Article
|
Open Access
Featured
-
-
Article
| Open Access Transient loss of Polycomb components induces an epigenetic cancer fate
Transient loss of Polycomb components induces an epigenetic cancer fate
A transient perturbation of transcriptional silencing mediated by Polycomb proteins is sufficient to induce an epigenetic cancer cell fate in Drosophila in the absence of driver mutations.
- V. Parreno
- , V. Loubiere
- & G. Cavalli
-
Article
| Open Access Transient naive reprogramming corrects hiPS cells functionally and epigenetically
Transient naive reprogramming corrects hiPS cells functionally and epigenetically
A new reprogramming strategy used to produce human induced pluripotent stem cells from somatic cells results in epigenetic and functional profiles that are highly similar to those of human embryonic stem cells.
- Sam Buckberry
- , Xiaodong Liu
- & Ryan Lister
-
Article |
 Glucose-driven TOR–FIE–PRC2 signalling controls plant development
Glucose-driven TOR–FIE–PRC2 signalling controls plant development
Glucose signalling via TOR controls growth and differentiation through regulation of genome-wide histone methylation via FERTILIZATION-INDEPENDENT ENDOSPERM (FIE).
- Ruiqiang Ye
- , Meiyue Wang
- & Jen Sheen
-
Review Article |
 Inflammatory memory and tissue adaptation in sickness and in health
Inflammatory memory and tissue adaptation in sickness and in health
A Review on inflammatory memory in non-immune cells of different epithelia and neurons, and the potential mechanisms controlling these epigenetic memories and their implications in human health and disease.
- Shruti Naik
- & Elaine Fuchs
-
Article
| Open Access HP1 drives de novo 3D genome reorganization in early Drosophila embryos
HP1 drives de novo 3D genome reorganization in early Drosophila embryos
The heterochromatin protein HP1 has an essential role in establishing several features of the 3D nuclear organization of the genome during early embryonic development in Drosophila.
- Fides Zenk
- , Yinxiu Zhan
- & Nicola Iovino
-
Article |
 N6-methyladenine in DNA antagonizes SATB1 in early development
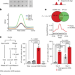
N6-methyladenine in DNA antagonizes SATB1 in early development
The DNA modification N6-methyladenine regulates gene expression during mouse trophoblast development by depositing at the boundaries of active chromatin and preventing its spread by antagonizing the chromatin organizer SATB1.
- Zheng Li
- , Shuai Zhao
- & Andrew Z. Xiao
-
Article |
 poly(UG)-tailed RNAs in genome protection and epigenetic inheritance
poly(UG)-tailed RNAs in genome protection and epigenetic inheritance
In Caenorhabditis elegans, the ribonucleotidyltransferase RDE-3 adds alternating uridine and guanosine ribonucleotides to the 3′ termini of RNAs, a key step in RNA interference and thus epigenetic inheritance in the C. elegans germline.
- Aditi Shukla
- , Jenny Yan
- & Scott Kennedy
-
Article |
 Parental-to-embryo switch of chromosome organization in early embryogenesis
Parental-to-embryo switch of chromosome organization in early embryogenesis
Single-cell allelic HiC analysis, combined with allelic gene expression and chromatin states, reveals parent-of-origin-specific dynamics of chromosome organization and gene expression during mouse preimplantation development.
- Samuel Collombet
- , Noémie Ranisavljevic
- & Edith Heard
-
Article |
 Multi-omics profiling of mouse gastrulation at single-cell resolution
Multi-omics profiling of mouse gastrulation at single-cell resolution
Single-cell mapping of chromatin accessibility, DNA methylation and RNA expression during gastrulation in mouse embryos shows characteristic epigenetic changes that accompany formation of the primary germ layers.
- Ricard Argelaguet
- , Stephen J. Clark
- & Wolf Reik
-
Letter |
 Maternal vitamin C regulates reprogramming of DNA methylation and germline development
Maternal vitamin C regulates reprogramming of DNA methylation and germline development
The effect of vitamin C deprivation on mouse germline development is analysed, revealing that maternal vitamin C is required for proper DNA demethylation and the development of fetal oocytes, whereas the lack of vitamin C during pregnancy leads to reduced female fecundity in the offspring.
- Stephanie P. DiTroia
- , Michelle Percharde
- & Miguel Ramalho-Santos
-
Letter |
 Genome–lamina interactions are established de novo in the early mouse embryo
Genome–lamina interactions are established de novo in the early mouse embryo
Spatial genome organization into lamina-associated domains is first established in the mouse zygote immediately after fertilization without inheritance from the maternal germline—with the paternal and maternal pronucleus exhibiting different organization, which subsequently converges prior to implantation of the embryo.
- Máté Borsos
- , Sara M. Perricone
- & Jop Kind
-
Letter |
 Chromatin analysis in human early development reveals epigenetic transition during ZGA
Chromatin analysis in human early development reveals epigenetic transition during ZGA
By applying an optimized ATAC-seq protocol to human early embryos, distinct accessible chromatin landscapes are found before and after zygotic genome activation, revealing a marked epigenetic transition during zygotic genome activation and putative regulatory elements wiring human early development.
- Jingyi Wu
- , Jiawei Xu
- & Yingpu Sun
-
Analysis
| Open Access Adolescence and the next generation
Adolescence and the next generation
Investing in adolescents as the parents of the next generation is important for the wellbeing of current and future generations.
- George C. Patton
- , Craig A. Olsson
- & Susan M. Sawyer
-
Letter |
 Derivation of ground-state female ES cells maintaining gamete-derived DNA methylation
Derivation of ground-state female ES cells maintaining gamete-derived DNA methylation
Derivation of female mouse embryonic stem cells under certain conditions induces a loss of DNA methylation and erasure of genomic imprints, which are not recovered and that may contribute to observed impaired development.
- Masaki Yagi
- , Satoshi Kishigami
- & Yasuhiro Yamada
-
Article |
 Maternal H3K27me3 controls DNA methylation-independent imprinting
Maternal H3K27me3 controls DNA methylation-independent imprinting
Analysis of parental allele-specific chromatin accessibility genome-wide in mouse zygotes and morula embryos, and investigation of the epigenetic mechanisms underlying these allelic sites, identifying maternal H3K27me3 as a DNA methylation-independent mechanism for genomic imprinting.
- Azusa Inoue
- , Lan Jiang
- & Yi Zhang
-
Letter |
 Pioneer factors govern super-enhancer dynamics in stem cell plasticity and lineage choice
Pioneer factors govern super-enhancer dynamics in stem cell plasticity and lineage choice
An analysis of mouse skin reveals that super-enhancers are critical to identity, lineage commitment and plasticity of adult stem cells; dynamic super-enhancer remodelling in new niches is dependent on the levels of pioneer transcription factor SOX9, which is identified as a key regulator of super-enhancer chromatin for hair follicle stem cells.
- Rene C. Adam
- , Hanseul Yang
- & Elaine Fuchs
-
Article |
 Deconstructing transcriptional heterogeneity in pluripotent stem cells
Deconstructing transcriptional heterogeneity in pluripotent stem cells
This study uses single-cell expression profiling of pluripotent stem cells after various perturbations, and uncovers a high degree of variability that can be inherited through cell divisions—modulating microRNA or external signalling pathways induces a ground state with reduced gene expression heterogeneity and a distinct chromatin profile.
- Roshan M. Kumar
- , Patrick Cahan
- & James J. Collins
-
Letter |
 Epigenetic reprogramming that prevents transgenerational inheritance of the vernalized state
Epigenetic reprogramming that prevents transgenerational inheritance of the vernalized state
The Arabidopsis thaliana floral repressor FLC is epigenetically silenced by prolonged cold in a process called vernalization and then is reactivated before the completion of seed development; a histone demethylase, ELF6, is now shown to be involved in reactivating FLC in reproductive tissues, allowing the resetting of FLC expression and thus the requirement for vernalization in each generation.
- Pedro Crevillén
- , Hongchun Yang
- & Caroline Dean
-
Letter |
 DNA methylation dynamics of the human preimplantation embryo
DNA methylation dynamics of the human preimplantation embryo
Genome-scale DNA methylation maps over early human embryogenesis and embryonic stem cell derivation provide insights into shared and unique modes of regulation when compared to the mouse model, including relationships to gene expression, transposable element activity, and maternal-specific methylation.
- Zachary D. Smith
- , Michelle M. Chan
- & Alexander Meissner
-
Letter |
 Dynamic and static maintenance of epigenetic memory in pluripotent and somatic cells
Dynamic and static maintenance of epigenetic memory in pluripotent and somatic cells
Using a new method to estimate DNA methylation turnover rate, embryonic stem cells are shown to lack clonal transmission of methylation but still maintain a stable epigenetic state, whereas somatic cells transmit methylation clonally but lose epigenetic state coherence owing to the persistence of accumulated methylation errors.
- Zohar Shipony
- , Zohar Mukamel
- & Amos Tanay
-
Article |
 Abnormalities in human pluripotent cells due to reprogramming mechanisms
Abnormalities in human pluripotent cells due to reprogramming mechanisms
Genome-wide analysis of matched human IVF embryonic stem cells (IVF ES cells), induced pluripotent stem cells (iPS cells) and nuclear transfer ES cells (NT ES cells) derived by somatic cell nuclear transfer (SCNT) reveals that human somatic cells can be faithfully reprogrammed to pluripotency by SCNT; NT ES cells and iPS cells derived from the same somatic cells contain comparable numbers of
de novo copy number variations, but whereas DNA methylation and transcriptome profiles of NT ES cells and IVF ES cells are similar, iPS cells have residual patterns typical of parental somatic cells.- Hong Ma
- , Robert Morey
- & Shoukhrat Mitalipov
-
Article |
 Genome-defence small RNAs exapted for epigenetic mating-type inheritance
Genome-defence small RNAs exapted for epigenetic mating-type inheritance
The molecular basis for mating-type determination in the ciliate Paramecium has been elucidated, revealing a novel function for a class of small RNAs — these scnRNAs are typically involved in reprogramming the Paramecium genome during sexual reproduction by recognizing and excising transposable elements, but they are now found to be co-opted to switch off expression of the newly identified mating-type gene mtA by excising its promoter, and to mediate epigenetic inheritance of mating types across sexual generations.
- Deepankar Pratap Singh
- , Baptiste Saudemont
- & Eric Meyer
-
Letter |
 AID stabilizes stem-cell phenotype by removing epigenetic memory of pluripotency genes
AID stabilizes stem-cell phenotype by removing epigenetic memory of pluripotency genes
Fibroblasts deficient in the activation-induced cytidine deaminase (AID) enzyme are shown to fail to stabilize in the pluripotent state, despite initiating the expression of pluripotency genes.
- Ritu Kumar
- , Lauren DiMenna
- & Todd Evans
-
Article |
 Transgenerational epigenetic inheritance of longevity in Caenorhabditis elegans
Transgenerational epigenetic inheritance of longevity in Caenorhabditis elegans
- Eric L. Greer
- , Travis J. Maures
- & Anne Brunet
-
Letter |
 A Polycomb-based switch underlying quantitative epigenetic memory
A Polycomb-based switch underlying quantitative epigenetic memory
- Andrew Angel
- , Jie Song
- & Martin Howard
-
Article |
 Epigenetic memory in induced pluripotent stem cells
Epigenetic memory in induced pluripotent stem cells
Pluripotent stem cells can be generated in the laboratory through somatic cell nuclear transfer (generating nuclear transfer embryonic stem cells, ntESCs) or transcription-factor-based reprogramming (producing induced pluripotent stem cells, iPSCs). These methods reset the methylation signature of the genome — but to what extent? Here it is found that mouse iPSCs 'remember' the methylation status of their tissue of origin, but the methylation of ntESCs is more similar to that of naturally produced ES cells.
- K. Kim
- , A. Doi
- & G. Q. Daley

 Paternal microbiome perturbations impact offspring fitness
Paternal microbiome perturbations impact offspring fitness