Article
|
Open Access
Featured
-
-
Article
| Open Access A signal processing and deep learning framework for methylation detection using Oxford Nanopore sequencing
A signal processing and deep learning framework for methylation detection using Oxford Nanopore sequencing
The authors present DeepMod2, a deep-learning based computational method that allows fast and accurate detection of DNA methylation and epihaplotypes from Oxford Nanopore sequencing data.
- Mian Umair Ahsan
- , Anagha Gouru
- & Kai Wang
-
Article
| Open Access TFvelo: gene regulation inspired RNA velocity estimation
TFvelo: gene regulation inspired RNA velocity estimation
Most RNA velocity models extract dynamics from the phase delay between unspliced and spliced mRNA for each gene. Here, authors propose TFvelo, broadening RNA velocity beyond splicing information to include gene regulation. TFvelo accurately models genes dynamics and infers cell pseudo-time from RNA abundance data.
- Jiachen Li
- , Xiaoyong Pan
- & Hong-Bin Shen
-
Article
| Open Access MAIVeSS: streamlined selection of antigenically matched, high-yield viruses for seasonal influenza vaccine production
MAIVeSS: streamlined selection of antigenically matched, high-yield viruses for seasonal influenza vaccine production
Vaccines combat global influenza threats, relying on timely selection of optimal seed viruses. Here, authors introduce MAIVeSS, a machine learning assisted framework to streamline vaccine seed virus selection using genomic sequence, expediting seasonal flu vaccine production and supply.
- Cheng Gao
- , Feng Wen
- & Xiu-Feng Wan
-
Article
| Open Access Protein structure generation via folding diffusion
Protein structure generation via folding diffusion
The ability to engineer novel protein structures has tremendous scientific and therapeutic impact. Here, authors develop a generative model acting upon an angular representation of protein structures to create high quality protein backbones.
- Kevin E. Wu
- , Kevin K. Yang
- & Ava P. Amini
-
Article
| Open Access The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
Cell type annotation for single-cell data is challenging. Here, authors explore active and self-supervised learning and introduce adaptive reweighting as a tailored heuristic, demonstrating competitive performance and showing that incorporating prior knowledge enhances cell type annotation accuracy.
- Michael J. Geuenich
- , Dae-won Gong
- & Kieran R. Campbell
-
Article
| Open Access Congenital heart disease detection by pediatric electrocardiogram based deep learning integrated with human concepts
Congenital heart disease detection by pediatric electrocardiogram based deep learning integrated with human concepts
Congenital heart disease is life threatening, and its screening is complex and costly. Here, authors use AI to detect the disease based on pediatric electrocardiogram, suggesting superior performance over cardiologists.
- Jintai Chen
- , Shuai Huang
- & Huiying Liang
-
Article
| Open Access scDisInFact: disentangled learning for integration and prediction of multi-batch multi-condition single-cell RNA-sequencing data
scDisInFact: disentangled learning for integration and prediction of multi-batch multi-condition single-cell RNA-sequencing data
Here the authors propose a deep learning model that integrates multi-condition, multi-batch single-cell RNA-sequencing datasets. The model disentangles biological variation (condition effect) from technical confounders (batch effect) and overcomes some limitations of existing approaches.
- Ziqi Zhang
- , Xinye Zhao
- & Xiuwei Zhang
-
Article
| Open Access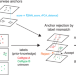 Semi-supervised integration of single-cell transcriptomics data
Semi-supervised integration of single-cell transcriptomics data
Batch effects hinder multi-sample single-cell data analyses. Here, authors present STACAS, a scalable single-cell RNA-seq data integration tool that uses prior cell type knowledge to preserve biological variability, demonstrating robustness to noisy input cell type labels.
- Massimo Andreatta
- , Léonard Hérault
- & Santiago J. Carmona
-
Article
| Open Access Compositional and temporal division of labor modulates mixed sugar fermentation by an engineered yeast consortium
Compositional and temporal division of labor modulates mixed sugar fermentation by an engineered yeast consortium
Synthetic microbial communities are suitable for mixed substrates fermentation and long metabolic pathway engineering. Here, the authors combine fermentation experiments with mathematical modeling to reveal the effect of compositional and temporal changes on division of labor in cellulosic ethanol production using two yeast strains.
- Jonghyeok Shin
- , Siqi Liao
- & Yong-Su Jin
-
Article
| Open Access SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
Cells simultaneously encode multiple signals, some harder to recover. Here, authors introduce SiFT (Signal FilTering), a kernel-based projection method, revealing underlying biological processes in single-cell data.
- Zoe Piran
- & Mor Nitzan
-
Article
| Open Access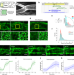 Emergence of periodic circumferential actin cables from the anisotropic fusion of actin nanoclusters during tubulogenesis
Emergence of periodic circumferential actin cables from the anisotropic fusion of actin nanoclusters during tubulogenesis
Periodic circumferential cytoskeletons support biological tube formation. Here, the authors show that self-assembled actin nanoclusters undergo biased fusion and develop into periodic cables in response to the membrane anisotropy of the expanding Drosophila tracheal tube.
- Sayaka Sekine
- , Mitsusuke Tarama
- & Shigeo Hayashi
-
Article
| Open Access Rational strain design with minimal phenotype perturbation
Rational strain design with minimal phenotype perturbation
No consensus exists on the computationally tractable use of dynamic models for strain design. To tackle this, the authors report a framework, nonlinear-dynamic-model-assisted rational metabolic engineering design, for efficiently designing robust, artificially engineered cellular organisms.
- Bharath Narayanan
- , Daniel Weilandt
- & Vassily Hatzimanikatis
-
Article
| Open Access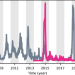 Shifting patterns of dengue three years after Zika virus emergence in Brazil
Shifting patterns of dengue three years after Zika virus emergence in Brazil
Dengue virus circulation was unusually low in Brazil in 2015-2018 following the emergence of Zika virus, but subsequently resurged causing large outbreaks with a lower mean age of infection. Here, the authors use mathematical modelling to investigate the links between dengue dynamics and prior Zika infection.
- Francesco Pinotti
- , Marta Giovanetti
- & José Lourenço
-
Article
| Open Access Determinants of epidemic size and the impacts of lulls in seasonal influenza virus circulation
Determinants of epidemic size and the impacts of lulls in seasonal influenza virus circulation
Seasonal influenza levels were unusually low when non-pharmaceutical interventions for COVID-19 were in place. Here, the authors analyse serological and epidemiological evidence for the hypothesis that such lulls in influenza transmission lead to reduced immunity and therefore larger epidemics in subsequent seasons.
- Simon P. J. de Jong
- , Zandra C. Felix Garza
- & Colin A. Russell
-
Article
| Open Access PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
Understanding biological mechanisms requires a thorough exploration of spatiotemporal transcriptional patterns in complex tissues. Here, authors present PROST to quantify spatial gene expression patterns and detect spatial domains using spatial transcriptomics data of varying resolutions.
- Yuchen Liang
- , Guowei Shi
- & Zhonghui Tang
-
Article
| Open Access BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
BIDCell: Biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data
Subcellular in situ spatial transcriptomics offers the promise to address biological problems that were previously inaccessible but requires accurate cell segmentation to uncover insights. Here, authors present BIDCell, a biologically informed, deep learning-based cell segmentation framework.
- Xiaohang Fu
- , Yingxin Lin
- & Jean Y. H. Yang
-
Article
| Open Access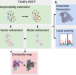 Cryo-EM structure and B-factor refinement with ensemble representation
Cryo-EM structure and B-factor refinement with ensemble representation
Cryo-EM is the go-to method for visualizing large, flexible biomolecules. Here, authors introduce a new Gaussian mixture modelling method for cryo-EM modelling tasks, including refinement, composite map generation and ensemble representation.
- Joseph G. Beton
- , Thomas Mulvaney
- & Maya Topf
-
Article
| Open Access Mechanism-centric regulatory network identifies NME2 and MYC programs as markers of Enzalutamide resistance in CRPC
Mechanism-centric regulatory network identifies NME2 and MYC programs as markers of Enzalutamide resistance in CRPC
Heterogeneous response to Enzalutamide remains a critical issue in castration-resistant prostate cancer (CRPC). Here, the authors reconstruct a CRPC-specific mechanism-centric regulatory network to identify signatures of Enzalutamide response and predict patients at risk of Enzalutamide resistance.
- Sukanya Panja
- , Mihai Ioan Truica
- & Antonina Mitrofanova
-
Article
| Open Access MarsGT: Multi-omics analysis for rare population inference using single-cell graph transformer
MarsGT: Multi-omics analysis for rare population inference using single-cell graph transformer
Identifying rare cell populations is key to understanding cancer progression and response to therapy. Here, authors introduce MarsGT, an end-to-end deep learning model for rare cell population identification from scMulti-omics data.
- Xiaoying Wang
- , Maoteng Duan
- & Qin Ma
-
Article
| Open Access Enhancing geometric representations for molecules with equivariant vector-scalar interactive message passing
Enhancing geometric representations for molecules with equivariant vector-scalar interactive message passing
Utilising geometric information and reducing computational costs are key challenges in the molecular modelling field. Here, authors propose ViSNet, which efficiently extracts geometric features, accurately predicts molecular properties, and drives simulations with interpretability.
- Yusong Wang
- , Tong Wang
- & Tie-Yan Liu
-
Article
| Open Access MENDER: fast and scalable tissue structure identification in spatial omics data
MENDER: fast and scalable tissue structure identification in spatial omics data
Identifying tissue structure in large-scale spatial omics datasets from multiple slices is challenging. Here, authors present MENDER, an optimisation-free spatial clustering method that can scale to million-level spatial data, enabling efficient analysis of spatial cell atlases.
- Zhiyuan Yuan
-
Article
| Open Access The juxtamembrane linker of synaptotagmin 1 regulates Ca2+ binding via liquid-liquid phase separation
The juxtamembrane linker of synaptotagmin 1 regulates Ca2+ binding via liquid-liquid phase separation
Synaptotagmin (syt) 1 is a calcium sensor for neuronal exocytosis. Here, the authors show that the juxtamembrane linker of this integral membrane protein negatively regulates its calcium sensing activity by mediating self-association via liquid-liquid phase separation.
- Nikunj Mehta
- , Sayantan Mondal
- & Edwin R. Chapman
-
Article
| Open Access Tuning parameters for polygenic risk score methods using GWAS summary statistics from training data
Tuning parameters for polygenic risk score methods using GWAS summary statistics from training data
Some polygenic risk score (PRS) methods for predicting genetic risk for common diseases require an external individual-level dataset for parameter tuning, posing privacy-related concerns. Here, the authors present an empirical Bayes method that tunes PRS models using only summary statistics from the training data.
- Wei Jiang
- , Ling Chen
- & Hongyu Zhao
-
Article
| Open Access Revealing hidden patterns in deep neural network feature space continuum via manifold learning
Revealing hidden patterns in deep neural network feature space continuum via manifold learning
Existing feature visualisation methods are not well-suited for regression tasks. Here, authors introduce a method to learn the manifold topology related to deep neural network output and target labels and provide insightful visualisations of the high-dimensional features while preserving the local geometry.
- Md Tauhidul Islam
- , Zixia Zhou
- & Lei Xing
-
Article
| Open Access Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Clustering-based analysis has limited power in highly dynamic single-cell data, which is a common situation in tumour samples. Here, authors introduce GSDensity, enabling pathway-centric analysis for the direct integration of data with their domain knowledge.
- Qingnan Liang
- , Yuefan Huang
- & Ken Chen
-
Article
| Open Access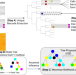 LinRace: cell division history reconstruction of single cells using paired lineage barcode and gene expression data
LinRace: cell division history reconstruction of single cells using paired lineage barcode and gene expression data
Inferring lineage trees while incorporating gene expressions and lineage barcodes is a challenging task. Here, authors present LinRace, which infers improved cell lineage trees and ancestral cell states using the proposed asymmetric division model.
- Xinhai Pan
- , Hechen Li
- & Xiuwei Zhang
-
Article
| Open Access A computational toolbox for the assembly yield of complex and heterogeneous structures
A computational toolbox for the assembly yield of complex and heterogeneous structures
Predicting the effective assembly of a set of proteins into a desired structure has traditionally been a challenging task. Here, authors demonstrate that advancements in automatic differentiation make it possible to address this problem using classical statistical mechanics.
- Agnese I. Curatolo
- , Ofer Kimchi
- & Michael P. Brenner
-
Article
| Open Access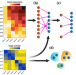 Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Here, the authors develop MaCroDNA, an algorithm to integrate single-cell DNA and RNA sequencing data from the same tissue. They use MaCroDNA to show—in agreement with previous studies—that copy number changes can predict progression from Barrett’s esophagus to esophageal adenocarcinoma.
- Mohammadamin Edrisi
- , Xiru Huang
- & Luay Nakhleh
-
Article
| Open Access GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
Reconstructing transcriptome-wide spatially-resolved gene expressions requires modelling nonlinear patterns and spatial structures in RNA profiling data. Here, authors introduce a graph-guided neural hierarchical tensor decomposition model that incorporates spatial and functional relations for the task.
- Tianci Song
- , Charles Broadbent
- & Rui Kuang
-
Article
| Open Access An archetype and scaling of developmental tissue dynamics across species
An archetype and scaling of developmental tissue dynamics across species
Limb tissue dynamics until basic skeletal pattern establishment exhibit a high degree of conservation between chick and frog after proper rescaling of spacetime, suggesting the presence of a species-independent archetype of morphogenetic dynamics.
- Yoshihiro Morishita
- , Sang-Woo Lee
- & Aiko Kawasumi-Kita
-
Article
| Open Access UniKP: a unified framework for the prediction of enzyme kinetic parameters
UniKP: a unified framework for the prediction of enzyme kinetic parameters
Prediction of enzyme kinetic parameters is essential for designing and optimising enzymes for various biotechnological and industrial applications. Here, authors presented a prediction framework (UniKP), which improves the accuracy of predictions for three enzyme kinetic parameters.
- Han Yu
- , Huaxiang Deng
- & Xiaozhou Luo
-
Article
| Open Access STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
Spatial transcriptomics (ST) enables gene expression characterisation within tissue sections, but comparing across sections and technologies remains challenging. Here, authors develop STalign to spatially align ST data and demonstrate applications including aligning to common coordinate frameworks.
- Kalen Clifton
- , Manjari Anant
- & Jean Fan
-
Article
| Open Access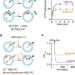 Phospholipids are imported into mitochondria by VDAC, a dimeric beta barrel scramblase
Phospholipids are imported into mitochondria by VDAC, a dimeric beta barrel scramblase
Mitochondria depend on phospholipids supplied by the endoplasmic reticulum. Here, using biochemical assays and molecular dynamics simulations, authors identify VDAC as a scramblase-type lipid transporter that catalyze lipid entry.
- Helene Jahn
- , Ladislav Bartoš
- & Anant K. Menon
-
Article
| Open Access Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Computational deconvolution with single-cell RNA sequencing data as a reference is pivotal for interpreting spatial transcriptomics data. Here, authors present Redeconve, which improves the resolution by more than 100-fold with higher accuracy and speed.
- Zixiang Zhou
- , Yunshan Zhong
- & Xianwen Ren
-
Article
| Open Access Engineered immunogens to elicit antibodies against conserved coronavirus epitopes
Engineered immunogens to elicit antibodies against conserved coronavirus epitopes
A pan-betacoronavirus vaccine will likely require the elicitation of antibodies against spike regions conserved across diverse coronaviruses. Here, authors computationally engineer and experimentally validate immunogens to elicit antibodies against two such spike regions.
- A. Brenda Kapingidza
- , Daniel J. Marston
- & Mihai L. Azoitei
-
Article
| Open Access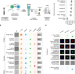 Explainable machine learning for profiling the immunological synapse and functional characterization of therapeutic antibodies
Explainable machine learning for profiling the immunological synapse and functional characterization of therapeutic antibodies
Therapeutic antibodies are crucial in treating severe diseases. Here, the authors introduce scifAI, an open-source explainable AI framework for analyzing imaging flow cytometry data, enabling rapid screening of therapeutic antibody candidates.
- Sayedali Shetab Boushehri
- , Katharina Essig
- & Fabian Schmich
-
Article
| Open Access TRS: a method for determining transcript termini from RNAtag-seq sequencing data
TRS: a method for determining transcript termini from RNAtag-seq sequencing data
TRS is a new method for determining 3’ transcript termini in bacteria, using data generated by the RNAtag-seq protocol. This methodology opens the door to study the evolution of transcription termini and their condition-dependent dynamics.
- Amir Bar
- , Liron Argaman
- & Hanah Margalit
-
Article
| Open Access Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Spatial transcriptomics (ST) is transforming tissue analysis but has limitations. Here, authors introduce SpatialScope, an integrated approach combining scRNA-seq and ST data using deep generative models, enabling comprehensive spatial characterisation at transcriptome-wide single-cell resolution.
- Xiaomeng Wan
- , Jiashun Xiao
- & Can Yang
-
Article
| Open Access PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
Here, authors present PhenoSV, a phenotype-aware machine-learning model for the functional interpretation of various types of structural variants (SVs) and genes within or outside SVs, facilitating the extraction of biological insights from coding and noncoding SVs.
- Zhuoran Xu
- , Quan Li
- & Kai Wang
-
Article
| Open Access scDREAMER for atlas-level integration of single-cell datasets using deep generative model paired with adversarial classifier
scDREAMER for atlas-level integration of single-cell datasets using deep generative model paired with adversarial classifier
Integration of single-cell datasets is essential to gain a comprehensive understanding of complex biological systems. Here, the authors develop scDREAMER, a deep generative framework for performing unsupervised and supervised atlas-level integration, demonstrating improved bio-conservation and batch-correction.
- Ajita Shree
- , Musale Krushna Pavan
- & Hamim Zafar
-
Article
| Open Access SPACEL: deep learning-based characterization of spatial transcriptome architectures
SPACEL: deep learning-based characterization of spatial transcriptome architectures
Spatial transcriptomics (ST) technologies detect transcript distribution in space. Here, authors present a deep learning based method SPACEL for cell type deconvolution, spatial domain identification and 3D alignment, showcasing it as a valuable toolkit for ST data analysis
- Hao Xu
- , Shuyan Wang
- & Kun Qu
-
Article
| Open Access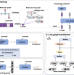 A knowledge-guided pre-training framework for improving molecular representation learning
A knowledge-guided pre-training framework for improving molecular representation learning
Accurate property prediction relies on effective molecular representation. Here, the authors introduce KPGT, a knowledge-guided self-supervised framework that improves molecular representation, leading to superior predictions of molecular properties and advancing AI-driven drug discovery.
- Han Li
- , Ruotian Zhang
- & Jianyang Zeng
-
Article
| Open Access Evaluation of the US COVID-19 Scenario Modeling Hub for informing pandemic response under uncertainty
Evaluation of the US COVID-19 Scenario Modeling Hub for informing pandemic response under uncertainty
The US COVID-19 Scenario Modeling Hub produced medium to long term projections based on different epidemic scenarios. In this study, the authors evaluate 14 rounds of projections by comparing them to the epidemic trajectories that occurred, and discuss lessons learned for future similar projects.
- Emily Howerton
- , Lucie Contamin
- & Justin Lessler
-
Article
| Open Access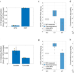 Inferring mitochondrial and cytosolic metabolism by coupling isotope tracing and deconvolution
Inferring mitochondrial and cytosolic metabolism by coupling isotope tracing and deconvolution
Studying metabolism in distinct subcellular compartments typically involves isolating organelles. Here, the authors demonstrate a quantitative approach to infer cytosolic and mitochondrial metabolic activities based on experiments with intact cells, maintaining physiological conditions.
- Alon Stern
- , Mariam Fokra
- & Tomer Shlomi
-
Article
| Open Access scReadSim: a single-cell RNA-seq and ATAC-seq read simulator
scReadSim: a single-cell RNA-seq and ATAC-seq read simulator
Benchmarking computational tools for analysis of single-cell sequencing data demands simulation of realistic sequencing reads. However, none of the few existing read simulators aim to mimic real data. Here, the authors introduce scReadSim, a single-cell RNA-seq and ATAC-seq read simulator that works by mimicking real data.
- Guanao Yan
- , Dongyuan Song
- & Jingyi Jessica Li
-
Article
| Open Access ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
Inverse Protein Folding is a critical component of protein design. Here, authors introduce ProRefiner, a deep-learning model for IPF that exhibits both high performance and memory efficiency, thereby contributing to advancements in protein design.
- Xinyi Zhou
- , Guangyong Chen
- & Pheng Ann Heng
-
Article
| Open Access Deep learning of human polyadenylation sites at nucleotide resolution reveals molecular determinants of site usage and relevance in disease
Deep learning of human polyadenylation sites at nucleotide resolution reveals molecular determinants of site usage and relevance in disease
The authors develop deep learning models to identify genome-wide polyA sites at nucleotide resolution and calculate site strength. They further examine genomic parameters regulating site usage and reveal genetic variants altering polyA activity.
- Emily Kunce Stroup
- & Zhe Ji
-
Article
| Open Access Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Identifying spatially variable genes (SVGs) is essential for linking molecular cell functions with tissue phenotypes. Here, authors introduce a non-parametric model that detects SVGs from two or three-dimensional spatial transcriptomics data by comparing gene expression patterns at granularities.
- Juexin Wang
- , Jinpu Li
- & Dong Xu
-
Article
| Open Access CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
Five-prime single-cell RNA-seq, especially the read 1, has precise capture of transcription start sites (TSS), but such information is often overlooked. Here, authors present a computational method suite, CamoTSS, to precisely identify TSS and quantify its expression, enabling effective detection of alternative TSS usage in different biological processes.
- Ruiyan Hou
- , Chung-Chau Hon
- & Yuanhua Huang

 Iterative design of training data to control intricate enzymatic reaction networks
Iterative design of training data to control intricate enzymatic reaction networks