to organic and inorganic carbon resources.
- Jiangtao Li
- , Paraskevi Mara
- & Virginia P. Edgcomb
-
Article |
 Microbiome analyses of blood and tissues suggest cancer
diagnostic approach
Microbiome analyses of blood and tissues suggest cancer
diagnostic approach
Microbial nucleic acids are detected in samples of tissues and blood from more than 10,000 patients with cancer, and machine learning is used to show that these can be used to discriminate between and among different types of cancer, suggesting a new microbiome-based diagnostic approach.
- Gregory D. Poore
- , Evguenia Kopylova
- & Rob Knight
-
Article
| Open Access Clades of huge phages from across Earth’s ecosystems
Clades of huge phages from across Earth’s ecosystems
Genomic analyses of major clades of huge phages sampled from across Earth’s ecosystems show that they have diverse genetic inventories, including a variety of CRISPR–Cas systems and translation-relevant genes.
- Basem Al-Shayeb
- , Rohan Sachdeva
- & Jillian F. Banfield
-
Letter |
 Stunted microbiota and opportunistic pathogen colonization in caesarean-section birth
Stunted microbiota and opportunistic pathogen colonization in caesarean-section birth
Delivery via caesarean section, maternal antibiotic prophylaxis and colonization by opportunistic pathogens associated with the hospital environment affect the composition of the gut microbiota of children from birth until infancy.
- Yan Shao
- , Samuel C. Forster
- & Trevor D. Lawley
-
Letter |
 Microbiota-derived lantibiotic restores resistance against vancomycin-resistant Enterococcus
Microbiota-derived lantibiotic restores resistance against vancomycin-resistant Enterococcus
The gut commensal Blautia producta secretes a lantibiotic that reduces colonization of the gut by the major pathogen vancomycin-resistant Enterococcus faecium, and transplantation of microbiota with high abundance of the lantibiotic gene enhances resistance to colonization in mice.
- Sohn G. Kim
- , Simone Becattini
- & Eric G. Pamer
-
Article |
 Human placenta has no microbiome but can contain potential pathogens
Human placenta has no microbiome but can contain potential pathogens
The human placenta does not have a microbiota, suggesting that bacterial infection of the placenta is not a common cause of adverse pregnancy outcome, but group B Streptococcus is found in approximately 5% of placental samples.
- Marcus C. de Goffau
- , Susanne Lager
- & Gordon C. S. Smith
-
Letter |
 Anaerobic oxidation of ethane by archaea from a marine hydrocarbon seep
Anaerobic oxidation of ethane by archaea from a marine hydrocarbon seep
An archaeon, ‘Candidatus Argoarchaeum ethanivorans’, which is involved in the oxidation of ethane observed in anoxic marine habitats, is identified and metabolically characterized.
- Song-Can Chen
- , Niculina Musat
- & Florin Musat
-
Article
| Open Access A new genomic blueprint of the human gut microbiota
A new genomic blueprint of the human gut microbiota
The known species repertoire of the collective human gut microbiota is substantially expanded with the discovery of 1,952 uncultured bacterial species that greatly improve classification of understudied African and South American samples.
- Alexandre Almeida
- , Alex L. Mitchell
- & Robert D. Finn
-
Letter
| Open Access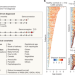 The human gut microbiome in early-onset type 1 diabetes from the TEDDY study
The human gut microbiome in early-onset type 1 diabetes from the TEDDY study
An analysis of more than 10,000 metagenomes from the TEDDY study provides a detailed functional profile of the gut microbiome in relation to islet autoimmunity, and supports the protective effects of short-chain fatty acids in early-onset type 1 diabetes.
- Tommi Vatanen
- , Eric A. Franzosa
- & Ramnik J. Xavier
-
Article |
 Genome-centric view of carbon processing in thawing permafrost
Genome-centric view of carbon processing in thawing permafrost
Analysis of more than 1,500 microbial genomes sheds light on the processing of carbon released as permafrost thaws.
- Ben J. Woodcroft
- , Caitlin M. Singleton
- & Gene W. Tyson
-
Letter |
 Novel soil bacteria possess diverse genes for secondary metabolite biosynthesis
Novel soil bacteria possess diverse genes for secondary metabolite biosynthesis
Metagenomic and soil microcosm analyses identify abundant biosynthetic gene clusters in genomes of microorganisms from a northern Californian grassland ecosystem that provide a potential source for the future development of bacterial natural products.
- Alexander Crits-Christoph
- , Spencer Diamond
- & Jillian F. Banfield
-
Letter |
 Deep mitochondrial origin outside the sampled alphaproteobacteria
Deep mitochondrial origin outside the sampled alphaproteobacteria
Genome data for thirteen alphaproteobacteria-related clades expand the coverage of alphaproteobacterial diversity and suggest that mitochondria diverged from Alphaproteobacteria before the diversification of all currently known alphaproteobacterial lineages.
- Joran Martijn
- , Julian Vosseberg
- & Thijs J. G. Ettema
-
Letter
| Open Access Atmospheric trace gases support primary production in Antarctic desert surface soil
Atmospheric trace gases support primary production in Antarctic desert surface soil
Metagenomic and biochemical analyses of soil samples from Antarctic desert regions provides evidence that bacteria in these soils derive carbon and energy from atmospheric CO, H2 and CO2.
- Mukan Ji
- , Chris Greening
- & Belinda C. Ferrari
-
Article
| Open Access Strains, functions and dynamics in the expanded Human Microbiome Project
Strains, functions and dynamics in the expanded Human Microbiome Project
Updates from the Human Microbiome Project analyse the largest known body-wide metagenomic profile of human microbiome personalization.
- Jason Lloyd-Price
- , Anup Mahurkar
- & Curtis Huttenhower
-
Article |
 Commensal bacteria make GPCR ligands that mimic human signalling molecules
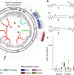
Commensal bacteria make GPCR ligands that mimic human signalling molecules
Commensal bacteria have N-acyl amide synthase genes that encode signalling molecules (N-acyl amides) that can interact with G-protein-coupled receptors and elicit host cellular responses similar to eukaryotic N-acyl amides.
- Louis J. Cohen
- , Daria Esterhazy
- & Sean F. Brady
-
Article |
 Asgard archaea illuminate the origin of eukaryotic cellular complexity
Asgard archaea illuminate the origin of eukaryotic cellular complexity
This work describes the Asgard superphylum, an assemblage of diverse archaea that comprises Odinarchaeota, Heimdallarchaeota, Lokiarchaeota and Thorarchaeota, offering insights into the earliest days of eukaryotic cells and their complex features.
- Katarzyna Zaremba-Niedzwiedzka
- , Eva F. Caceres
- & Thijs J. G. Ettema
-
Letter |
 Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses
Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses
The assembly and analysis of complete genomes and large genomic fragments have tripled the number of known ocean viruses and uncovered the potentially important roles they play in nitrogen and sulfur cycling.
- Simon Roux
- , Jennifer R. Brum
- & Matthew B. Sullivan
-
Letter |
 Mobile genes in the human microbiome are structured from global to individual scales
Mobile genes in the human microbiome are structured from global to individual scales
Mobile genes, which can be transferred between bacterial species in the microbiome to impart properties such as antibiotic resistance, are reflective of human activity and local diets.
- I. L. Brito
- , S. Yilmaz
- & E. J. Alm
-
Article |
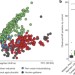 Interconnected microbiomes and resistomes in low-income human habitats
Interconnected microbiomes and resistomes in low-income human habitats
An analysis of bacterial community structure and antibiotic resistance gene content of interconnected human faecal and environmental samples from two low-income communities in Latin America was carried out using a combination of functional metagenomics, 16S sequencing and shotgun sequencing; resistomes across habitats are generally structured along ecological gradients, but key resistance genes can cross these boundaries, and the authors assessed the usefulness of excreta management protocols in the prevention of resistance gene dissemination.
- Erica C. Pehrsson
- , Pablo Tsukayama
- & Gautam Dantas
-
Letter
| Open Access Culturing of ‘unculturable’ human microbiota reveals novel taxa and extensive sporulation
Culturing of ‘unculturable’ human microbiota reveals novel taxa and extensive sporulation
A novel approach is used to cultivate a substantial proportion of the human gut microbiota, representing an important step forward in characterizing the role of these bacteria in health and disease.
- Hilary P. Browne
- , Samuel C. Forster
- & Trevor D. Lawley
-
Letter |
 Complete nitrification by a single microorganism
Complete nitrification by a single microorganism
Until now, the oxidation steps necessary for complete nitrification had always been observed to occur in two separate microorganisms in a cross-feeding interaction; here, together with the study by Daims et al., van Kessel et al. report the enrichment and characterization of Nitrospira species that encode all of the enzymes necessary to catalyse complete nitrification, a phenotype referred to as ‘comammox’ (for complete ammonia oxidation).
- Maartje A. H. J. van Kessel
- , Daan R. Speth
- & Sebastian Lücker
-
Letter |
 Unusual biology across a group comprising more than 15% of domain Bacteria
Unusual biology across a group comprising more than 15% of domain Bacteria
More than 15% of the bacterial domain consists of a radiation of phyla about which very little is known; here, metagenomics is used to reconstruct 8 complete and 789 draft genomes from more than 35 of these phyla, revealing a shared evolutionary history, metabolic limitations, and unusual ribosome compositions.
- Christopher T. Brown
- , Laura A. Hug
- & Jillian F. Banfield
-
Article |
 Complex archaea that bridge the gap between prokaryotes and eukaryotes
Complex archaea that bridge the gap between prokaryotes and eukaryotes
This study identifies a clade of archaea that is the immediate sister group of eukaryotes in phylogenetic analyses, and that also has a repertoire of proteins otherwise characteristic of eukaryotes—proteins that would have provided the first eukaryotes with a ‘starter kit’ for the genomic and cellular complexity characteristic of the eukaryotic cell.
- Anja Spang
- , Jimmy H. Saw
- & Thijs J. G. Ettema
-
Article |
 In vivo genome editing using Staphylococcus aureus Cas9
In vivo genome editing using Staphylococcus aureus Cas9
The physical size of the commonly used Cas9 from Streptococcus pyogenes poses challenges for CRISPR-Cas genome editing systems that use the adeno-associated virus as a delivery vehicle; here, smaller Cas9 orthologues are characterized, and Cas9 from Staphylococcus aureus allowed targeting of the cholesterol regulatory gene Pcsk9 in the mouse liver.
- F. Ann Ran
- , Le Cong
- & Feng Zhang
-
Letter |
 Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile
Precision microbiome reconstitution restores bile acid mediated resistance to Clostridium difficile
A fraction of the intestinal microbiota as precise as a single bacterial species confers infection resistance by synthesizing Clostridium difficile-inhibiting metabolites from host-derived bile salts.
- Charlie G. Buffie
- , Vanni Bucci
- & Eric G. Pamer
-
Article |
 Biogeography and individuality shape function in the human skin metagenome
Biogeography and individuality shape function in the human skin metagenome
Previous work has shown that human skin is home to a rich and varied microbiota; here a metagenomic approach for samples from physiologically diverse body sites illuminates that the skin microbiota, including bacterial, fungal and viral members, is shaped by the local biogeography and yet marked by strong individuality.
- Julia Oh
- , Allyson L. Byrd
- & Julia A. Segre
-
Article |
 Artificial sweeteners induce glucose intolerance by altering the gut microbiota
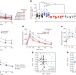
Artificial sweeteners induce glucose intolerance by altering the gut microbiota
Non-caloric artificial sweeteners (NAS), widely used food additives considered to be safe and beneficial alternatives to sugars, are shown here to lead to the development of glucose intolerance through compositional and functional changes in the gut microbiota of mice, and the deleterious metabolic effects are transferred to germ-free mice by faecal transplant; NAS-induced dysbiosis and glucose intolerance are also demonstrated in healthy human subjects.
- Jotham Suez
- , Tal Korem
- & Eran Elinav
-
Letter |
 Viral tagging reveals discrete populations in Synechococcus viral genome sequence space
Viral tagging reveals discrete populations in Synechococcus viral genome sequence space
The metagenome of uncultured, Pacific Ocean viruses linked to a ubiquitous cyanobacteria is characterized using viral-tagging, revealing discrete populations in viral sequence space that includes previously cultivated populations and new populations missed in isolate-based studies.
- Li Deng
- , J. Cesar Ignacio-Espinoza
- & Matthew B. Sullivan
-
Letter |
 Bacterial phylogeny structures soil resistomes across habitats
Bacterial phylogeny structures soil resistomes across habitats
Functional metagenomic selections for resistance to 18 antibiotics in 18 different soils reveal that bacterial community composition is the primary determinant of soil antibiotic resistance gene content.
- Kevin J. Forsberg
- , Sanket Patel
- & Gautam Dantas
-
Letter |
 Abundant SAR11 viruses in the ocean
Abundant SAR11 viruses in the ocean
Viruses are isolated from the SAR11 bacterial clade, the most abundant group of bacteria in the ocean, that were thought to be resistant to viral infection; because of the essential role of SAR11 in carbon cycling these viruses are also an important factor in biogeochemical cycling.
- Yanlin Zhao
- , Ben Temperton
- & Stephen J. Giovannoni
-
Article |
 Genomic variation landscape of the human gut microbiome
Genomic variation landscape of the human gut microbiome
A framework for metagenomic variation analysis to explore variation in the human microbiome is developed; the study describes SNPs, short indels and structural variants in 252 faecal metagenomes of 207 individuals from Europe and North America.
- Siegfried Schloissnig
- , Manimozhiyan Arumugam
- & Peer Bork
-
News & Views |
 Resident risks
Resident risks
An innovative method for probing the genomes of the vast community of microorganisms that inhabit the human gut provides an alternative approach to identifying risk factors for type 2 diabetes. See Letter p.55
- Julia Oh
- & Julia A. Segre
-
Article |
 A metagenome-wide association study of gut microbiota in type 2 diabetes
A metagenome-wide association study of gut microbiota in type 2 diabetes
The authors have developed a new method, metagenome-wide association study (MGWAS), to compare the combined genetic content of the faecal microbiota of healthy people versus patients with type 2 diabetes; they identify multiple microbial species and metabolic pathways that are associated with either cohort and show that some of these may be used as biomarkers.
- Junjie Qin
- , Yingrui Li
- & Jun Wang
-
News & Views |
 Learning about who we are
Learning about who we are
Microbial inhabitants outnumber our body's own cells by about ten to one. These residents have become the subject of intensive research, which is beginning to elucidate their roles in health and disease. See Articles p.207 & p.215
- David A. Relman
-
Article
| Open Access Structure, function and diversity of the healthy human microbiome
Structure, function and diversity of the healthy human microbiome
The Human Microbiome Project Consortium reports the first results of their analysis of microbial communities from distinct, clinically relevant body habitats in a human cohort; the insights into the microbial communities of a healthy population lay foundations for future exploration of the epidemiology, ecology and translational applications of the human microbiome.
- Curtis Huttenhower
- , Dirk Gevers
- & Owen White
-
Article
| Open Access A framework for human microbiome research
A framework for human microbiome research
The Human Microbiome Project Consortium has established a population-scale framework to study a variety of microbial communities that exist throughout the human body, enabling the generation of a range of quality-controlled data as well as community resources.
- Barbara A. Methé
- , Karen E. Nelson
- & Owen White
-
News |
 Microbiome sequencing offers hope for diagnostics
Microbiome sequencing offers hope for diagnostics
Scientists try to avoid the hype that dogs human-genome research.
- Ed Yong
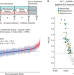
 Bioactive glycans in a microbiome-directed food for children with malnutrition
Bioactive glycans in a microbiome-directed food for children with malnutrition