Article
|
Open Access
Featured
-
-
Article
| Open Access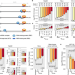 The regulatory impact of RNA-binding proteins on microRNA targeting
The regulatory impact of RNA-binding proteins on microRNA targeting
miRNAs are loaded into Argonaute protein and repress complementary mRNA targets. Here the authors show the unappreciated role of RNA binding proteins for efficient miRNA targeting and expand the current understanding of miRNA targeting.
- Sukjun Kim
- , Soyoung Kim
- & Daehyun Baek
-
Article
| Open Access Sc-compReg enables the comparison of gene regulatory networks between conditions using single-cell data
Sc-compReg enables the comparison of gene regulatory networks between conditions using single-cell data
Changes in cell state underlie the difference between health and disease. Here, the authors propose a computational framework for the integration of gene expression and chromatin-accessibility data from single cells to identify differences in gene regulation in cell types across two conditions.
- Zhana Duren
- , Wenhui Sophia Lu
- & Wing Hung Wong
-
Article
| Open Access Spearheading future omics analyses using dyngen, a multi-modal simulator of single cells
Spearheading future omics analyses using dyngen, a multi-modal simulator of single cells
To benchmark single cell bioinformatics tools, data simulators can provide a robust ground truth. Here the authors present dyngen, a multi-modal simulator, and apply it to aligning cell developmental trajectories, cell-specific regulatory network inference and estimation of RNA velocity.
- Robrecht Cannoodt
- , Wouter Saelens
- & Yvan Saeys
-
Article
| Open Access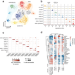 The regulatory landscape of Arabidopsis thaliana roots at single-cell resolution
The regulatory landscape of Arabidopsis thaliana roots at single-cell resolution
Existing studies of the chromatin accessibility, the primary mark of regulatory DNA, in Arabidopsis are based mainly on bulk samples. Here, the authors report the regulatory landscape of Arabidopsis thaliana roots at single-cell resolution.
- Michael W. Dorrity
- , Cristina M. Alexandre
- & Josh T. Cuperus
-
Article
| Open Access Single cell regulatory landscape of the mouse kidney highlights cellular differentiation programs and disease targets
Single cell regulatory landscape of the mouse kidney highlights cellular differentiation programs and disease targets
Epigenetic and transcriptional dynamics are critical for both tissue homeostasis and injury response in the kidney. Leveraging a single cell multiomics atlas of the developing mouse kidney, the authors reveal key events in chromatin regulation and gene expression dynamics during postnatal development.
- Zhen Miao
- , Michael S. Balzer
- & Katalin Susztak
-
Article
| Open Access Defining super-enhancer landscape in triple-negative breast cancer by multiomic profiling
Defining super-enhancer landscape in triple-negative breast cancer by multiomic profiling
Triple-negative breast cancer (TNBC) is an aggressive breast cancer subtype with poor prognostic outcomes. Here the authors characterize super-enhancer heterogeneity and they identify genes that are specifically regulated by TNBC-specific super-enhancers, including FOXC1, MET and ANLN.
- Hao Huang
- , Jianyang Hu
- & Y. Rebecca Chin
-
Article
| Open Access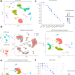 Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney
Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney
Single cell transcriptomic and epigenomic sequencing of human kidney highlight diverse cell types and states. These findings help characterize a novel population of injured proximal tubule cells and illustrate the power of multi-omic approaches to characterizing human tissue.
- Yoshiharu Muto
- , Parker C. Wilson
- & Benjamin D. Humphreys
-
Article
| Open Access A rationally engineered decoder of transient intracellular signals
A rationally engineered decoder of transient intracellular signals
Cells encode information by modulating signal dynamics. Here the authors developed a rapid prototyping tool, TopoDesign, to engineer a synthetic short-pulse decoder in yeast.
- Claude Lormeau
- , Fabian Rudolf
- & Jörg Stelling
-
Article
| Open Access A computer-guided design tool to increase the efficiency of cellular conversions
A computer-guided design tool to increase the efficiency of cellular conversions
Transcription factor over-expression-based cellular conversion methods often endure low conversion efficiency. Here the authors show how to increase conversion efficiency by combining a computational method for prioritizing more efficient TF combinations with a transposon-based genomic integration system for delivery.
- Sascha Jung
- , Evan Appleton
- & Antonio del Sol
-
Article
| Open Access PGC1/PPAR drive cardiomyocyte maturation at single cell level via YAP1 and SF3B2
PGC1/PPAR drive cardiomyocyte maturation at single cell level via YAP1 and SF3B2
Cardiomyocyte maturation and the acquisition of phenotypes is poorly understood at the single cell level. Here, the authors analyse the transcriptome of single cells from neonatal to adult heart and reveal that peroxisome proliferator-activated receptor coactivator-1 mediates the phenotypic shift.
- Sean A. Murphy
- , Matthew Miyamoto
- & Chulan Kwon
-
Article
| Open Access Single-cell transcriptional changes associated with drug tolerance and response to combination therapies in cancer
Single-cell transcriptional changes associated with drug tolerance and response to combination therapies in cancer
It has been proposed that resistance to targeted therapies in non-small cell lung carcinoma (NSCLC) is due to a nonhomogeneous cell population. Here the authors analyse preclinical NSCLC models using single-cell RNA-seq and identify drug tolerant cell states and subpopulations, as well as associated genes.
- Alexandre F. Aissa
- , Abul B. M. M. K. Islam
- & Elizaveta V. Benevolenskaya
-
Article
| Open Access Integrative pan cancer analysis reveals epigenomic variation in cancer type and cell specific chromatin domains
Integrative pan cancer analysis reveals epigenomic variation in cancer type and cell specific chromatin domains
The NCI-60 cancer cell line panel covers multiple cancer types but has not been extensively investigated at the epigenetic level. Here, the authors present H3K4me3, H3K27ac, H3K9me3, and H4K20me3 ChIP-Seq analysis of the cell lines, and describe features of chromatin states and integrative analyses of expression, epigenetic and genetic mutation data.
- Lijin K. Gopi
- & Benjamin L. Kidder
-
Article
| Open Access Causal network models of SARS-CoV-2 expression and aging to identify candidates for drug repurposing
Causal network models of SARS-CoV-2 expression and aging to identify candidates for drug repurposing
Given the severity of the SARS-CoV-2 pandemic, a major challenge is to rapidly repurpose existing approved drugs for clinical interventions. Here, the authors identify robust druggable protein targets within a principled causal framework that makes use of multiple data modalities and integrates aging signatures.
- Anastasiya Belyaeva
- , Louis Cammarata
- & Caroline Uhler
-
Article
| Open Access LETR1 is a lymphatic endothelial-specific lncRNA governing cell proliferation and migration through KLF4 and SEMA3C
LETR1 is a lymphatic endothelial-specific lncRNA governing cell proliferation and migration through KLF4 and SEMA3C
Long noncoding RNAs regulate tissue-specific gene expression. Here the authors profile lineage-specific lncRNAs in human dermal lymphatic and blood vascular endothelial cells (LECs and BECs) and show a role of LEC-specific lncRNA, LETR1, in cell proliferation and migration.
- Luca Ducoli
- , Saumya Agrawal
- & Michael Detmar
-
Article
| Open Access RUNX1/RUNX1T1 mediates alternative splicing and reorganises the transcriptional landscape in leukemia
RUNX1/RUNX1T1 mediates alternative splicing and reorganises the transcriptional landscape in leukemia
The fusion gene RUNX1/RUNX1T1 is oncogenic in acute myeloid leukemia. Here, the authors show that the fusion gene alters the transcriptional landscape of the cells by changing the structure of the 5’UTR, altering isoform expression, and controlling the expression of splicing factors.
- Vasily V. Grinev
- , Farnaz Barneh
- & Olaf Heidenreich
-
Article
| Open Access Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Here, the authors introduce Cell Heterogeneity–Adjusted cLonal Methylation (CHALM) as a methylation quantification method that considers the heterogeneity of sequenced bulk cells. They apply CHALM to methylation datasets to detect differentially methylated genes that exhibit distinct biological functions supporting underlying mechanisms.
- Jianfeng Xu
- , Jiejun Shi
- & Wei Li
-
Article
| Open Access Single cell transcriptomic analysis of human pluripotent stem cell chondrogenesis
Single cell transcriptomic analysis of human pluripotent stem cell chondrogenesis
Application of human induced pluripotent stem cells (hiPSCs) for tissue regeneration is hindered by off-target cell differentiation. Here, the authors use bulk and single cell RNA-sequencing to identify WNT and MITF as off-target hubs during chondrogenic differentiation; inhibiting these pathways enhanced homogeneity and yield.
- Chia-Lung Wu
- , Amanda Dicks
- & Farshid Guilak
-
Article
| Open Access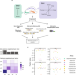 Differential contribution of transcriptomic regulatory layers in the definition of neuronal identity
Differential contribution of transcriptomic regulatory layers in the definition of neuronal identity
Post-transcriptional gene regulation is an important contributor to cell type-specific differences at the transcriptomic level. Here, the authors use a multiomics approach to characterize neuronal diversity in the mouse nervous system, analyzing the relative contributions of multiple layers of transcriptomic regulation in the specification of cell type identity.
- Kevin C. H. Ha
- , Timothy Sterne-Weiler
- & Benjamin J. Blencowe
-
Article
| Open Access Machine learning uncovers independently regulated modules in the Bacillus subtilis transcriptome
Machine learning uncovers independently regulated modules in the Bacillus subtilis transcriptome
The systems-level regulatory structure underlying gene expression in bacteria can be inferred using machine learning algorithms. Here we show this structure for Bacillus subtilis, present five hypotheses gleaned from it, and analyse the process of sporulation from its perspective.
- Kevin Rychel
- , Anand V. Sastry
- & Bernhard O. Palsson
-
Article
| Open Access Deep learning suggests that gene expression is encoded in all parts of a co-evolving interacting gene regulatory structure
Deep learning suggests that gene expression is encoded in all parts of a co-evolving interacting gene regulatory structure
Regulatory and coding regions of genes are shaped by evolution to control expression levels. Here, the authors use deep learning to identify rules controlling gene expression levels and suggest that all parts of the gene regulatory structure interact in this.
- Jan Zrimec
- , Christoph S. Börlin
- & Aleksej Zelezniak
-
Article
| Open Access Transcription-dependent cohesin repositioning rewires chromatin loops in cellular senescence
Transcription-dependent cohesin repositioning rewires chromatin loops in cellular senescence
Senescence is a state of stable proliferative arrest. Here, the authors perform Hi-C analysis on oncogenic RAS-induced senescence in human fibroblasts and characterize the changes in the 3D genome folding associated with the senescence-specific gene expression profile, which are mediated in part through cohesin redistribution on chromatin.
- Ioana Olan
- , Aled J. Parry
- & Masashi Narita
-
Article
| Open Access Interpretation of morphogen gradients by a synthetic bistable circuit
Interpretation of morphogen gradients by a synthetic bistable circuit
Morphogen gradients can be dynamic and transient yet give rise to stable cellular patterns. Here the authors show that a synthetic morphogen-induced mutual inhibition circuit produces stable boundaries when the spatial average of morphogens falls within the region of bistability.
- Paul K. Grant
- , Gregory Szep
- & Andrew Phillips
-
Article
| Open Access Network-based machine learning in colorectal and bladder organoid models predicts anti-cancer drug efficacy in patients
Network-based machine learning in colorectal and bladder organoid models predicts anti-cancer drug efficacy in patients
Cancer patient classification using predictive biomarkers for anti-cancer drug responses is essential for improving therapeutic outcomes. Here, the authors present a machine-learning framework to identify robust drug biomarkers by taking advantage of network-based analyses using pharmacogenomic data.
- JungHo Kong
- , Heetak Lee
- & Sanguk Kim
-
Article
| Open Access Nuclear gene proximity and protein interactions shape transcript covariations in mammalian single cells
Nuclear gene proximity and protein interactions shape transcript covariations in mammalian single cells
Gene expression covariation can be studied by single-cell RNA sequencing. Here the authors analyze intrinsically covarying gene pairs by eliminating the confounding effects in single-cell experiments and observe covariation of proximal genes and miRNA-induced covariation of target mRNAs.
- Marcel Tarbier
- , Sebastian D. Mackowiak
- & Marc R. Friedländer
-
Article
| Open Access Reconstructing the maize leaf regulatory network using ChIP-seq data of 104 transcription factors
Reconstructing the maize leaf regulatory network using ChIP-seq data of 104 transcription factors
Transcriptional factors (TFs) bind in a combinatorial fashion to specify the on-and-off states of genes in a complex and redundant regulatory network. Here, the authors construct the transcription regulatory network in maize leaf using 104 TFs ChIP-seq data and train machine learning models to predict TF binding and colocalization.
- Xiaoyu Tu
- , María Katherine Mejía-Guerra
- & Silin Zhong
-
Article
| Open Access Integrative genomics identifies a convergent molecular subtype that links epigenomic with transcriptomic differences in autism
Integrative genomics identifies a convergent molecular subtype that links epigenomic with transcriptomic differences in autism
Autism spectrum disorder (ASD) is a neurodevelopmental disorder characterized by impaired social interactions with repetitive and restrictive behaviours. Here the authors integrate mRNA expression, miRNA expression, DNA methylation, and histone acetylation datasets from a collection of post mortem brain tissues and identify a convergent molecular subtype of ASD.
- Gokul Ramaswami
- , Hyejung Won
- & Daniel H. Geschwind
-
Article
| Open Access Multiplatform genomic profiling and magnetic resonance imaging identify mechanisms underlying intratumor heterogeneity in meningioma
Multiplatform genomic profiling and magnetic resonance imaging identify mechanisms underlying intratumor heterogeneity in meningioma
Meningiomas are heterogeneous tumours. Here, the authors analysed genetic, epigenetic, and transcriptomic features across spatially-distinct meningioma samples to identify molecular programs underlying tumorigenesis that can be detected preoperatively using magnetic resonance imaging.
- Stephen T. Magill
- , Harish N. Vasudevan
- & David R. Raleigh
-
Article
| Open Access Amalgamated cross-species transcriptomes reveal organ-specific propensity in gene expression evolution
Amalgamated cross-species transcriptomes reveal organ-specific propensity in gene expression evolution
Multicellularity requires complex coordinated gene expression. Fukushima and Pollock find that gene expression in different organs is likely to constrain future patterns of gene expression evolution, particularly following gene duplication.
- Kenji Fukushima
- & David D. Pollock
-
Article
| Open Access A clustering-independent method for finding differentially expressed genes in single-cell transcriptome data
A clustering-independent method for finding differentially expressed genes in single-cell transcriptome data
How cell clusters are defined in single-cell sequencing data has important consequences for downstream analyses and the interpretation of results, but is often not straightforward. Here, the authors present a new approach that enables the prediction of differentially expressed genes without relying on explicit clustering of cells.
- Alexis Vandenbon
- & Diego Diez
-
Article
| Open Access An integrative ENCODE resource for cancer genomics
An integrative ENCODE resource for cancer genomics
ENCODE is a resource comprising thousands of functional genomic datasets. Here, the authors present custom annotation within ENCODE for cancer, highlighting a workflow that can help prioritise key elements in oncogenesis.
- Jing Zhang
- , Donghoon Lee
- & Mark Gerstein
-
Article
| Open Access A supervised learning framework for chromatin loop detection in genome-wide contact maps
A supervised learning framework for chromatin loop detection in genome-wide contact maps
Predicting chromatin loops from genome-wide interaction matrices such as Hi-C data provides insight into gene regulation events. Here, the authors present Peakachu, a Random Forest classification framework that predicts chromatin loops from genome-wide contact maps, and apply it to systematically predict chromatin loops in 56 Hi-C datasets, with results available at the 3D Genome Browser.
- Tarik J. Salameh
- , Xiaotao Wang
- & Feng Yue
-
Article
| Open Access Intron retention is a hallmark and spliceosome represents a therapeutic vulnerability in aggressive prostate cancer
Intron retention is a hallmark and spliceosome represents a therapeutic vulnerability in aggressive prostate cancer
Dysregulation of mRNA alternative splicing is prevalent in cancers. Here, the authors characterized the landscape of aberrant alternative splicing during the development of prostate cancer, progression and therapeutic resistance and show that splicing modulator, E7107, reduces growth in castration-resistant prostate cancer.
- Dingxiao Zhang
- , Qiang Hu
- & Dean G. Tang
-
Article
| Open Access Dimensionality reduction by UMAP to visualize physical and genetic interactions
Dimensionality reduction by UMAP to visualize physical and genetic interactions
Dimensionality reduction is often used to visualize expression profiling data in order to find relationships among cells. Here, the authors use Uniform Manifold Approximation and Projection (UMAP) on published expression data of gene deletions of S. cerevisiae to find novel protein interactions.
- Michael W. Dorrity
- , Lauren M. Saunders
- & Cole Trapnell
-
Article
| Open Access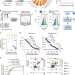 Pan-cancer analysis reveals cooperativity of both strands of microRNA that regulate tumorigenesis and patient survival
Pan-cancer analysis reveals cooperativity of both strands of microRNA that regulate tumorigenesis and patient survival
5p and 3p miRNA strands have different mRNA-targeting sequences and may both functionally impact gene expression in cancer. Here, the authors undertake a pan-cancer analysis that indicates 5p/3p miRNA strands function together to regulate tumorigenic processes.
- Ramkrishna Mitra
- , Clare M. Adams
- & Christine M. Eischen
-
Article
| Open Access In silico prediction of high-resolution Hi-C interaction matrices
In silico prediction of high-resolution Hi-C interaction matrices
Existing computational approaches to predict long-range regulatory interactions do not fully exploit high-resolution Hi-C datasets. Here the authors present a Random Forests regression-based approach to predict high-resolution Hi-C counts using one-dimensional regulatory genomic signals.
- Shilu Zhang
- , Deborah Chasman
- & Sushmita Roy
-
Article
| Open Access Stem-cell-ubiquitous genes spatiotemporally coordinate division through regulation of stem-cell-specific gene networks
Stem-cell-ubiquitous genes spatiotemporally coordinate division through regulation of stem-cell-specific gene networks
Stem-cell-specific genes regulate processes such as maintenance, identity and/or division. Here, the authors show that in the Arabidopsis root TCX2, a gene expressed across different stem cell populations (a stem-cell-ubiquitous gene), controls division and identity by regulating stem-cell-type-specific networks.
- Natalie M. Clark
- , Eli Buckner
- & Rossangela Sozzani
-
Article
| Open Access The Escherichia coli transcriptome mostly consists of independently regulated modules
The Escherichia coli transcriptome mostly consists of independently regulated modules
Mechanistic insight into the regulation of transcriptional modules remains scarce. Here, the authors identify statistically independent gene sets by applying independent component analysis to a high-quality E. coli RNA-seq data compendium and find that most gene sets represent the effects of specific transcriptional regulators.
- Anand V. Sastry
- , Ye Gao
- & Bernhard O. Palsson
-
Article
| Open Access Single-cell RNA-sequencing of herpes simplex virus 1-infected cells connects NRF2 activation to an antiviral program
Single-cell RNA-sequencing of herpes simplex virus 1-infected cells connects NRF2 activation to an antiviral program
Herpes simplex virus 1 (HSV-1) induces major reprogramming of the host cell environment. Here, the authors profile the transcriptomes of individual HSV-1-infected human fibroblasts and provide evidence of an antiviral program mediated by the activation of the host transcription factor NRF2.
- Emanuel Wyler
- , Vedran Franke
- & Markus Landthaler
-
Article
| Open Access IRF2 is a master regulator of human keratinocyte stem cell fate
IRF2 is a master regulator of human keratinocyte stem cell fate
Epidermal homeostasis requires long term stem cell function. Here, the authors apply transcriptional circuitry analysis based on integrated epigenomic profiling of primary human keratinocytes with high and low stem cell function to identify IRF2 as a negative regulator of stemness.
- Nicolas Mercado
- , Gabi Schutzius
- & Susan Kirkland
-
Article
| Open Access Chromatin-informed inference of transcriptional programs in gynecologic and basal breast cancers
Chromatin-informed inference of transcriptional programs in gynecologic and basal breast cancers
Epigenomic data on chromatin accessibility and transcription factor occupancy can reveal enhancer landscapes in cancer. Here, the authors develop a computational strategy called PSIONIC (patient-specific inference of networks informed by chromatin) to model the impact of enhancers on transcriptional programs in gynecologic and basal breast cancers.
- Hatice U. Osmanbeyoglu
- , Fumiko Shimizu
- & Christina S. Leslie
-
Article
| Open Access Cancer-associated mutations in DICER1 RNase IIIa and IIIb domains exert similar effects on miRNA biogenesis
Cancer-associated mutations in DICER1 RNase IIIa and IIIb domains exert similar effects on miRNA biogenesis
DICER is involved in the processing of miRNAs, where the RNase IIIa and IIIb domains are thought to cut the 3p and 5p hairpin arms, respectively. Here, in endometrial cancer, the authors identify an RNase IIIa mutation, which phenocopies mutations in the RNase IIIb domain.
- Jeffrey Vedanayagam
- , Walid K. Chatila
- & Eric C. Lai
-
Article
| Open Access Pluripotency reprogramming by competent and incompetent POU factors uncovers temporal dependency for Oct4 and Sox2
Pluripotency reprogramming by competent and incompetent POU factors uncovers temporal dependency for Oct4 and Sox2
Oct4, along with Sox2 and Klf4 can induce pluripotency, but structurally similar factors like Oct6 cannot. Here, using pluripotency competent and incompetent factors, the authors show that Sox2 plays a dominant role in facilitating chromatin opening at Oct4 bound DNA early during reprogramming to pluripotency.
- Vikas Malik
- , Laura V. Glaser
- & Ralf Jauch
-
Article
| Open Access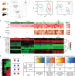 Defective HNF4alpha-dependent gene expression as a driver of hepatocellular failure in alcoholic hepatitis
Defective HNF4alpha-dependent gene expression as a driver of hepatocellular failure in alcoholic hepatitis
Alcoholic hepatitis, a common cause of liver failure, lacks effective treatment. Here, the authors show altered hepatic HNF4a isoform expression and hypermethylation of its target genes in patients. HNF4a dysregulation is improved in vitro by TGFb or PPARg modulation suggesting potential therapeutic avenues.
- Josepmaria Argemi
- , Maria U. Latasa
- & Ramon Bataller
-
Article
| Open Access Chemical genomics reveals histone deacetylases are required for core regulatory transcription
Chemical genomics reveals histone deacetylases are required for core regulatory transcription
Core regulatory transcription factors are usually regulated by cell-type specific super enhancers (SEs). Here, the authors screen for chemical probes able to distinguish between SE-driven and promoter-driven transcription and find that histone deacetylases are selectively required for core regulatory transcription.
- Berkley E. Gryder
- , Lei Wu
- & Javed Khan
-
Article
| Open Access Feed-forward regulation adaptively evolves via dynamics rather than topology when there is intrinsic noise
Feed-forward regulation adaptively evolves via dynamics rather than topology when there is intrinsic noise
Feed‐forward loops (FFLs) can filter out noise, but whether their overrepresentation in GRNs reflects adaptive evolution for this function is debated. Here, the authors develop a null model of regulatory evolution and find that FFLs evolve readily under selection for the noise filtering function.
- Kun Xiong
- , Alex K. Lancaster
- & Joanna Masel
-
Article
| Open Access Optimal control nodes in disease-perturbed networks as targets for combination therapy
Optimal control nodes in disease-perturbed networks as targets for combination therapy
Synergistic interactions may arise between regulators in complex molecular networks. Here, the authors develop OptiCon, a computational method for de novo identification of synergistic key regulators and investigate their potential roles as candidate targets for combination therapy.
- Yuxuan Hu
- , Chia-hui Chen
- & Kai Tan
-
Article
| Open Access Dynamic control of enhancer activity drives stage-specific gene expression during flower morphogenesis
Dynamic control of enhancer activity drives stage-specific gene expression during flower morphogenesis
Enhancer elements can control spatial and temporal patterns of gene expression. Here the authors profile chromatin accessibility, histone modifications and gene expression during Arabidopsis flower development providing evidence for sets of distal enhancers acting in a highly stage-specific manner.
- Wenhao Yan
- , Dijun Chen
- & Kerstin Kaufmann
-
Article
| Open Access Competitive endogenous RNA is an intrinsic component of EMT regulatory circuits and modulates EMT
Competitive endogenous RNA is an intrinsic component of EMT regulatory circuits and modulates EMT
Competitive endogenous RNAs help to regulate biological processes by regulating miRNA activity levels. Here the author show TGFBI acts as a ceRNA for miR-21 in the epithelial-to-mesenchymal transition.
- Yuwei Liu
- , Mengzhu Xue
- & Peng Wang
-
Article
| Open Access Network Walking charts transcriptional dynamics of nitrogen signaling by integrating validated and predicted genome-wide interactions
Network Walking charts transcriptional dynamics of nitrogen signaling by integrating validated and predicted genome-wide interactions
Temporal control of transcriptional networks enables organisms to adapt to changing environment. Here, the authors use a scaled-up cell-based assay to identify direct targets of nitrogen-early responsive transcription factors and validate a network path mediating dynamic nitrogen signaling in Arabidopsis.
- Matthew D. Brooks
- , Jacopo Cirrone
- & Gloria M. Coruzzi
 The basis of easy controllability in Boolean networks
The basis of easy controllability in Boolean networks