Featured
-
-
Article
| Open Access A generative adversarial network model alternative to animal studies for clinical pathology assessment
A generative adversarial network model alternative to animal studies for clinical pathology assessment
Generative AI has the potential to transform the way chemical and drug safety research is conducted. Here the authors show AnimalGAN, a model developed using Generative Adversarial Networks, which simulates virtual animal experiments to generate multidimensional rat clinical pathology measurements.
- Xi Chen
- , Ruth Roberts
- & Weida Tong
-
Article
| Open Access Dynamic characterization and interpretation for protein-RNA interactions across diverse cellular conditions using HDRNet
Dynamic characterization and interpretation for protein-RNA interactions across diverse cellular conditions using HDRNet
Predicting dynamic RNA-RBP interactions in diverse cell lines is an important challenge in unravelling RNA function and post-transcriptional regulatory mechanisms. Here, authors develop HDRNet, an end-to-end deep-learning-based framework for accurately predicting dynamic RBP binding events across various cellular conditions.
- Haoran Zhu
- , Yuning Yang
- & Xiangtao Li
-
Article
| Open Access Deep flanking sequence engineering for efficient promoter design using DeepSEED
Deep flanking sequence engineering for efficient promoter design using DeepSEED
Designing promoters with desired properties is crucial in synthetic biology. Here, authors introduce DeepSEED, an AI-aided flanking sequence optimisation framework which combines expert knowledge with deep learning techniques to efficiently design promoters in both eukaryotic and prokaryotic cells.
- Pengcheng Zhang
- , Haochen Wang
- & Xiaowo Wang
-
Article
| Open Access MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
The authors develop an integrative and scalable framework to eliminate systematic biases and retrieve high-quality metagenome-assembled genomes using either long-read or short-read metagenomic Hi-C data.
- Yuxuan Du
- & Fengzhu Sun
-
Article
| Open Access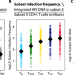 Estimating the contribution of CD4 T cell subset proliferation and differentiation to HIV persistence
Estimating the contribution of CD4 T cell subset proliferation and differentiation to HIV persistence
The authors used mathematical modeling of human data to study how HIV persists despite suppressive antiretroviral therapy. They found that when latently infected CD4+ T cells proliferate or differentiate, they can create HIV DNA and passage it into other subsets. More mature CD4 cell subsets then clear HIV DNA faster.
- Daniel B. Reeves
- , Charline Bacchus-Souffan
- & Peter W. Hunt
-
Article
| Open Access Patient-specific models link neurotransmitter receptor mechanisms with motor and visuospatial axes of Parkinson’s disease
Patient-specific models link neurotransmitter receptor mechanisms with motor and visuospatial axes of Parkinson’s disease
Neurotransmitter receptor distributions help explain structural and functional brain alterations in Parkinson’s disease. Distinct multi-receptor profiles are associated with the severity of motor, and visuospatial, psychiatric and memory symptoms.
- Ahmed Faraz Khan
- , Quadri Adewale
- & Yasser Iturria-Medina
-
Article
| Open Access TAGET: a toolkit for analyzing full-length transcripts from long-read sequencing
TAGET: a toolkit for analyzing full-length transcripts from long-read sequencing
Accurate long-read RNA sequencing facilitates analysis of full-length transcripts. Here the authors develop an integrative toolkit, optimised for Iso-Seq data analysis, that includes transcript alignment, annotation, quantification and gene fusion detection.
- Yuchao Xia
- , Zijie Jin
- & Ruibin Xi
-
Article
| Open Access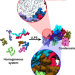 Thermodynamic forces from protein and water govern condensate formation of an intrinsically disordered protein domain
Thermodynamic forces from protein and water govern condensate formation of an intrinsically disordered protein domain
In this work, the authors report atomistic molecular dynamics simulations showing that solvation entropy and protein-protein interactions are the main thermodynamic driving forces for the formation of condensates of the intrinsically disordered domain of the protein FUS.
- Saumyak Mukherjee
- & Lars V. Schäfer
-
Article
| Open Access epiAneufinder identifies copy number alterations from single-cell ATAC-seq data
epiAneufinder identifies copy number alterations from single-cell ATAC-seq data
'Here the authors present epiAneufinder, an algorithm for the identification of single-cell copy number alterations from scATAC-seq data, and explore the clonal heterogeneity in cell populations.
- Akshaya Ramakrishnan
- , Aikaterini Symeonidi
- & Maria Colomé-Tatché
-
Article
| Open Access SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
Protein isoform quantification in shotgun proteomics is challenging due to the mapping of many peptides to multiple protein isoforms. Here, the authors present a computational method SEPepQuant and demonstrate its utility in revealing protein isoform level regulation in shotgun proteomics.
- Yongchao Dou
- , Yuejia Liu
- & Bing Zhang
-
Article
| Open Access The tumor microenvironment shows a hierarchy of cell-cell interactions dominated by fibroblasts
The tumor microenvironment shows a hierarchy of cell-cell interactions dominated by fibroblasts
The tumor microenvironment (TME) is complex and heterogenous, with cancer cells and diverse non-malignant cells interacting with each other. Here the authors define the network of interactions between different cell types in the TME of breast cancer, identifying and characterizing a two-cell circuit of cancer associated fibroblasts and macrophages.
- Shimrit Mayer
- , Tomer Milo
- & Ruth Scherz-Shouval
-
Article
| Open Access Natural plant growth and development achieved in the IPK PhenoSphere by dynamic environment simulation
Natural plant growth and development achieved in the IPK PhenoSphere by dynamic environment simulation
The PhenoSphere is a unique plant cultivation facility in which field-like environments can be simulated. Here, the authors find that a single season simulation is superior to an averaged season and to a climatized glasshouse cultivation to elicit field-like phenotypes evaluated in 11 maize lines.
- Marc C. Heuermann
- , Dominic Knoch
- & Thomas Altmann
-
Article
| Open Access Integrating end-to-end learning with deep geometrical potentials for ab initio RNA structure prediction
Integrating end-to-end learning with deep geometrical potentials for ab initio RNA structure prediction
Here the authors developed an open-source program (DRfold) for RNA tertiary structure prediction from sequence. Through a unique combination of end-to-end learning and geometry restraint guided simulations, the method demonstrates advantage over peer methods.
- Yang Li
- , Chengxin Zhang
- & Yang Zhang
-
Article
| Open Access Performance of tumour microenvironment deconvolution methods in breast cancer using single-cell simulated bulk mixtures
Performance of tumour microenvironment deconvolution methods in breast cancer using single-cell simulated bulk mixtures
Multiple computational approaches have been developed for the deconvolution of cells in the tumour microenvironment (TME) using bulk RNA-seq data. Here, the authors use breast cancer single-cell RNA-seq data to produce simulated bulk data, with which they compare the performance of nine TME deconvolution methods.
- Khoa A. Tran
- , Venkateswar Addala
- & Nicola Waddell
-
Article
| Open Access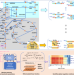 Local flux coordination and global gene expression regulation in metabolic modeling
Local flux coordination and global gene expression regulation in metabolic modeling
Genome-scale metabolic networks (GSMs) are a representation of a cell’s stoichiometrically balanced reactions. Here the authors report Decrem, a GSM reconstruction method, by integrating locally coupled reactions and global transcriptional regulation of metabolism by cell state.
- Gaoyang Li
- , Li Liu
- & Huansheng Cao
-
Article
| Open Access SiGra: single-cell spatial elucidation through an image-augmented graph transformer
SiGra: single-cell spatial elucidation through an image-augmented graph transformer
Recent advances have pushed spatial transcriptomics to subcellular resolution. Here, the authors propose SiGra, a graph artificial intelligence model designed for high-throughput spatial molecular imaging.
- Ziyang Tang
- , Zuotian Li
- & Qianqian Song
-
Article
| Open Access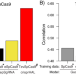 A generalizable Cas9/sgRNA prediction model using machine transfer learning with small high-quality datasets
A generalizable Cas9/sgRNA prediction model using machine transfer learning with small high-quality datasets
Current bacterial sgRNA activity models struggle with accurate predictions and generalizations. Here the authors report crisprHAL, a machine learning architecture that can be trained on existing datasets, and shows good sgRNA activity prediction accuracy can generalize predictions to different bacteria.
- Dalton T. Ham
- , Tyler S. Browne
- & David R. Edgell
-
Article
| Open Access Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Deep neural networks hold significant promise in capturing the complexity of biological systems. However, they suffer from a lack of interpretability. Here, authors present a generalizable method for developing, interpreting, and visualizing biologically informed neural networks for proteomics data.
- Erik Hartman
- , Aaron M. Scott
- & Johan Malmström
-
Article
| Open Access Addressing mechanism bias in model-based impact forecasts of new tuberculosis vaccines
Addressing mechanism bias in model-based impact forecasts of new tuberculosis vaccines
The complex transmission chain of tuberculosis (TB) forces mathematical modelers to make mechanistic assumptions when modelling vaccine effects. Here, authors posit a Bayesian formalism that unlocks mechanism-agnostic impact forecasts for TB vaccines.
- M. Tovar
- , Y. Moreno
- & J. Sanz
-
Article
| Open Access Effects of public-health measures for zeroing out different SARS-CoV-2 variants
Effects of public-health measures for zeroing out different SARS-CoV-2 variants
China maintained a ‘zero-COVID’ policy from early in the pandemic until late 2022 that employed various public health interventions with the aim of COVID-19 containment. Here, the authors use data from 131 outbreaks in China to estimate the effects of a range of interventions against different SARS-CoV-2 variants in diverse settings.
- Yong Ge
- , Xilin Wu
- & Shengjie Lai
-
Article
| Open Access Thermodynamic principle to enhance enzymatic activity using the substrate affinity
Thermodynamic principle to enhance enzymatic activity using the substrate affinity
Currently, there is no well-defined strategy to increase the activity of enzymes. Here, the authors provide mathematical evidence that adjusting the Michaelis-Menten constant to the substrate concentration maximizes enzymatic activity.
- Hideshi Ooka
- , Yoko Chiba
- & Ryuhei Nakamura
-
Article
| Open Access CD98hc is a target for brain delivery of biotherapeutics
CD98hc is a target for brain delivery of biotherapeutics
New delivery platforms are needed to allow broader application of biotherapeutics for CNS diseases. Here, the authors show enhanced CNS delivery with a transport vehicle engineered to bind CD98hc, a highly expressed target at the blood-brain barrier.
- Kylie S. Chew
- , Robert C. Wells
- & Mihalis S. Kariolis
-
Article
| Open Access A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
A deep learning method for replicate-based analysis of chromosome conformation contacts using Siamese neural networks
Siamese neural networks are a powerful deep learning approach for image analysis. Here, the authors adapt this method to the replicate-based analysis of Hi-C data and find that it successfully discriminates technical noise from biological variation.
- Ediem Al-jibury
- , James W. D. King
- & Daniel Rueckert
-
Article
| Open Access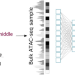 Whole genome deconvolution unveils Alzheimer’s resilient epigenetic signature
Whole genome deconvolution unveils Alzheimer’s resilient epigenetic signature
The authors present a deep learning method that deconvolutes ATAC-seq samples into cell type-specific chromatin accessibility profiles. Applied on 191 samples, the method unveils cell type-specific pathways and nominates potential epigenetic mediators underlying resilience to Alzheimer’s disease.
- Eloise Berson
- , Anjali Sreenivas
- & Thomas J. Montine
-
Article
| Open Access COMPASS: joint copy number and mutation phylogeny reconstruction from amplicon single-cell sequencing data
COMPASS: joint copy number and mutation phylogeny reconstruction from amplicon single-cell sequencing data
Understanding the evolution of a tumor is important for predicting its resistance to treatment. This paper presents a new computational method, COMPASS, for inferring the joint phylogeny of single nucleotide variants and copy number alterations from targeted scDNAseq data.
- Etienne Sollier
- , Jack Kuipers
- & Katharina Jahn
-
Article
| Open Access Fast kernel-based association testing of non-linear genetic effects for biobank-scale data
Fast kernel-based association testing of non-linear genetic effects for biobank-scale data
We have developed FastKAST, a highly-scalable algorithm to identify non-linear genetic effects on complex traits in large datasets. Applied to 300K UK Biobank individuals, we successfully detected significant non-linear effects across 53 traits.
- Boyang Fu
- , Ali Pazokitoroudi
- & Sriram Sankararaman
-
Article
| Open Access Characterizing cancer metabolism from bulk and single-cell RNA-seq data using METAFlux
Characterizing cancer metabolism from bulk and single-cell RNA-seq data using METAFlux
Metabolic reprogramming is a common indicator of the tumour microenvironment. Here the authors develop the METAflux framework to predict metabolic fluxes from single cell RNA-seq data.
- Yuefan Huang
- , Vakul Mohanty
- & Ken Chen
-
Article
| Open Access Cell-type-specific co-expression inference from single cell RNA-sequencing data
Cell-type-specific co-expression inference from single cell RNA-sequencing data
Inferring co-expressions with scRNA-seq data is challenging, and existing methods suffer from inflated false positives and biases. Here, the authors proposed CS-CORE, which yields unbiased estimates and identifies co-expressions that are more reproducible and biologically relevant for scRNA-seq data.
- Chang Su
- , Zichun Xu
- & Jingfei Zhang
-
Article
| Open Access A machine-learning approach to human ex vivo lung perfusion predicts transplantation outcomes and promotes organ utilization
A machine-learning approach to human ex vivo lung perfusion predicts transplantation outcomes and promotes organ utilization
Ex vivo perfusion is a unique platform to study isolated human lungs. Here, authors show that a machine learning model, InsighTx, derived from data generated during ex vivo lung perfusion can accurately predict transplant outcomes and increase organ utilization rates.
- Andrew T. Sage
- , Laura L. Donahoe
- & Shaf Keshavjee
-
Article
| Open Access Proteomics and constraint-based modelling reveal enzyme kinetic properties of Chlamydomonas reinhardtii on a genome scale
Proteomics and constraint-based modelling reveal enzyme kinetic properties of Chlamydomonas reinhardtii on a genome scale
Closing a major gap in photosynthetic metabolic modelling, the authors provide over 500 estimates of in vivo enzyme catalytic rate in C. reinhardtii, which considerably improves predictions on how enzyme mass is allocated to different pathways.
- Marius Arend
- , David Zimmer
- & Zoran Nikoloski
-
Article
| Open Access Experimental validation of the free-energy principle with in vitro neural networks
Experimental validation of the free-energy principle with in vitro neural networks
Empirical applications of the free-energy principle entail a commitment to a particular process theory. Here, the authors reverse engineered generative models from neural responses of in vitro networks and demonstrated that the free-energy principle could predict how neural networks reorganized in response to external stimulation.
- Takuya Isomura
- , Kiyoshi Kotani
- & Karl J. Friston
-
Article
| Open Access SONAR enables cell type deconvolution with spatially weighted Poisson-Gamma model for spatial transcriptomics
SONAR enables cell type deconvolution with spatially weighted Poisson-Gamma model for spatial transcriptomics
Spatial transcriptomics reveal cellular profiles with spatial context. Here the authors present SONAR, a computational model that utilizes spatial information to decipher cell types in tissues and validate on various spatial patterns and fine-mapped cell types in complex tissues.
- Zhiyuan Liu
- , Dafei Wu
- & Liang Ma
-
Article
| Open Access Functional annotation of proteins for signaling network inference in non-model species
Functional annotation of proteins for signaling network inference in non-model species
An artificial-intelligence network is used to generate highly accurate predictions of proteins’ functionality. The predictions on the identity of regulatory proteins is used to create regulatory networks and make discoveries about complex biological systems.
- Lisa Van den Broeck
- , Dinesh Kiran Bhosale
- & Rosangela Sozzani
-
Article
| Open Access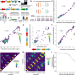 A multiplexed bacterial two-hybrid for rapid characterization of protein–protein interactions and iterative protein design
A multiplexed bacterial two-hybrid for rapid characterization of protein–protein interactions and iterative protein design
Protein-protein interactions (PPIs) are crucial for biological functions and have applications ranging from drug design to synthetic cell circuits. Here the authors develop an assay and computational methods to identify more orthogonal coiled-coil pairs, critical for biological processes and drug design.
- W. Clifford Boldridge
- , Ajasja Ljubetič
- & Sriram Kosuri
-
Article
| Open Access Preventing antimalarial drug resistance with triple artemisinin-based combination therapies
Preventing antimalarial drug resistance with triple artemisinin-based combination therapies
Triple artemisinin-based combination therapies have shown high efficacy for treatment of malaria in preliminary studies. Here, the authors use mathematical modelling to assess whether these therapies could also delay the emergence and spread of antimalarial drug resistance when compared against frontline therapies.
- Tran Dang Nguyen
- , Bo Gao
- & Ricardo Aguas
-
Article
| Open Access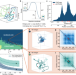 Partition complex structure can arise from sliding and bridging of ParB dimers
Partition complex structure can arise from sliding and bridging of ParB dimers
In many bacteria and plasmids, DNA segregation is controlled by the ParABS system, an essential component of which is the formation of a nucleoprotein complex. Here, making use of recent discoveries, the authors develop a sliding and bridging model to predict the fine structure of this complex.
- Lara Connolley
- , Lucas Schnabel
- & Seán M. Murray
-
Article
| Open Access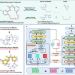 Macrocyclization of linear molecules by deep learning to facilitate macrocyclic drug candidates discovery
Macrocyclization of linear molecules by deep learning to facilitate macrocyclic drug candidates discovery
Macrocyclization of bioactive acyclic molecules provides a potential avenue to yield novel chemical scaffolds with improved pharmacological properties. Here, the authors propose a deep learning based macrocyclization method to generate diverse macrocycles from a given acyclic molecule.
- Yanyan Diao
- , Dandan Liu
- & Honglin Li
-
Article
| Open Access Nutritional redundancy in the human diet and its application in phenotype association studies
Nutritional redundancy in the human diet and its application in phenotype association studies
Studying human diet may help us identify measures to treat or prevent chronic diseases. Here, the authors discover the nutritional redundancy phenomenon in human diet and demonstrate its association with cardiovascular disease and type 2 diabetes.
- Xu-Wen Wang
- , Yang Hu
- & Yang-Yu Liu
-
Article
| Open Access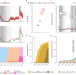 Epidemiological drivers of transmissibility and severity of SARS-CoV-2 in England
Epidemiological drivers of transmissibility and severity of SARS-CoV-2 in England
The COVID-19 pandemic has been characterised by periods of dominance of different SARS-CoV-2 variants. In this mathematical modelling study, the authors investigate the epidemiological properties of successive variants in England until early 2022 and quantify the impacts of control measures.
- Pablo N. Perez-Guzman
- , Edward Knock
- & Marc Baguelin
-
Article
| Open Access Sequence-based drug design as a concept in computational drug design
Sequence-based drug design as a concept in computational drug design
Conventional structure-based drug design pipeline is a complex, human-engineered process with multiple independently optimized steps. Here, the authors report a sequence-to-drug concept that discovers drug-like small molecule modulators directly from protein sequences.
- Lifan Chen
- , Zisheng Fan
- & Mingyue Zheng
-
Article
| Open Access Impact of misclassified defective proviruses on HIV reservoir measurements
Impact of misclassified defective proviruses on HIV reservoir measurements
Quantifying intact proviruses is key to understanding decreases in HIV reservoirs but results can differ depending on the method. To balance sensitivity and specificity of two assays, the authors use mathematical models and measurements of intact and defective proviruses to assess how misclassification can impact estimates of natural and therapeutic reservoir reduction.
- Daniel B. Reeves
- , Christian Gaebler
- & Michel C. Nussenzweig
-
Article
| Open Access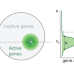 A model for organization and regulation of nuclear condensates by gene activity
A model for organization and regulation of nuclear condensates by gene activity
Through a physics-based model framework, the authors propose a central role for the nonequilibrium processes underling gene activity in shaping morphology, dynamics, and regulation of diverse nuclear condensates.
- Halima H. Schede
- , Pradeep Natarajan
- & Krishna Shrinivas
-
Article
| Open Access DNA 5-methylcytosine detection and methylation phasing using PacBio circular consensus sequencing
DNA 5-methylcytosine detection and methylation phasing using PacBio circular consensus sequencing
Existing methods for detecting DNA methylation (5mC) are less accurate and robust. Here, the authors develop a deep learning tool ccsmeth and a Nextflow pipeline ccsmethphase for genome-wide 5mCpG detection and phasing with high accuracy from CCS reads in human.
- Peng Ni
- , Fan Nie
- & Jianxin Wang
-
Article
| Open Access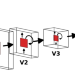 Learning-induced reorganization of number neurons and emergence of numerical representations in a biologically inspired neural network
Learning-induced reorganization of number neurons and emergence of numerical representations in a biologically inspired neural network
How the brain represents numbers remains poorly understood. Here, the authors uncover the emergence of absolute and relative magnitude representations of quantity in a biologically-inspired neural network, mirroring observations in children during numerical skill acquisition.
- Percy K. Mistry
- , Anthony Strock
- & Vinod Menon
-
Article
| Open Access Cell facilitation promotes growth and survival under drug pressure in breast cancer
Cell facilitation promotes growth and survival under drug pressure in breast cancer
In cancer, interactions between treatment-sensitive and resistant cells can influence the effectiveness of therapies. Here, the authors use experimental and mathematical models to explore interactions between ER+ breast cancer cell lineages that are sensitive or resistant to CDK4/6 inhibition, revealing the role of facilitative growth.
- Rena Emond
- , Jason I. Griffiths
- & Andrea H. Bild
-
Article
| Open Access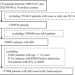 Predicting in-hospital outcomes of patients with acute kidney injury
Predicting in-hospital outcomes of patients with acute kidney injury
Early prediction of AKI-related clinical events and timely intervention for high-risk patients could improve outcomes. Here, the authors show a deep learning model that can identify patients with acute kidney injury (AKI) who are at high risk of death or dialysis at certain time points.
- Changwei Wu
- , Yun Zhang
- & Guisen Li
-
Article
| Open Access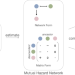 Joint inference of exclusivity patterns and recurrent trajectories from tumor mutation trees
Joint inference of exclusivity patterns and recurrent trajectories from tumor mutation trees
Understanding cancer evolution is crucial for developing effective therapies. Here, authors present TreeMHN, a probabilistic model for inferring exclusivity patterns of genomic events and evolutionary trajectories from intra-tumor phylogenetic trees.
- Xiang Ge Luo
- , Jack Kuipers
- & Niko Beerenwinkel
-
Article
| Open Access Biomedical knowledge graph learning for drug repurposing by extending guilt-by-association to multiple layers
Biomedical knowledge graph learning for drug repurposing by extending guilt-by-association to multiple layers
Computational drug repurposing models that leverage biomedical knowledge graphs to associate drugs to diseases, are biased to genes. Here, the authors present DREAMwalk, which extends guilt-by-association for multi-layer knowledge graph learning using a semantic information-guided random walk.
- Dongmin Bang
- , Sangsoo Lim
- & Sun Kim
-
Article
| Open Access A convolutional neural network STIFMap reveals associations between stromal stiffness and EMT in breast cancer
A convolutional neural network STIFMap reveals associations between stromal stiffness and EMT in breast cancer
The link between stiffness heterogeneity and tumor cell heterogeneity remains poorly understood. Here, authors propose an AI-informed method that reveals correlations between stromal stiffness and breast cancer cells with a heterogeneous EMT phenotype.
- Connor Stashko
- , Mary-Kate Hayward
- & Valerie M. Weaver

 Probabilities of developing HIV-1 bNAb sequence features in uninfected and chronically infected individuals
Probabilities of developing HIV-1 bNAb sequence features in uninfected and chronically infected individuals