Article
|
Open Access
Featured
-
-
Article
| Open Access Global role of the bacterial post-transcriptional regulator CsrA revealed by integrated transcriptomics
Global role of the bacterial post-transcriptional regulator CsrA revealed by integrated transcriptomics
The RNA-binding protein CsrA regulates the expression of hundreds of bacterial genes. Here, Potts et al. use several approaches to assess the contribution of CsrA to global gene expression in E. coli, revealing new binding targets and physiological roles such as in envelope function and iron homeostasis.
- Anastasia H. Potts
- , Christopher A. Vakulskas
- & Tony Romeo
-
Article
| Open Access The multiple antibiotic resistance operon of enteric bacteria controls DNA repair and outer membrane integrity
The multiple antibiotic resistance operon of enteric bacteria controls DNA repair and outer membrane integrity
Transcription factors MarR and MarA confer multidrug resistance in enteric bacteria by modulating efflux pump and porin expression. Here, Sharma et al. show that MarA also upregulates genes required for lipid trafficking and DNA repair, thus reducing antibiotic entry and quinolone-induced DNA damage.
- Prateek Sharma
- , James R. J. Haycocks
- & David C. Grainger
-
Article
| Open Access Dissemination of antibiotic resistance genes from antibiotic producers to pathogens
Dissemination of antibiotic resistance genes from antibiotic producers to pathogens
Some antibiotic resistance genes found in pathogenic bacteria might derive from antibiotic-producing actinobacteria. Here, Jianget al. provide bioinformatic and experimental evidence supporting this hypothesis, and propose a specific mechanism for the transfer of these genes between bacterial phyla.
- Xinglin Jiang
- , Mostafa M. Hashim Ellabaan
- & Sang Yup Lee
-
Article
| Open Access Evolutionary dynamics and genomic features of the Elizabethkingia anophelis 2015 to 2016 Wisconsin outbreak strain
Evolutionary dynamics and genomic features of the Elizabethkingia anophelis 2015 to 2016 Wisconsin outbreak strain
Elizabethkingia anophelis is an emerging pathogen of high antimicrobial resistance. Perrin and colleagues sequenced isolates of a 2015/2016 E. anophelis outbreak in Wisconsin and found substantial genetic diversity, accelerated evolutionary rate and a disruptive mutation in the DNA repair gene mutY.
- Amandine Perrin
- , Elise Larsonneur
- & Sylvain Brisse
-
Article
| Open Access Determining the bacterial cell biology of Planctomycetes
Determining the bacterial cell biology of Planctomycetes
Several unusual features have been reported for bacteria of the phylum Planctomycetes, such as cytosolic compartmentalization and an endocytosis-like process. Here, Boedekeret al. provide evidence supporting a Gram-negative cell plan and the absence of endocytosis-like processes in these organisms.
- Christian Boedeker
- , Margarete Schüler
- & Christian Jogler
-
Article
| Open Access Management of E. coli sister chromatid cohesion in response to genotoxic stress
Management of E. coli sister chromatid cohesion in response to genotoxic stress
Homologous recombination of DNA lesions in bacteria involves sister chromatid pairing. Here, the authors show that RecN promotes contacts between sister chromatids and facilitates repair.
- Elise Vickridge
- , Charlene Planchenault
- & Olivier Espéli
-
Article
| Open Access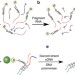 Global repositioning of transcription start sites in a plant-fermenting bacterium
Global repositioning of transcription start sites in a plant-fermenting bacterium
Bacteria may respond to a change in environment by using alternative transcriptional start sites. Here, the authors use a novel genome-wide capture and reverse transcription method to find substrate-specific start sites for hundreds of genes at single base resolution inClostridium phytofermentans.
- Magali Boutard
- , Laurence Ettwiller
- & Andrew C. Tolonen
-
Article
| Open Access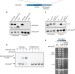 Chaperone addiction of toxin–antitoxin systems
Chaperone addiction of toxin–antitoxin systems
Some bacterial toxin-antitoxin systems consist of a labile antitoxin that inhibits a toxin, and a chaperone that stabilizes the antitoxin. Here, Bordes et al. identify a sequence within the antitoxin to which the chaperone binds and which can be transferred to other proteins to make them chaperone-dependent.
- Patricia Bordes
- , Ambre Julie Sala
- & Pierre Genevaux
-
Article
| Open Access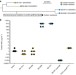 The evolution of antimicrobial peptide resistance in Pseudomonas aeruginosa is shaped by strong epistatic interactions
The evolution of antimicrobial peptide resistance in Pseudomonas aeruginosa is shaped by strong epistatic interactions
Colistin is an antibiotic used in the treatment of Pseudomonas aeruginosa infections in cystic fibrosis patients. Here, Jochumsen et al. reconstruct the pathways for the molecular evolution of colistin resistance in P. aeruginosaand show that the number of pathways is highly constrained by interactions among genes.
- Nicholas Jochumsen
- , Rasmus L. Marvig
- & Anders Folkesson
-
Article
| Open Access Sequence element enrichment analysis to determine the genetic basis of bacterial phenotypes
Sequence element enrichment analysis to determine the genetic basis of bacterial phenotypes
Plasticity and clonal population structure in bacterial genomes can hinder traditional SNP-based genetic association studies. Here, Corander and colleagues present a method to identify variable-length sequence elements enriched in a phenotype of interest, and demonstrate its use in human pathogens.
- John A. Lees
- , Minna Vehkala
- & Jukka Corander
-
Article
| Open Access Molecular logic of the Zur-regulated zinc deprivation response in Bacillus subtilis
Molecular logic of the Zur-regulated zinc deprivation response in Bacillus subtilis
The transcription factor Zur controls the zinc deprivation response in Bacillus subtilis. Here, Shin and Helmann show that Zur-regulated genes are derepressed in three waves in response to zinc deprivation, and this is linked to the biochemistry of zinc sensing by Zur.
- Jung-Ho Shin
- & John D. Helmann
-
Article
| Open Access A rheostat mechanism governs the bifurcation of carbon flux in mycobacteria
A rheostat mechanism governs the bifurcation of carbon flux in mycobacteria
Microbes survive in dynamic environments by modulating their intracellular metabolism. Here, the authors reveal that mycobacteria employ a rheostat-like mechanism to regulate carbon flux between the oxidative TCA cycle and the glyoxylate shunt during glucose-acetate diauxic shift.
- Paul Murima
- , Michael Zimmermann
- & John D. McKinney
-
Article
| Open Access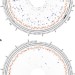 Genome-wide analysis of chromosomal import patterns after natural transformation of Helicobacter pylori
Genome-wide analysis of chromosomal import patterns after natural transformation of Helicobacter pylori
Uptake and integration of exogenous DNA into the bacterial genome play an important role in the evolution of the pathogen Helicobacter pylori. Here, the authors describe a bimodal pattern of chromosomal integration and show how restriction-modification systems limit the import of heterologous DNA.
- Sebastian Bubendorfer
- , Juliane Krebes
- & Sebastian Suerbaum
-
Article
| Open Access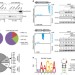 The CsrA-FliW network controls polar localization of the dual-function flagellin mRNA in Campylobacter jejuni
The CsrA-FliW network controls polar localization of the dual-function flagellin mRNA in Campylobacter jejuni
The CsrA protein binds to and represses translation of certain bacterial mRNAs. Here, Dugar et al. show for the human pathogen Campylobacter jejunithat the major flagellin mRNA acts as both a target and a regulatory 'sponge' for CsrA, and is localized at the cell poles in a translation-dependent manner.
- Gaurav Dugar
- , Sarah L. Svensson
- & Cynthia M. Sharma
-
Article
| Open Access A single gene of a commensal microbe affects host susceptibility to enteric infection
A single gene of a commensal microbe affects host susceptibility to enteric infection
The interactions between gut bacteria and enteric pathogens are poorly understood. Here, Yoon et al. show that subinhibitory antibiotic treatment in a mouse model leads to overgrowth of an E. coli strain carrying a catalase-encoding gene that enhances infection with the human pathogen Vibrio cholerae.
- Mi Young Yoon
- , Kyung Bae Min
- & Sang Sun Yoon
-
Article
| Open Access Chlamydia trachomatis from Australian Aboriginal people with trachoma are polyphyletic composed of multiple distinctive lineages
Chlamydia trachomatis from Australian Aboriginal people with trachoma are polyphyletic composed of multiple distinctive lineages
Chlamydia trachomatis isolates causing a blinding disease (trachoma) form a single lineage that is different from the lineages causing urogenital infections. Here, Andersson et al. show however that trachoma isolates from Australia are more closely related to urogenital strains than to other trachoma isolates.
- Patiyan Andersson
- , Simon R. Harris
- & Philip M. Giffard
-
Article
| Open Access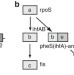 The propagation of perturbations in rewired bacterial gene networks
The propagation of perturbations in rewired bacterial gene networks
Expression of transcription factors to alter gene regulation can cause substantial changes to expression across a genome. Here the authors ‘rewire’ E. coliand analyse the global transcriptome alterations to identify novel network interactions.
- Rebecca Baumstark
- , Sonja Hänzelmann
- & Mark Isalan
-
Article
| Open Access Adaptive immunity increases the pace and predictability of evolutionary change in commensal gut bacteria
Adaptive immunity increases the pace and predictability of evolutionary change in commensal gut bacteria
The mechanisms underlying host-commensal coevolution are incompletely understood. Here the authors show that host adaptive immunity directs the evolution of Escherichia coliin the mouse gut towards host benefit by influencing the microbiome composition.
- João Barroso-Batista
- , Jocelyne Demengeot
- & Isabel Gordo
-
Article |
 Mycobacteria modulate host epigenetic machinery by Rv1988 methylation of a non-tail arginine of histone H3
Mycobacteria modulate host epigenetic machinery by Rv1988 methylation of a non-tail arginine of histone H3
Epigenetic modulation of hosts by pathogenic bacteria is underexplored. Here, Yaseen et al. show that protein Rv1988 from Mycobacterium tuberculosisenhances microbial survival by methylating histone H3 in the host cell nucleus and thus altering host gene expression.
- Imtiyaz Yaseen
- , Prabhjot Kaur
- & Sanjeev Khosla
-
Article
| Open Access Expanding the biotechnology potential of lactobacilli through comparative genomics of 213 strains and associated genera
Expanding the biotechnology potential of lactobacilli through comparative genomics of 213 strains and associated genera
Lactobacillus is a lactic acid bacteria and has a wide range of application from use in probiotic food production to biotherapeutics. Here, the authors sequence and compare the genomes of 213 different Lactobacillusstrains and related genera, and provide new insight into phylogenomic organization and adaptive immunity elements in this bacteria family.
- Zhihong Sun
- , Hugh M. B. Harris
- & Paul W. O’Toole
-
Article
| Open Access Decoding genome-wide GadEWX-transcriptional regulatory networks reveals multifaceted cellular responses to acid stress in Escherichia coli
Decoding genome-wide GadEWX-transcriptional regulatory networks reveals multifaceted cellular responses to acid stress in Escherichia coli
GadEWX regulons play a critical role in transcription regulation in response to acid stress. By reconstructing genome-wide GadEWX transcriptional network, here the authors show how GadEWX simultaneously coordinates many other cellular processes to produce the overall response of E. colito acid stress.
- Sang Woo Seo
- , Donghyuk Kim
- & Bernhard O. Palsson
-
Article
| Open Access A biphasic epigenetic switch controls immunoevasion, virulence and niche adaptation in non-typeable Haemophilus influenzae
A biphasic epigenetic switch controls immunoevasion, virulence and niche adaptation in non-typeable Haemophilus influenzae
Non-typeable Haemophilus influenzae, which causes ear and lung infections, has a DNA methyltransferase encoded by alternative alleles that are subject to random ON/OFF switching. Here, Atack et al.show that this epigenetic switch controls the expression of key proteins involved in virulence.
- John M. Atack
- , Yogitha N. Srikhanta
- & Michael P. Jennings
-
Article
| Open Access Single molecule-level detection and long read-based phasing of epigenetic variations in bacterial methylomes
Single molecule-level detection and long read-based phasing of epigenetic variations in bacterial methylomes
Bacterial DNA methylation is involved in many processes, from host defense to antibiotic resistance, however current methods for examining methylated genomes lack single-cell resolution. Here Beaulaurier et al. present Single Molecule Modification Analysis of Long Reads, a new tool for de novodetection of epigenetic heterogeneity.
- John Beaulaurier
- , Xue-Song Zhang
- & Gang Fang
-
Article
| Open Access Phasing of single DNA molecules by massively parallel barcoding
Phasing of single DNA molecules by massively parallel barcoding
DNA phasing information — the determination of which specific sequences belong to the same DNA molecule—is not easily obtained from sequencing applications that rely on short reads. Here the authors develop a phasing method based on massively parallel barcoding of single DNA molecules.
- Erik Borgström
- , David Redin
- & Afshin Ahmadian
-
Article
| Open Access Characterization of genome-wide ordered sequence-tagged Mycobacterium mutant libraries by Cartesian Pooling-Coordinate Sequencing
Characterization of genome-wide ordered sequence-tagged Mycobacterium mutant libraries by Cartesian Pooling-Coordinate Sequencing
The generation of characterized panels of specific mutants is an essential but time-consuming step of reverse genetic studies. Here Vandewalle et al. describe CP-CSeq, an easy to implement parallel sequencing method for rapid library construction.
- Kristof Vandewalle
- , Nele Festjens
- & Nico Callewaert
-
Article
| Open Access Interactions between horizontally acquired genes create a fitness cost in Pseudomonas aeruginosa
Interactions between horizontally acquired genes create a fitness cost in Pseudomonas aeruginosa
Horizontal gene transfer is important for bacterial evolution but the molecular basis of its fitness costs remain unclear. Here the authors show that fitness costs produced by a plasmid in P. aeruginosaare alleviated by mutations in recently acquired genes encoded in mobile genetic elements.
- Alvaro San Millan
- , Macarena Toll-Riera
- & R. Craig MacLean
-
Article
| Open Access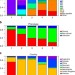 Genomic signatures of human and animal disease in the zoonotic pathogen Streptococcus suis
Genomic signatures of human and animal disease in the zoonotic pathogen Streptococcus suis
The bacterium Streptococcus suiscauses respiratory tract infections in pigs and meningitis in humans. Here, the authors show that human disease isolates are limited to a single virulent population and find no consistent genomic differences between pig and human isolates.
- Lucy A. Weinert
- , Roy R. Chaudhuri
- & Vanessa Terra
-
Article
| Open Access The DNA-binding network of Mycobacterium tuberculosi s
The DNA-binding network of Mycobacterium tuberculosi s
Adaptation of Mycobacterium tuberculosis to the host environment is principally mediated through its transcription factors. Here, the authors report the DNA binding and transcriptional profile of ~80% of all predicted M. tuberculosistranscription factors, and find wide-spread dormant DNA binding.
- Kyle J. Minch
- , Tige R. Rustad
- & David R. Sherman
-
Article |
 Multiple enzymatic activities of ParB/Srx superfamily mediate sexual conflict among conjugative plasmids
Multiple enzymatic activities of ParB/Srx superfamily mediate sexual conflict among conjugative plasmids
Conjugative plasmids block translocation of rival plasmids using fertility inhibition factors (FINs). Here Maindola et al.present the structure of the FIN Osa and show that it contains a ParB/Sulfiredoxin fold with both ATPase and DNase activity, with general functional implications for this fold.
- Priyank Maindola
- , Rahul Raina
- & Arulandu Arockiasamy
-
Article
| Open Access A robust SNP barcode for typing Mycobacterium tuberculosis complex strains
A robust SNP barcode for typing Mycobacterium tuberculosis complex strains
Genetic variation in Mycobacterium tuberculosiscomplex (MTBC) bacteria is responsible for differences in factors such as virulence and transmissibility. Here, the authors analyse the genomes of 1,601 MTBC isolates from diverse geographic locations and identify 62 SNPs that may be used to resolve lineages and sublineages of these strains.
- Francesc Coll
- , Ruth McNerney
- & Taane G. Clark
-
Article |
 Adaptation in bacterial CRISPR-Cas immunity can be driven by defective phages
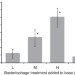
Adaptation in bacterial CRISPR-Cas immunity can be driven by defective phages
The bacterial ‘adaptive’ immune system known as CRISPR-Cas destroys foreign DNA molecules, such as viral genomes, to which the cells have previously been exposed. Here, Hynes et al.show that this gain of immunity is favoured by exposure to defective viruses, a result reminiscent of vaccination.
- Alexander P. Hynes
- , Manuela Villion
- & Sylvain Moineau
-
Article |
 Mutants of Cre recombinase with improved accuracy
Mutants of Cre recombinase with improved accuracy
Cre recombinase is widely used to precisely manipulate genes and chromosomes, but it often displays off-target activity. Here, the authors improve the accuracy of Cre-mediated recombination by introducing specific mutations in the enzyme’s dimerization surface.
- Nikolai Eroshenko
- & George M. Church
-
Article
| Open Access Genome sequence and functional genomic analysis of the oil-degrading bacterium Oleispira antarctica
Genome sequence and functional genomic analysis of the oil-degrading bacterium Oleispira antarctica
Oleispira antarctica is an oil-degrading bacterium found in the cold and deep sea. Here Kube et al. report the genome sequence of O. antarcticaand provide a comprehensive functional genetic and protein structural analysis, revealing insights into how this organism has adapted to its cold environment.
- Michael Kube
- , Tatyana N. Chernikova
- & Peter N. Golyshin
-
Article
| Open Access R-loops and nicks initiate DNA breakage and genome instability in non-growing Escherichia coli
R-loops and nicks initiate DNA breakage and genome instability in non-growing Escherichia coli
DNA double-strand breaks commonly occur in all replicating cells. Wimberly and colleagues show that in non-replicating cells, aborted transcription/translation forms RNA/DNA hybrid R-loops that prime origin-independent replication, leading to DNA breakage, point mutations and chromosomal rearrangements.
- Hallie Wimberly
- , Chandan Shee
- & P. J. Hastings
-
Article |
 Insertion sequence-excision enhancer removes transposable elements from bacterial genomes and induces various genomic deletions
Insertion sequence-excision enhancer removes transposable elements from bacterial genomes and induces various genomic deletions
Insertion sequences are transposable elements that are found in the genomes of many bacteria. Here, the authors identify an enhancer element that results in a high frequency of excision of insertion elements, and suggest that the excision enhancer element coevolved with the insertion sequences.
- Masahiro Kusumoto
- , Tadasuke Ooka
- & Tetsuya Hayashi

 Identification of regulatory targets for the bacterial Nus factor complex
Identification of regulatory targets for the bacterial Nus factor complex