Featured
-
-
Article
| Open Access Subtelomeric assembly of a multi-gene pathway for antimicrobial defense compounds in cereals
Subtelomeric assembly of a multi-gene pathway for antimicrobial defense compounds in cereals
The genomic organization and origin of the avenacin biosynthetic gene cluster remain unknown. Here, the authors assemble the genome of diploid oat Avena strigosa, reveal the structure and organization of the consecutive genes, characterize the last two missing pathway steps, and investigate the origin of the pathway in cereals.
- Yan Li
- , Aymeric Leveau
- & Anne Osbourn
-
Article
| Open Access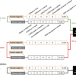 Learning a genome-wide score of human–mouse conservation at the functional genomics level
Learning a genome-wide score of human–mouse conservation at the functional genomics level
Understanding conserved functional genomic properties between human and mouse provides important context for mouse model studies. Here, the authors present a genome-wide conservation score integrating epigenomic, transcription factor binding, and transcriptomic data from mouse and human genomes.
- Soo Bin Kwon
- & Jason Ernst
-
Article
| Open Access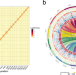 Chromosome-scale assembly and analysis of biomass crop Miscanthus lutarioriparius genome
Chromosome-scale assembly and analysis of biomass crop Miscanthus lutarioriparius genome
The genus Miscanthus has great potential for bio-energy production due to its high biomass yield and strong stress resistance. Here, the authors report the genome assembly of the diploid M. lutarioriparius, showing it has an allotetraploid origin and an expanded number of genes in families related to stress resistance.
- Jiashun Miao
- , Qi Feng
- & Bin Han
-
Article
| Open Access Apparent nosocomial adaptation of Enterococcus faecalis predates the modern hospital era
Apparent nosocomial adaptation of Enterococcus faecalis predates the modern hospital era
Enterococcus faecalis is a commensal microorganism of animals, insects and humans, but also a nosocomial pathogen. Here, the authors analyse genomic sequences from E. faecalis isolates from animals and humans, and find that the last common ancestors of multiple hospital-associated lineages date to the pre-antibiotic era.
- Anna K. Pöntinen
- , Janetta Top
- & Jukka Corander
-
Article
| Open Access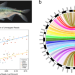 A chromosome-level genome of Astyanax mexicanus surface fish for comparing population-specific genetic differences contributing to trait evolution
A chromosome-level genome of Astyanax mexicanus surface fish for comparing population-specific genetic differences contributing to trait evolution
Mexican Tetra cavefish have long been of interest in understanding adaptation to severe environmental change. Here the authors present a chromosome-level genome for the proxy-ancestral surface fish, and use CRISPR gene-editing to show the role of the rx3 gene in eye size.
- Wesley C. Warren
- , Tyler E. Boggs
- & Nicolas Rohner
-
Article
| Open Access Genome evolution and the emergence of pathogenicity in avian Escherichia coli
Genome evolution and the emergence of pathogenicity in avian Escherichia coli
It is unclear how gut-dwelling E. coli bacteria often emerge to cause systemic infection in chickens. Here, Mageiros et al. use population genomics and pangenome-wide association studies to identify genetic elements associated with pathogenicity in avian E. coli.
- Leonardos Mageiros
- , Guillaume Méric
- & Samuel K. Sheppard
-
Article
| Open Access Scalable multiple whole-genome alignment and locally collinear block construction with SibeliaZ
Scalable multiple whole-genome alignment and locally collinear block construction with SibeliaZ
Multiple whole-genome alignment is a challenging problem in bioinformatics, especially when computational resources are limited. Here the authors present SibeliaZ, an algorithm and software based on analysis of de Bruijn graphs, which provides improved computational efficiency and scalability.
- Ilia Minkin
- & Paul Medvedev
-
Article
| Open Access Identification and characterization of constrained non-exonic bases lacking predictive epigenomic and transcription factor binding annotations
Identification and characterization of constrained non-exonic bases lacking predictive epigenomic and transcription factor binding annotations
Genome-wide maps of evolutionary constraint and large-scale compendia of epigenomic and transcription factor data provide complementary information for genome annotation. Here, the authors develop the Constrained Non-Exonic Predictor (CNEP) that enables better understanding of their relationship.
- Olivera Grujic
- , Tanya N. Phung
- & Jason Ernst
-
Article
| Open Access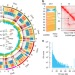 Genome of Solanum pimpinellifolium provides insights into structural variants during tomato breeding
Genome of Solanum pimpinellifolium provides insights into structural variants during tomato breeding
Solanum pimpinellifolium (SP) is the progenitor of cultivated tomato and an important germplasm. Here, the authors assemble SP genome, identify structural variants (SVs) by comparing with modern cultivar, reveal SVs associated with important breeding traits, and detect SVs harboring master regulators of fruit quality traits.
- Xin Wang
- , Lei Gao
- & Zhangjun Fei
-
Article
| Open Access Chromosome-level genome assembly of a parent species of widely cultivated azaleas
Chromosome-level genome assembly of a parent species of widely cultivated azaleas
Azaleas are one of the most diverse ornamental plants and have cultural and economic importance. Here, the authors report a chromosome-scale genome assembly for the primary ancestor of the azalea cultivar Rhododendro simsi and identify transcription factors that may function in flower coloration at different stages.
- Fu-Sheng Yang
- , Shuai Nie
- & Jian-Feng Mao
-
Article
| Open Access The structural variation landscape in 492 Atlantic salmon genomes
The structural variation landscape in 492 Atlantic salmon genomes
This study presents and validates a novel approach to reliably identify structural variations (SVs) in non-model genomes using whole genome sequencing, which was used to detect 15,483 SVs in 492 Atlantic salmon, shedding light on their roles in genome evolution and the genetic architecture of domestication.
- Alicia C. Bertolotti
- , Ryan M. Layer
- & Daniel J. Macqueen
-
Article
| Open Access Resequencing of 1,143 indica rice accessions reveals important genetic variations and different heterosis patterns
Resequencing of 1,143 indica rice accessions reveals important genetic variations and different heterosis patterns
Hybrid rice cultivars are widely planted around the world. Here, the authors resequence 1,143 indica accessions, focusing on the parents of superior hybrid rice lines in China, and reveal genetic loci that are associated with heterosis via measuring frequency of parental variation difference (FPVD).
- Qiming Lv
- , Weiguo Li
- & Dingyang Yuan
-
Article
| Open Access Retrospective evaluation of whole exome and genome mutation calls in 746 cancer samples
Retrospective evaluation of whole exome and genome mutation calls in 746 cancer samples
With the generation of large pan-cancer whole-exome and whole-genome sequencing projects, a question remains about how comparable these datasets are. Here, using The Cancer Genome Atlas samples analysed as part of the Pan-Cancer Analysis of Whole Genomes project, the authors explore the concordance of mutations called by whole exome sequencing and whole genome sequencing techniques.
- Matthew H. Bailey
- , William U. Meyerson
- & Christian von Mering
-
Article
| Open Access Gradual polyploid genome evolution revealed by pan-genomic analysis of Brachypodium hybridum and its diploid progenitors
Gradual polyploid genome evolution revealed by pan-genomic analysis of Brachypodium hybridum and its diploid progenitors
Existing plant pan-genomic studies usually report considerable intraspecific whole gene presence-absence variation. Here, the authors use pan-genomic approach to reveal gradual polyploid genome evolution by analyzing of Brachypodium hybridum and its diploid progenitors.
- Sean P. Gordon
- , Bruno Contreras-Moreira
- & John P. Vogel
-
Article
| Open Access Tracing animal genomic evolution with the chromosomal-level assembly of the freshwater sponge Ephydatia muelleri
Tracing animal genomic evolution with the chromosomal-level assembly of the freshwater sponge Ephydatia muelleri
Reconstructing the early molecular evolution of animals requires genomic resources for non-bilaterian animals. Here, the authors present the chromosome-level genome of a freshwater sponge together with analyses of its genome architecture, methylation, developmental gene expression, and microbiome.
- Nathan J. Kenny
- , Warren R. Francis
- & Sally P. Leys
-
Article
| Open Access Discovery and population genomics of structural variation in a songbird genus
Discovery and population genomics of structural variation in a songbird genus
Structural genomic variation can fuel evolution. Here, authors present genome data from seven Corvus species and unearth structural variants that vary between incipient crow species in Europe, with implications for premating isolation involving plumage patterning.
- Matthias H. Weissensteiner
- , Ignas Bunikis
- & Jochen B. W. Wolf
-
Article
| Open Access The Seminavis robusta genome provides insights into the evolutionary adaptations of benthic diatoms
The Seminavis robusta genome provides insights into the evolutionary adaptations of benthic diatoms
Available genomics studies have mostly focused on planktonic centric diatom. Here, the authors report the genome assembly of the marine biofilm-forming diatom Seminavis robusta and the resequencing data of a panel of accessions to reveal their evolutionary adaptations.
- Cristina Maria Osuna-Cruz
- , Gust Bilcke
- & Klaas Vandepoele
-
Article
| Open Access Genome and single-cell RNA-sequencing of the earthworm Eisenia andrei identifies cellular mechanisms underlying regeneration
Genome and single-cell RNA-sequencing of the earthworm Eisenia andrei identifies cellular mechanisms underlying regeneration
The mechanisms regulating regeneration of the earthworm are unclear. Here, the authors use genomic and transcriptomic analysis of the earthworm Eisenia andrei together with Hi-C analysis to identify genes involved and show activation of LINE2 transposable elements on regeneration.
- Yong Shao
- , Xiao-Bo Wang
- & Dong-Dong Wu
-
Article
| Open Access Exceptional subgenome stability and functional divergence in the allotetraploid Ethiopian cereal teff
Exceptional subgenome stability and functional divergence in the allotetraploid Ethiopian cereal teff
Teff is an indigenous cereal critical to food security in the Horn of Africa. Here, the authors report an improved genome assembly and observe the surprisingly low levels of large-scale structural rearrangement, homoeologous exchanges, or bias gene loss after the formation of this tetraploid species.
- Robert VanBuren
- , Ching Man Wai
- & Todd P. Michael
-
Article
| Open Access Genomic regions under selection in the feralization of the dingoes
Genomic regions under selection in the feralization of the dingoes
Dingoes evolved in isolation from both their domesticated and wild ancestors. Here, the authors investigate the genomic basis of the feralization of dingoes and trace their origin to domestic dogs that migrated to Australia approximately 8300 years ago.
- Shao-jie Zhang
- , Guo-Dong Wang
- & Ya-ping Zhang
-
Article
| Open Access Mikania micrantha genome provides insights into the molecular mechanism of rapid growth
Mikania micrantha genome provides insights into the molecular mechanism of rapid growth
Mikania micrantha is an extremely fast-growing invasive plant species that can cause serious damage to natural ecosystems. Here, the authors assemble its chromosome-scale reference genome and explore possible mechanisms that contribute to its rapid growth.
- Bo Liu
- , Jian Yan
- & Fanghao Wan
-
Article
| Open Access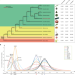 The wax gourd genomes offer insights into the genetic diversity and ancestral cucurbit karyotype
The wax gourd genomes offer insights into the genetic diversity and ancestral cucurbit karyotype
Cucurbits fruits have diverse shapes and sizes, but their genomes evolution and genetic basis of diversity are unclear. Here, the authors show that the wax gourd genome has the most ancestral karyotype among cucurbits and identify candidate genes which contribute to large fruit size by comparative and population genomics analyses.
- Dasen Xie
- , Yuanchao Xu
- & Zhonghua Zhang
-
Article
| Open Access Expansion of phycobilisome linker gene families in mesophilic red algae
Expansion of phycobilisome linker gene families in mesophilic red algae
Widely distributed red algae have experienced massive genome reduction during evolution. Here, using an improved genome assembly of Porphyridium purpureum, Lee et al. show the role of endosymbiotic gene transfer in plastid evolution and the correlation between phycobilisome linker diversification and the red algal radiation.
- JunMo Lee
- , Dongseok Kim
- & Hwan Su Yoon
-
Article
| Open Access Long-read assembly of the Chinese rhesus macaque genome and identification of ape-specific structural variants
Long-read assembly of the Chinese rhesus macaque genome and identification of ape-specific structural variants
Comparative genomic analysis of human and primate relatives can reveal important biological and evolutionary insights. Here, the authors present a long-read assembly of the Chinese rhesus macaque genome and identify ape-specific structural variants.
- Yaoxi He
- , Xin Luo
- & Bing Su
-
Article
| Open Access Comparative genomics reveals the origin of fungal hyphae and multicellularity
Comparative genomics reveals the origin of fungal hyphae and multicellularity
Hyphae are a major innovation in fungi associated with transitions to multicellularity. Here, Kiss and colleagues use comparative genomic analyses to reconstruct the evolutionary origins of hyphae and the molecular evolution of hypha morphogenesis genes.
- Enikő Kiss
- , Botond Hegedüs
- & László G. Nagy
-
Article
| Open Access A streamlined and predominantly diploid genome in the tiny marine green alga Chloropicon primus
A streamlined and predominantly diploid genome in the tiny marine green alga Chloropicon primus
The Chloropicophyceae represent an important group of green algae in tropical oceans, but there is only limited genomic resource available. Here, the authors present the genome sequence of Chloropicon primus, revealing a diploid structure and the presence of a propionate detoxification pathway.
- Claude Lemieux
- , Monique Turmel
- & Jean-François Pombert
-
Article
| Open Access Population dynamics of an Escherichia coli ST131 lineage during recurrent urinary tract infection
Population dynamics of an Escherichia coli ST131 lineage during recurrent urinary tract infection
Recurrent urinary tract infections occur in ~ 25% of women. Here, Beatson and colleagues use whole genome sequencing to track the dynamics of an E. coli ST131 clone in a single patient over a 5-year period. This study provides unique insights into pathogen evolution during recurrent urinary infection.
- Brian M. Forde
- , Leah W. Roberts
- & Scott A. Beatson
-
Article
| Open Access Extensive intraspecific gene order and gene structural variations in upland cotton cultivars
Extensive intraspecific gene order and gene structural variations in upland cotton cultivars
While multiple cotton genomes are available, genome wide variation comparison between allotetraploid upland cotton cultivars remain unexplored. Here, the authors assemble two upland cotton cultivars and reveal large scale structural variations on chromosome A08.
- Zhaoen Yang
- , Xiaoyang Ge
- & Fuguang Li
-
Article
| Open Access Statistics of chromatin organization during cell differentiation revealed by heterogeneous cross-linked polymers
Statistics of chromatin organization during cell differentiation revealed by heterogeneous cross-linked polymers
Chromatin is folded into Topologically Associating domains (TADs), with the organization and folding hierarchy of the TADs being highly dynamic. Here the authors develop a parsimonious randomly cross-linked (RCL) polymer model that maps high frequency encounters present in Hi-C data within and between TADs and reconstruct TADs across cell differentiation, revealing local chromatin re-organization.
- O. Shukron
- , V. Piras
- & D. Holcman
-
Article
| Open Access A high-quality apple genome assembly reveals the association of a retrotransposon and red fruit colour
A high-quality apple genome assembly reveals the association of a retrotransposon and red fruit colour
Existing apple genome assemblies all derive from Golden Delicious. Here, the authors combine different sequencing technologies to assemble a high quality genome of an anther-derived homozygous genotype HFTH1 and find the association of a retrotransposon and red fruit colour.
- Liyi Zhang
- , Jiang Hu
- & Peihua Cong
-
Article
| Open Access The genome of broomcorn millet
The genome of broomcorn millet
Broomcorn millet is one of the earliest domesticated plants and has the highest water use efficiency among cereals. Here, the authors report its genome assembly and annotation, which provides a valuable resource for breeders and paves the way for studying plant drought tolerance and C4 photosynthesis.
- Changsong Zou
- , Leiting Li
- & Heng Zhang
-
Article
| Open Access Chromosome conformation capture resolved near complete genome assembly of broomcorn millet
Chromosome conformation capture resolved near complete genome assembly of broomcorn millet
Broomcorn millet is one of the oldest crops cultivated by human that has strong abiotic stress tolerance. To facilitate genome assisted breeding of this and related species, the authors report its genome assembly and conduct comparative genome structure and evolution analyses with foxtail millet.
- Junpeng Shi
- , Xuxu Ma
- & Jinsheng Lai
-
Article
| Open Access Chromosome-level assembly of the water buffalo genome surpasses human and goat genomes in sequence contiguity
Chromosome-level assembly of the water buffalo genome surpasses human and goat genomes in sequence contiguity
Despite technological advances, chromosome-level assemblies of mammalian genomes are still rare. Here, the authors use PacBio, Chicago and Hi-C approaches to generate a highly contiguous and partially-phased genome assembly for the water buffalo, Bubalus bubalis
- Wai Yee Low
- , Rick Tearle
- & John L. Williams
-
Article
| Open Access Selective single molecule sequencing and assembly of a human Y chromosome of African origin
Selective single molecule sequencing and assembly of a human Y chromosome of African origin
Due to various structural and sequence complexities, the human Y chromosome is challenging to sequence and characterize. Here, the authors develop a strategy to sequence native, unamplified flow sorted Y chromosomes with a nanopore sequencing platform, and report the first assembly of a human Y chromosome of African origin.
- Lukas F. K. Kuderna
- , Esther Lizano
- & Tomas Marques-Bonet
-
Article
| Open Access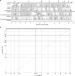 Genomic insights into multidrug-resistance, mating and virulence in Candida auris and related emerging species
Genomic insights into multidrug-resistance, mating and virulence in Candida auris and related emerging species
Candida auris is an emergent fungal pathogen that is resistant to multiple antifungals. Here, Muñoz et al. analyse genomic sequences for isolates from each of the four major C. auris clades and for three related species, and identify genetic features associated with virulence, antifungal resistance and mating.
- José F. Muñoz
- , Lalitha Gade
- & Christina A. Cuomo
-
Article
| Open Access Heart enhancers with deeply conserved regulatory activity are established early in zebrafish development
Heart enhancers with deeply conserved regulatory activity are established early in zebrafish development
During early embryogenesis, critical cardiac specification events occur. Here the authors isolate cardiac progenitor cells from early zebrafish embryos and characterize accessible chromatin regions specific to this cell population, finding that many of these regions overlap with conserved non-coding elements that are ortholgous to accessible chromatin regions in human.
- Xuefei Yuan
- , Mengyi Song
- & Michael D. Wilson
-
Article
| Open Access Phenotype loss is associated with widespread divergence of the gene regulatory landscape in evolution
Phenotype loss is associated with widespread divergence of the gene regulatory landscape in evolution
Cis-regulatory elements are important factors for morphological changes. Here, the authors show widespread divergence of limb and eye regulatory elements in limb loss in snakes and eye degeneration in subterranean mammals respectively.
- Juliana G. Roscito
- , Katrin Sameith
- & Michael Hiller
-
Article
| Open Access Genome sequences of two diploid wild relatives of cultivated sweetpotato reveal targets for genetic improvement
Genome sequences of two diploid wild relatives of cultivated sweetpotato reveal targets for genetic improvement
Sweetpotato is an important food security crop providing rich source of macro- and micronutrients including carbohydrates and vitamins. Here, the authors assemble of the two diploid relatives of cultivated sweetpotato and identify genes and alleles associated with carotenoid biosynthesis from breeding lines.
- Shan Wu
- , Kin H. Lau
- & Zhangjun Fei
-
Article
| Open Access A rapid rate of sex-chromosome turnover and non-random transitions in true frogs
A rapid rate of sex-chromosome turnover and non-random transitions in true frogs
The evolutionary forces that favour transitions in sex chromosomes are not well understood. Here, Jeffries and colleagues show a very high rate of sex chromosome turnover in true frogs, which may be driven by rapid mutation-load accumulation due to the low recombination rate in males.
- Daniel L. Jeffries
- , Guillaume Lavanchy
- & Nicolas Perrin
-
Article
| Open Access Genus-wide sequencing supports a two-locus model for sex-determination in Phoenix
Genus-wide sequencing supports a two-locus model for sex-determination in Phoenix
The origin and evolution of separate sexes in plants are long-standing questions. Here, the authors use genus-wide sequencing to identify sex determining candidate genes in the genus Phoenix and demonstrate the consistence with the previously proposed two-mutation model.
- Maria F. Torres
- , Lisa S. Mathew
- & Joel A. Malek
-
Article
| Open Access Horse Y chromosome assembly displays unique evolutionary features and putative stallion fertility genes
Horse Y chromosome assembly displays unique evolutionary features and putative stallion fertility genes
The rapidly evolving Y chromosome accumulates male-benefit genes but is often poorly characterized in many mammals. Here, the authors assemble the male specific region of the horse Y chromosome and investigate its evolution and function.
- Jan E. Janečka
- , Brian W. Davis
- & Terje Raudsepp
-
Article
| Open Access Squamate reptiles challenge paradigms of genomic repeat element evolution set by birds and mammals
Squamate reptiles challenge paradigms of genomic repeat element evolution set by birds and mammals
Large-scale patterns of genomic repeat element evolution have been studied mainly in birds and mammals. Here, the authors analyze the genomes of over 60 squamate reptiles and show high variation in repeat elements compared to mammals and birds, and particularly high microsatellite seeding in snakes.
- Giulia I. M. Pasquesi
- , Richard H. Adams
- & Todd A. Castoe
-
Article
| Open Access Large-scale gene losses underlie the genome evolution of parasitic plant Cuscuta australis
Large-scale gene losses underlie the genome evolution of parasitic plant Cuscuta australis
Dodders (Cuscuta spp., Convolvulaceae) are root- and leafless parasitic plants. Here, the authors sequence the genome of Cuscuta australis and find remarkable gene loss associated with parasitic lifestyle and large changes in body plan.
- Guiling Sun
- , Yuxing Xu
- & Jianqiang Wu
-
Article
| Open Access Parallels between experimental and natural evolution of legume symbionts
Parallels between experimental and natural evolution of legume symbionts
It is unclear if experimental evolution is a good model for natural processes. Here, Clerissi et al. find parallels between the evolution of symbiosis in rhizobia after horizontal transfer of a plasmid over 10 million years ago and experimentally evolved symbionts.
- Camille Clerissi
- , Marie Touchon
- & Eduardo P. C. Rocha
-
Article
| Open Access Diversity and evolution of the emerging Pandoraviridae family
Diversity and evolution of the emerging Pandoraviridae family
Giant viruses are visible by light microscopy and have unusually long genomes. Here, the authors report three new members of the Pandoraviridae family and investigate their evolution and diversity.
- Matthieu Legendre
- , Elisabeth Fabre
- & Jean-Michel Claverie
-
Article
| Open Access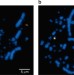 Reconstruction of the diapsid ancestral genome permits chromosome evolution tracing in avian and non-avian dinosaurs
Reconstruction of the diapsid ancestral genome permits chromosome evolution tracing in avian and non-avian dinosaurs
Ancient diapsids diverged into the lineages leading to turtles and birds over 250 million years ago. Here, the authors use genomic and molecular cytogenetic analyses of modern species to infer the genome structure of the diapsid common ancestor (DCA) and the changes occurring along the lineage to birds through theropod dinosaurs.
- Rebecca E. O’Connor
- , Michael N. Romanov
- & Darren K. Griffin
-
Article
| Open Access Reconstruction of the ancestral metazoan genome reveals an increase in genomic novelty
Reconstruction of the ancestral metazoan genome reveals an increase in genomic novelty
Animals, the Metazoa, co-opted numerous unicellular genes in their transition to multicellularity. Here, the authors use phylogenomic analyses to infer the genome composition of the ancestor of extant animals and show there was also a burst of novel gene groups associated with this transition.
- Jordi Paps
- & Peter W. H. Holland
-
Article
| Open Access Extreme haplotype variation in the desiccation-tolerant clubmoss Selaginella lepidophylla
Extreme haplotype variation in the desiccation-tolerant clubmoss Selaginella lepidophylla
Selaginella lepidophylla is a clubmoss with extreme desiccation tolerance. Here, the authors assemble its highly heterozygotic haplotypes and examine gene expression changes during desiccation, which shed light on the mechanisms for maintaining a small genome size and adaptation to extreme drying.
- Robert VanBuren
- , Ching Man Wai
- & Todd P. Michael
-
Article
| Open Access RNA sequencing provides insights into the evolution of lettuce and the regulation of flavonoid biosynthesis
RNA sequencing provides insights into the evolution of lettuce and the regulation of flavonoid biosynthesis
Horticultural lettuce varieties vary considerably in phenotype. Here, via RNA-seq of 240 different lettuce accessions, the authors identify loci and expression patterns associated with flavonoid and anthocyanin content and show that cultivated lettuce likely arose via a single domestication event.
- Lei Zhang
- , Wenqing Su
- & Hanhui Kuang

 SARS-CoV-2 gene content and COVID-19 mutation impact by comparing 44 Sarbecovirus genomes
SARS-CoV-2 gene content and COVID-19 mutation impact by comparing 44 Sarbecovirus genomes