Article
|
Open Access
Featured
-
-
Article
| Open Access A combined experimental and modelling approach for the Weimberg pathway optimisation
A combined experimental and modelling approach for the Weimberg pathway optimisation
Metabolic engineering is often hampered by non-linear kinetics and allosteric regulatory mechanisms. Here, the authors construct a quantitative model for the pentose degradation Weimberg pathway in Caulobacter crescentus and demonstrate its biotechnological applications in cell-free system and standard metabolic engineering.
- Lu Shen
- , Martha Kohlhaas
- & Bettina Siebers
-
Article
| Open Access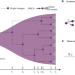 Measuring single cell divisions in human tissues from multi-region sequencing data
Measuring single cell divisions in human tissues from multi-region sequencing data
Quantifying somatic evolutionary processes in cancer and healthy tissue is a challenge. Here, the authors use single time point multi-region sampling of cancer and normal tissue, combined with evolutionary theory, to quantify in vivo mutation and cell survival rates per cell division.
- Benjamin Werner
- , Jack Case
- & Andrea Sottoriva
-
Article
| Open Access Predicting clinical benefit of immunotherapy by antigenic or functional mutations affecting tumour immunogenicity
Predicting clinical benefit of immunotherapy by antigenic or functional mutations affecting tumour immunogenicity
Predicting response to cancer immunotherapy is still a challenge. Here, the authors show that their method of predicting MHC-binding peptides, combined with profiling anti-immunogenic mutations, can better predict the clinical benefit of immunotherapy.
- Kwoneel Kim
- , Hong Sook Kim
- & Jung Kyoon Choi
-
Article
| Open Access Reconstructing evolutionary trajectories of mutation signature activities in cancer using TrackSig
Reconstructing evolutionary trajectories of mutation signature activities in cancer using TrackSig
Cancers evolve as they progress under differing selective pressures. Here, as part of the ICGC/TCGA Pan-Cancer Analysis of Whole Genomes (PCAWG) Consortium, the authors present the method TrackSig the estimates evolutionary trajectories of somatic mutational processes from single bulk tumour data.
- Yulia Rubanova
- , Ruian Shi
- & Christian von Mering
-
Article
| Open Access Surface protein imputation from single cell transcriptomes by deep neural networks
Surface protein imputation from single cell transcriptomes by deep neural networks
Cell-surface proteins serve as phenotypic cell markers and in many cases are more indicative of cellular function than the transcriptome. Here, the authors introduce a transfer learning framework to impute surface protein abundances from scRNA-seq data.
- Zilu Zhou
- , Chengzhong Ye
- & Nancy R. Zhang
-
Article
| Open Access The ETFL formulation allows multi-omics integration in thermodynamics-compliant metabolism and expression models
The ETFL formulation allows multi-omics integration in thermodynamics-compliant metabolism and expression models
Accounting for the effects of genetic expression in genome-scale metabolic models is challenging. Here, the authors introduce a model formulation that efficiently simulates thermodynamic-compliant fluxes, enzyme and mRNA concentration levels, allowing omics integration and broad analysis of in silico cellular physiology.
- Pierre Salvy
- & Vassily Hatzimanikatis
-
Article
| Open Access Realistic in silico generation and augmentation of single-cell RNA-seq data using generative adversarial networks
Realistic in silico generation and augmentation of single-cell RNA-seq data using generative adversarial networks
Low sample numbers often limit the robustness of analyses in biomedical research. Here, the authors introduce a method to generate realistic scRNA-seq data using GANs that learn gene expression dependencies from complex samples, and show that augmenting spare cell populations improves downstream analyses.
- Mohamed Marouf
- , Pierre Machart
- & Stefan Bonn
-
Article
| Open Access De novo generation of hit-like molecules from gene expression signatures using artificial intelligence
De novo generation of hit-like molecules from gene expression signatures using artificial intelligence
High quality hit identification remains a considerable challenge in de novo drug design. Here, the authors train a generative adversarial network with transcriptome profiles induced by a large set of compounds, enabling it to design molecules that are likely to induce desired expression profiles.
- Oscar Méndez-Lucio
- , Benoit Baillif
- & Joerg Wichard
-
Article
| Open Access In silico prediction of high-resolution Hi-C interaction matrices
In silico prediction of high-resolution Hi-C interaction matrices
Existing computational approaches to predict long-range regulatory interactions do not fully exploit high-resolution Hi-C datasets. Here the authors present a Random Forests regression-based approach to predict high-resolution Hi-C counts using one-dimensional regulatory genomic signals.
- Shilu Zhang
- , Deborah Chasman
- & Sushmita Roy
-
Article
| Open Access Estimating heritability and genetic correlations from large health datasets in the absence of genetic data
Estimating heritability and genetic correlations from large health datasets in the absence of genetic data
Disease heritability and genetic correlations between traits depend on genetics, the environment and their interaction. Here, Jia et al. compute disease prevalence curves and disease embeddings from electronic health records and impute heritability for hundreds of diseases and genetic correlations for thousands of disease pairs.
- Gengjie Jia
- , Yu Li
- & Andrey Rzhetsky
-
Article
| Open Access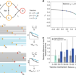 Bridging the gap between efficacy trials and model-based impact evaluation for new tuberculosis vaccines
Bridging the gap between efficacy trials and model-based impact evaluation for new tuberculosis vaccines
One measurement of tuberculosis vaccine efficacy in clinical trials is prevention of disease, but different mechanisms can underlie disease prevention. Here, the authors develop a mathematical model that allows to identify mechanisms of action of a vaccine preventing TB disease.
- Mario Tovar
- , Sergio Arregui
- & Yamir Moreno
-
Article
| Open Access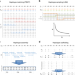 Accurate, scalable and integrative haplotype estimation
Accurate, scalable and integrative haplotype estimation
Haplotype information inferred by phasing is useful in genetic and genomic analysis. Here, the authors develop SHAPEIT4, a phasing method that exhibits sub-linear running time, provides accurate haplotypes and enables integration of external phasing information.
- Olivier Delaneau
- , Jean-François Zagury
- & Emmanouil T. Dermitzakis
-
Article
| Open Access RNA secondary structure prediction using an ensemble of two-dimensional deep neural networks and transfer learning
RNA secondary structure prediction using an ensemble of two-dimensional deep neural networks and transfer learning
The limited availability of high-resolution 3D RNA structures for model training limits RNA secondary structure prediction. Here, the authors overcome this challenge by pre-training a DNN on a large set of predicted RNA structures and using transfer learning with high-resolution structures.
- Jaswinder Singh
- , Jack Hanson
- & Yaoqi Zhou
-
Article
| Open Access Ranking of non-coding pathogenic variants and putative essential regions of the human genome
Ranking of non-coding pathogenic variants and putative essential regions of the human genome
Whole genome sequencing (WGS) holds promise to solve a subset of Mendelian disease cases for which exome sequencing did not provide a genetic diagnosis. Here, Wells et al. report a supervised machine learning model trained on functional, mutational and structural features for rank-scoring and interpreting variants in non-coding regions from WGS.
- Alex Wells
- , David Heckerman
- & Julia di Iulio
-
Article
| Open Access A Bayesian mixture model for the analysis of allelic expression in single cells
A Bayesian mixture model for the analysis of allelic expression in single cells
Allele-specific expression at single-cell resolution can reveal stochastic and dynamic features of gene expression in greater detail. The authors propose scBASE, a soft zero-and-one inflated model that improves estimation of cellular allelic proportions by pooling information across cells.
- Kwangbom Choi
- , Narayanan Raghupathy
- & Gary A. Churchill
-
Article
| Open Access Expanding the limits of the second genetic code with ribozymes
Expanding the limits of the second genetic code with ribozymes
Flexizymes have been used to expand the scope of chemical substrates for ribosome-directed polymerization in vitro. Here the authors deduce design rules of Flexizyme-mediated tRNA acylation that more effectively predict the incorporation of new monomers into peptides.
- Joongoo Lee
- , Kenneth E. Schwieter
- & Michael C. Jewett
-
Article
| Open Access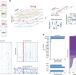 Genomic data integration by WON-PARAFAC identifies interpretable factors for predicting drug-sensitivity in vivo
Genomic data integration by WON-PARAFAC identifies interpretable factors for predicting drug-sensitivity in vivo
Integrative analyses that link molecular data to treatment sensitivity are essential for precision medicine. Here the authors introduce WON-PARAFAC to integrate multiple genomics data to identify sparse and interpretable factors.
- Yongsoo Kim
- , Tycho Bismeijer
- & Daniel J. Vis
-
Article
| Open Access Principles of meiotic chromosome assembly revealed in S. cerevisiae
Principles of meiotic chromosome assembly revealed in S. cerevisiae
During meiotic prophase chromosomes organise into a series of chromatin loops, but the mechanisms of assembly remain unclear. Here the authors use Saccharomyces cerevisiae to elucidate how this elaborate three-dimensional chromosome organisation is linked to genomic sequence, and demonstrate an essential role for cohesin during this process.
- Stephanie A. Schalbetter
- , Geoffrey Fudenberg
- & Matthew J. Neale
-
Article
| Open Access DC3 is a method for deconvolution and coupled clustering from bulk and single-cell genomics data
DC3 is a method for deconvolution and coupled clustering from bulk and single-cell genomics data
Single-cell omics analysis can reveal heterogeneity among individual cells at different levels. Here, the authors develop DC3, a computational method for joint analysis of various bulk and single-cell data from the same heterogeneous cell population.
- Wanwen Zeng
- , Xi Chen
- & Wing Hung Wong
-
Article
| Open Access A tunable dual-input system for on-demand dynamic gene expression regulation
A tunable dual-input system for on-demand dynamic gene expression regulation
Cellular systems have numerous mechanisms to control gene expression. Here the authors build a Tet-On system with conditional destablising elements to regulate gene expression and protein stability, allowing fine modulation of mESC signalling pathways.
- Elisa Pedone
- , Lorena Postiglione
- & Lucia Marucci
-
Article
| Open Access Simultaneous measurement of excitation-contraction coupling parameters identifies mechanisms underlying contractile responses of hiPSC-derived cardiomyocytes
Simultaneous measurement of excitation-contraction coupling parameters identifies mechanisms underlying contractile responses of hiPSC-derived cardiomyocytes
Cardiomyocytes obtained from human induced pluripotent stem cells are increasingly used for drug testing, but they are not always predictive of the heart contractile responses. Here the authors develop a method to measure cytosolic calcium, action potentials and contraction simultaneously, to achieve higher sensitivity for drug screenings.
- Berend J. van Meer
- , Ana Krotenberg
- & Christine L. Mummery
-
Article
| Open Access Single-cell transcriptomics reveals multi-step adaptations to endocrine therapy
Single-cell transcriptomics reveals multi-step adaptations to endocrine therapy
The development of resistance to endocrine therapy is a significant, clinical problem in breast cancer. Here, the authors identify a rare subpopulation of cells that drive resistance following transcriptional reprogramming.
- Sung Pil Hong
- , Thalia E. Chan
- & Luca Magnani
-
Article
| Open Access Changes in gene expression predictably shift and switch genetic interactions
Changes in gene expression predictably shift and switch genetic interactions
Non-additive genetic interactions are plastic and can complicate genetic prediction. Here, using deep mutagenesis of the lambda repressor, Li et al. reveal that changes in gene expression can alter the strength and direction of genetic interactions between mutations in many genes and develop mathematical models for predicting them.
- Xianghua Li
- , Jasna Lalić
- & Ben Lehner
-
Article
| Open Access The pause-initiation limit restricts transcription activation in human cells
The pause-initiation limit restricts transcription activation in human cells
Gene activation requires an increase of successful initiation events. Here, by employing a genome-wide kinetic analysis of transcription, the authors showed that gene activation generally requires a decrease in RNA Polymerase II (Pol II) promoter-proximal pausing while transcription of enhancer elements is not limited by Pol II pausing.
- Saskia Gressel
- , Björn Schwalb
- & Patrick Cramer
-
Article
| Open Access A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism
A consensus S. cerevisiae metabolic model Yeast8 and its ecosystem for comprehensively probing cellular metabolism
Genome-scale metabolic models provide a platform to study metabolism through simulations and analysis of omics data. Here the authors introduce Yeast8 with its model ecosystem, a comprehensive computational resource for simulating the metabolism of Saccharomyces cerevisiae.
- Hongzhong Lu
- , Feiran Li
- & Jens Nielsen
-
Article
| Open Access Cell-type-specific resolution epigenetics without the need for cell sorting or single-cell biology
Cell-type-specific resolution epigenetics without the need for cell sorting or single-cell biology
Compared to bulk data, cell-type-specific DNA methylation data provide higher resolution of epigenetic variation. Here, the authors introduce Tensor Composition Analysis, a novel computational approach for learning cell-type-specific DNA methylation from tissue-level bulk data, and show its application in epigenome-wide association studies.
- Elior Rahmani
- , Regev Schweiger
- & Eran Halperin
-
Article
| Open Access Profiling host ANP32A splicing landscapes to predict influenza A virus polymerase adaptation
Profiling host ANP32A splicing landscapes to predict influenza A virus polymerase adaptation
Polymorphisms in the avian influenza A virus (IAV) polymerase restrict its host range during transmission from birds to mammals. Here, the authors investigate differences in the host chromatin regulator ANP32A regarding IAV polymerase adaptation, and profile ANP32A splicing to predict avian species associated with pre-adaptive human-signatures in the virus.
- Patricia Domingues
- , Davide Eletto
- & Benjamin G. Hale
-
Article
| Open Access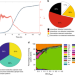 Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli
Regulatory mechanisms underlying coordination of amino acid and glucose catabolism in Escherichia coli
Bacteria must adapt their metabolism in the face of dynamically changing nutrient availability. Here, using their constraint-based modeling approach the authors analyze E. coli exometabolome data during growth in complex medium, revealing temporal coordination of glucose and amino acid catabolism.
- Mattia Zampieri
- , Manuel Hörl
- & Uwe Sauer
-
Article
| Open Access Estimating dispensable content in the human interactome
Estimating dispensable content in the human interactome
The fraction of protein-protein interactions (PPIs) that can be disrupted without fitness effect is unknown. Here, the authors model how disease-causing mutations and common mutations carried by healthy people perturb the interactome, and estimate that <20% of human PPIs are completely dispensable.
- Mohamed Ghadie
- & Yu Xia
-
Article
| Open Access NASC-seq monitors RNA synthesis in single cells
NASC-seq monitors RNA synthesis in single cells
Sequencing of newly synthesised RNA can reveal the transcriptional dynamics in a population of cells. Here the authors develop NASC-seq to bring this sensitivity and temporal resolution to single-cell analysis.
- Gert-Jan Hendriks
- , Lisa A. Jung
- & Rickard Sandberg
-
Comment
| Open AccessUsing “outbreak science” to strengthen the use of models during epidemics
Infectious disease modeling has played a prominent role in recent outbreaks, yet integrating these analyses into public health decision-making has been challenging. We recommend establishing ‘outbreak science’ as an inter-disciplinary field to improve applied epidemic modeling.
- Caitlin Rivers
- , Jean-Paul Chretien
- & Simon Pollett
-
Article
| Open Access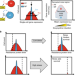 Role of network-mediated stochasticity in mammalian drug resistance
Role of network-mediated stochasticity in mammalian drug resistance
The role of gene expression noise in the evolution of drug resistance in mammalian cells is unclear. Here, by uncoupling noise from mean expression of a drug resistance gene in CHO cells the authors show that noisy expression aids adaptation to high drug levels, but delays it at low drug levels.
- Kevin S. Farquhar
- , Daniel A. Charlebois
- & Gábor Balázsi
-
Article
| Open Access Simulating multiple faceted variability in single cell RNA sequencing
Simulating multiple faceted variability in single cell RNA sequencing
Simulated single cell RNA sequencing data is useful for method development and comparison. Here, the authors developed SymSim, a simulator that explicitly models the main factors of variation in single cell data.
- Xiuwei Zhang
- , Chenling Xu
- & Nir Yosef
-
Article
| Open Access Detection of DNA base modifications by deep recurrent neural network on Oxford Nanopore sequencing data
Detection of DNA base modifications by deep recurrent neural network on Oxford Nanopore sequencing data
DNA modification generates unique electric signals in Oxford Nanopore sequencing data but the signals can be complicated to decipher. Here, the authors develop a deep learning framework, DeepMod, to detect DNA base modifications including 5mC and 6mA using Nanopore sequencing data
- Qian Liu
- , Li Fang
- & Kai Wang
-
Article
| Open Access Optimal control nodes in disease-perturbed networks as targets for combination therapy
Optimal control nodes in disease-perturbed networks as targets for combination therapy
Synergistic interactions may arise between regulators in complex molecular networks. Here, the authors develop OptiCon, a computational method for de novo identification of synergistic key regulators and investigate their potential roles as candidate targets for combination therapy.
- Yuxuan Hu
- , Chia-hui Chen
- & Kai Tan
-
Article
| Open Access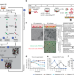 Experimental and computational analyses reveal that environmental restrictions shape HIV-1 spread in 3D cultures
Experimental and computational analyses reveal that environmental restrictions shape HIV-1 spread in 3D cultures
Here, using an integrative experimental and computational approach, Imle et al. show how cell motility and density affect HIV cell-associated transmission in a three-dimensional tissue-like culture system of CD4+ T cells and collagen, and how different collagen matrices restrict infection by cell-free virions.
- Andrea Imle
- , Peter Kumberger
- & Oliver T. Fackler
-
Article
| Open Access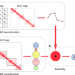 Integrating Hi-C and FISH data for modeling of the 3D organization of chromosomes
Integrating Hi-C and FISH data for modeling of the 3D organization of chromosomes
Methodological advances have increased our understanding of chromatin structure through chromosome conformation capture techniques and high resolution imaging, but integration of these datasets is challenging. Here the authors propose GEM-FISH, a method for reconstructing the 3D models of chromosomes through systematically integrating both Hi-C and FISH data with the prior biophysical knowledge of a polymer model.
- Ahmed Abbas
- , Xuan He
- & Jianyang Zeng
-
Article
| Open Access Microbial coexistence through chemical-mediated interactions
Microbial coexistence through chemical-mediated interactions
The drivers of coexistence between species with different growth rates are of interest in both ecology and applied microbial science. The authors show, via modelling, that species interactions moderated by consumption or degradation of chemicals can allow coexistence.
- Lori Niehaus
- , Ian Boland
- & Babak Momeni
-
Article
| Open Access An additive Gaussian process regression model for interpretable non-parametric analysis of longitudinal data
An additive Gaussian process regression model for interpretable non-parametric analysis of longitudinal data
Longitudinal data are common in biomedical research, but their analysis is often challenging. Here, the authors present an additive Gaussian process regression model specifically designed for statistical analysis of longitudinal experimental data.
- Lu Cheng
- , Siddharth Ramchandran
- & Harri Lähdesmäki
-
Article
| Open Access Mapping vaccination coverage to explore the effects of delivery mechanisms and inform vaccination strategies
Mapping vaccination coverage to explore the effects of delivery mechanisms and inform vaccination strategies
The success of vaccination programs depends largely on the mechanisms used in vaccine delivery. Here, the authors evaluate the relative effectiveness of two major vaccine delivery strategies, namely routine immunization and supplementary immunization activities in five study countries.
- C. Edson Utazi
- , Julia Thorley
- & Andrew J. Tatem
-
Article
| Open Access Memory and relatedness of transcriptional activity in mammalian cell lineages
Memory and relatedness of transcriptional activity in mammalian cell lineages
Phenotypically identical mammalian cells often display considerable variability in transcript levels of individual genes. Here the authors document how different genes propagate their expression levels in cell lineages and suggest a potential role of transcriptional memory for generating spatial patterns of gene expression.
- Nicholas E. Phillips
- , Aleksandra Mandic
- & David M. Suter
-
Article
| Open Access Inter-annual variation in seasonal dengue epidemics driven by multiple interacting factors in Guangzhou, China
Inter-annual variation in seasonal dengue epidemics driven by multiple interacting factors in Guangzhou, China
In 2014 Guangzhou, China experienced its worse dengue epidemic on record. To determine the reasons for this the authors model historical data under combinations of four time-varying factors and find that past epidemics were limited by one or more unfavourable conditions, but the 2014 epidemic faced none of these restraints.
- Rachel J. Oidtman
- , Shengjie Lai
- & Hongjie Yu
-
Article
| Open Access Model selection may not be a mandatory step for phylogeny reconstruction
Model selection may not be a mandatory step for phylogeny reconstruction
Model selection is a time-intensive step of molecular phylogenetic analysis. Here, Abadi, Azouri and colleagues show that all model selection criteria lead to similar inferences, and that for topology and ancestral sequence reconstruction, using the GTR+I+G model is as accurate.
- Shiran Abadi
- , Dana Azouri
- & Itay Mayrose
-
Article
| Open Access On the predictability of infectious disease outbreaks
On the predictability of infectious disease outbreaks
Forecasting of infectious disease outbreaks can inform appropriate intervention measures, but whether fundamental limits to accurate prediction exist is unclear. Here, the authors use permutation entropy as a model independent measure of predictability to study limitations across a broad set of infectious diseases.
- Samuel V. Scarpino
- & Giovanni Petri
-
Article
| Open Access Towards a data-integrated cell
Towards a data-integrated cell
Integration of omics data remains a challenge. Here, the authors introduce iCell, a framework to integrate tissue-specific protein–protein interaction, co-expression and genetic interaction data, enabling identification of the most rewired genes in cancer, unidentifiable in individual data layers.
- Noël Malod-Dognin
- , Julia Petschnigg
- & Nataša Pržulj
-
Article
| Open Access Optimal dosing of dihydroartemisinin-piperaquine for seasonal malaria chemoprevention in young children
Optimal dosing of dihydroartemisinin-piperaquine for seasonal malaria chemoprevention in young children
Seasonal malaria chemoprevention provides substantial benefit for young children, but resistance to used drugs will likely develop. Here, Chotsiri et al. evaluate the use of dihydroartemisinin-piperaquine as a regimen in 179 children, and population-based simulations suggest that small children would benefit from a higher and extended dosage.
- Palang Chotsiri
- , Issaka Zongo
- & Joel Tarning
-
Article
| Open Access Single-cell RNA-seq denoising using a deep count autoencoder
Single-cell RNA-seq denoising using a deep count autoencoder
Single-cell RNA sequencing is a powerful method to study gene expression, but noise in the data can obstruct analysis. Here the authors develop a denoising method based on a deep count autoencoder network that scales linearly with the number of cells, and therefore is compatible with large data sets.
- Gökcen Eraslan
- , Lukas M. Simon
- & Fabian J. Theis
-
Article
| Open Access Image-based modeling of kidney branching morphogenesis reveals GDNF-RET based Turing-type mechanism and pattern-modulating WNT11 feedback
Image-based modeling of kidney branching morphogenesis reveals GDNF-RET based Turing-type mechanism and pattern-modulating WNT11 feedback
Many organs develop through branching morphogenesis, but whether the underlying mechanisms are shared is unknown. Here, the authors show that a ligand-receptor based Turing mechanisms, similar to that observed in lung development, likely underlies branching morphogenesis of the kidney.
- Denis Menshykau
- , Odyssé Michos
- & Dagmar Iber
-
Article
| Open Access Kinetic analysis of multistep USP7 mechanism shows critical role for target protein in activity
Kinetic analysis of multistep USP7 mechanism shows critical role for target protein in activity
Deubiquitinating enzymes (DUBs) are critical regulators of cellular processes by removing ubiquitin from specific targets. Here global kinetic modelling reveals the mechanism by which the low intrinsic activity of USP7 is substantially enhanced on a specific physiological target.
- Robbert Q. Kim
- , Paul P. Geurink
- & Titia K. Sixma

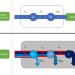 An analytical theory of balanced cellular growth
An analytical theory of balanced cellular growth