Featured
-
-
Article
| Open Access Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
Dynamics of extended-spectrum cephalosporin resistance genes in Escherichia coli from Europe and North America
Extended-spectrum cephalosporin resistance genes in Escherichia coli have spread worldwide. Here, the authors dissect the emergence and distribution of these genes over time, and across geographic location and host species, to better understand their dynamics and mechanisms of transmission.
- Roxana Zamudio
- , Patrick Boerlin
- & Alison E. Mather
-
Article
| Open Access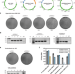 Genetic manipulation of the human gut bacterium Eggerthella lenta reveals a widespread family of transcriptional regulators
Genetic manipulation of the human gut bacterium Eggerthella lenta reveals a widespread family of transcriptional regulators
Eggerthella lenta is a prominent human gut bacterium implicated in several physiological processes, but its study has remained limited. Here, by developing a genetic toolbox for E. lenta, the authors provide insights into how the bacterium regulates drug and dietary compound metabolism.
- Xueyang Dong
- , Ben G. H. Guthrie
- & Emily P. Balskus
-
Article
| Open Access A comprehensive update to the Mycobacterium tuberculosis H37Rv reference genome
A comprehensive update to the Mycobacterium tuberculosis H37Rv reference genome
H37Rv is the most widely used Mycobacterium tuberculosis strain, and its genome is the reference sequence for this pathogen. Here, Chitale et al. present a bioinformatic pipeline for accurate assembly of bacterial genome sequences, and use it to provide important updates to the M. tuberculosis reference genome.
- Poonam Chitale
- , Alexander D. Lemenze
- & David Alland
-
Article
| Open Access Promiscuity of response regulators for thioredoxin steers bacterial virulence
Promiscuity of response regulators for thioredoxin steers bacterial virulence
The response regulator SsrB, a master activator of the Salmonella pathogenicity island-2 gene cluster, is under allosteric control of thioredoxin. Authors utilise in vitro and in vivo models to investigate if other members of the response regulator family might have adopted thioredoxin as a regulator.
- Ju-Sim Kim
- , Alexandra Born
- & Andrés Vázquez-Torres
-
Article
| Open Access Transcontinental spread and evolution of Mycobacterium tuberculosis W148 European/Russian clade toward extensively drug resistant tuberculosis
Transcontinental spread and evolution of Mycobacterium tuberculosis W148 European/Russian clade toward extensively drug resistant tuberculosis
An outbreak of the multidrug-resistant Mycobacterium tuberculosis lineage W148 has spread widely across Russia, Central Asia and Europe. Here, the authors use whole genome sequences of ~700 isolates of this lineage collected over ~20 years to analyze its spread, evolution of drug resistance, and impact of compensatory mutations.
- Matthias Merker
- , Jean-Philippe Rasigade
- & Thierry Wirth
-
Article
| Open Access Rapid evolution of mutation rate and spectrum in response to environmental and population-genetic challenges
Rapid evolution of mutation rate and spectrum in response to environmental and population-genetic challenges
How rapidly the mutation rate responds evolutionarily to ecological and population-genetic factors over time is unclear. Here, the authors show that the evolution of mutation rates in E. coli proceeds rapidly in response to these factors with substantial bidirectional shifts.
- Wen Wei
- , Wei-Chin Ho
- & Michael Lynch
-
Article
| Open Access Adaptation to novel spatially-structured environments is driven by the capsule and alters virulence-associated traits
Adaptation to novel spatially-structured environments is driven by the capsule and alters virulence-associated traits
Phenotypic and genotypic evolution in worrisome Klebsiella spp. is influenced by the capsule. Here the authors show that adaptation outside the host can impact virulence-associated traits, including de novo emergence of hypermucoviscosity.
- Amandine Nucci
- , Eduardo P. C. Rocha
- & Olaya Rendueles
-
Article
| Open Access Combined comparative genomics and clinical modeling reveals plasmid-encoded genes are independently associated with Klebsiella infection
Combined comparative genomics and clinical modeling reveals plasmid-encoded genes are independently associated with Klebsiella infection
Patient variables, such as comorbidities, partially explain which patients will progress to Klebsiella infection, with colonization of the gut acting as a reservoir. Little is known, however, regarding Klebsiella genes that may increase risk of disease in colonized individuals. Here, authors conduct a comparative genomics study to identify genes associated with progression from colonisation to infection.
- Jay Vornhagen
- , Emily K. Roberts
- & Michael A. Bachman
-
Article
| Open Access Molecular mechanism of toxin neutralization in the HipBST toxin-antitoxin system of Legionella pneumophila
Molecular mechanism of toxin neutralization in the HipBST toxin-antitoxin system of Legionella pneumophila
Here, the authors demonstrate that the Legionella pneumophila T4SS effector protein Lpg2370 is a Ser/Thr kinase and a toxin of a tripartite HipBST toxin-antitoxin (TA) system. Structural data and biochemical analysis provide detailed insights into the toxin neutralization mechanism in the HipBST TA.
- Xiangkai Zhen
- , Yongyu Wu
- & Songying Ouyang
-
Article
| Open Access Paeniclostridium sordellii hemorrhagic toxin targets TMPRSS2 to induce colonic epithelial lesions
Paeniclostridium sordellii hemorrhagic toxin targets TMPRSS2 to induce colonic epithelial lesions
Paeniclostridium sordellii is an opportunistic pathogen that can occur and be fatal in women undergoing abortion or childbirth. The pathogenesis of a hemorrhagic toxin, TcsH, produced by this bacteria, remains unknown. Here, authors carry out genome-wide screens to identify pathologically relevant host factors of TcsH.
- Xingxing Li
- , Liuqing He
- & Liang Tao
-
Article
| Open Access A proteolytically activated antimicrobial toxin encoded on a mobile plasmid of Bacteroidales induces a protective response
A proteolytically activated antimicrobial toxin encoded on a mobile plasmid of Bacteroidales induces a protective response
The bacterium Phocaeicola vulgatus is commonly found in the human gut. Here, the authors show that the microorganism produces an antibacterial toxin that targets the LPS core glycan of closely related species and induces a response that partially protects cells from multiple antimicrobial toxins.
- Jordan C. Evans
- , Valentina Laclare McEneany
- & Laurie E. Comstock
-
Article
| Open Access Development of a novel core genome MLST scheme for tracing multidrug resistant Staphylococcus capitis
Development of a novel core genome MLST scheme for tracing multidrug resistant Staphylococcus capitis
Staphylococcus capitis is a common causative agent of bloodstream infections in neonatal intensive care units, with multidrug resistant isolates complicating treatment. Authors aimed to establish a core genome multilocus sequence typing (cgMLST) scheme to document the transmission and dissemination of multidrug-resistant S. capitis isolates.
- Zhengan Wang
- , Chao Gu
- & Yunsong Yu
-
Article
| Open Access Population genomics of Group B Streptococcus reveals the genetics of neonatal disease onset and meningeal invasion
Population genomics of Group B Streptococcus reveals the genetics of neonatal disease onset and meningeal invasion
Group B Streptococcus (GBS) causes neonatal disease and mortality worldwide. Here, the authors use genome-wide association analyses to identify bacterial genetic signatures associated with disease onset time and meningeal tissue infection in acute invasive neonatal GBS disease.
- Chrispin Chaguza
- , Dorota Jamrozy
- & Stephen D. Bentley
-
Article
| Open Access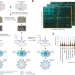 Auxotrophic and prototrophic conditional genetic networks reveal the rewiring of transcription factors in Escherichia coli
Auxotrophic and prototrophic conditional genetic networks reveal the rewiring of transcription factors in Escherichia coli
The bacterium E. coli has around 300 transcriptional factors, but the functions of many of them, and the interactions between their respective regulatory networks, are unclear. Here, the authors study genetic interactions among all transcription factor genes in E. coli, revealing condition-dependent interactions and roles for uncharacterized transcription factors.
- Alla Gagarinova
- , Ali Hosseinnia
- & Mohan Babu
-
Article
| Open Access A plasmid system with tunable copy number
A plasmid system with tunable copy number
The range of available copy numbers for cloning vectors is largely restricted to the handful of ORIs that have been isolated from plasmids found in nature. Here the authors introduce a plasmid system that allow for the continuous, finely-tuned control of plasmid copy number between 1 and 800 copies per cell.
- Miles V. Rouches
- , Yasu Xu
- & Guillaume Lambert
-
Article
| Open Access RNase III-CLASH of multi-drug resistant Staphylococcus aureus reveals a regulatory mRNA 3′UTR required for intermediate vancomycin resistance
RNase III-CLASH of multi-drug resistant Staphylococcus aureus reveals a regulatory mRNA 3′UTR required for intermediate vancomycin resistance
Regulatory small RNA (sRNA) interact with mRNAs to regulate their stability, transcription, and translation via diverse mechanisms. Here, Mediati et al. apply RNase III-CLASH to multidrug-resistant Staphylococcus aureus to characterise the network of RNA–RNA interactions associated with RNase III and identify a regulatory mRNA 3′UTR, named vigR-3′UTR, involved in the regulation of genes relevant for vancomycin sensitivity.
- Daniel G. Mediati
- , Julia L. Wong
- & Jai J. Tree
-
Article
| Open Access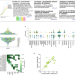 A genome-wide atlas of antibiotic susceptibility targets and pathways to tolerance
A genome-wide atlas of antibiotic susceptibility targets and pathways to tolerance
A lack of understanding in the development and emergence of antimicrobial resistance presents as a problem for accurate infection diagnosis and treatment. Here, authors utilize Streptococcus pneumoniae and build a genome-wide atlas to understand the genes and interactions that contribute to altered drug susceptibility.
- Dmitry Leshchiner
- , Federico Rosconi
- & Tim van Opijnen
-
Article
| Open Access Frequency modulation of a bacterial quorum sensing response
Frequency modulation of a bacterial quorum sensing response
Quorum-sensing bacteria produce and secrete autoinducers that trigger a behavioral change in the population when reaching a certain threshold. Here, Bettenworth et al. show that autoinducer synthase gene expression in Sinorhizobium meliloti occurs in asynchronous stochastic pulses, and that physiological cues modulate pulse frequency and, consequently, response behavior dynamics. Frequency-modulated pulsing in autoinducer synthase gene expression thus represents a time-based mechanism for information integration and collective decision-making.
- Vera Bettenworth
- , Simon van Vliet
- & Anke Becker
-
Matters Arising
| Open AccessTesting the adaptive hypothesis of lagging-strand encoding in bacterial genomes
- Haoxuan Liu
- & Jianzhi Zhang
-
Article
| Open Access cAMP and c-di-GMP synergistically support biofilm maintenance through the direct interaction of their effectors
cAMP and c-di-GMP synergistically support biofilm maintenance through the direct interaction of their effectors
Nucleotide second messengers, such as cAMP and c-di-GMP, regulate many physiological processes in bacteria, including biofilm formation. Here, the authors provide evidence of cross-talk between cAMP and c-di-GMP pathways through direct interaction of their effectors, showing that the cAMP receptor protein (CRP) can play regulatory roles at the post-translational level.
- Cong Liu
- , Di Sun
- & Weijie Liu
-
Article
| Open Access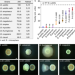 Resolving the conflict between antibiotic production and rapid growth by recognition of peptidoglycan of susceptible competitors
Resolving the conflict between antibiotic production and rapid growth by recognition of peptidoglycan of susceptible competitors
Microbial communities employ a variety of strategies to compete against competitors sharing their niche, for instance, by producing antibiotics. This study reveals that antibiotics produced by Bacillus subtilis act synergistically to eliminate phylogenetically distinct competitors and are regulated accordingly.
- Harsh Maan
- , Maxim Itkin
- & Ilana Kolodkin-Gal
-
Article
| Open Access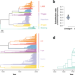 Genomic signatures of pre-resistance in Mycobacterium tuberculosis
Genomic signatures of pre-resistance in Mycobacterium tuberculosis
Signals of antimicrobial resistance in pathogen genomes may be detectable before the organism evolves an antimicrobial resistance phenotype. Here, the authors investigate this hypothesis using Mycobacterium tuberculosis data from Peru and identify candidate “pre-resistance” markers.
- Arturo Torres Ortiz
- , Jorge Coronel
- & Louis Grandjean
-
Article
| Open Access Dormant spores sense amino acids through the B subunits of their germination receptors
Dormant spores sense amino acids through the B subunits of their germination receptors
Germination of Bacillus subtilis spores in response to L-alanine requires a putative membrane receptor consisting of three proteins. Here, Artzi et al. use evolutionary co-variation analysis and functional assays of mutants to provide evidence that one of the proteins, GerAB, likely acts as the L-alanine sensor.
- Lior Artzi
- , Assaf Alon
- & David Z. Rudner
-
Article
| Open Access Bacterial chromosomal mobility via lateral transduction exceeds that of classical mobile genetic elements
Bacterial chromosomal mobility via lateral transduction exceeds that of classical mobile genetic elements
It is commonly thought that horizontal transfer of most bacterial chromosomal genes is limited, in comparison with the frequent transfer of mobile genetic elements. Humphrey et al. show that, actually, phage-mediated lateral transduction of core chromosomal genes can be more efficient than the transfer of mobile genetic elements via conjugation or generalized transduction.
- Suzanne Humphrey
- , Alfred Fillol-Salom
- & José R. Penadés
-
Comment
| Open Access Is the bacterial chromosome a mobile genetic element?
Is the bacterial chromosome a mobile genetic element?
An outcome of phage infection, lateral transduction, has been shown to mobilize chromosomal genes between bacterial cells at rates that exceed those of mobile genetic elements such as plasmids. Does this mean that the bacterial chromosome should be considered a mobile genetic element?
- James P. J. Hall
-
Article
| Open Access Naturally occurring fire coral clones demonstrate a genetic and environmental basis of microbiome composition
Naturally occurring fire coral clones demonstrate a genetic and environmental basis of microbiome composition
The microbiomes associated with reef corals are complex and diverse. Here, the authors investigate fire coral clones naturally occurring in distinct habitats as a model system to disentangle the contribution of host genotype and environment on their microbiome, and predict genomic functions based on taxonomic profiles.
- C. E. Dubé
- , M. Ziegler
- & C. R. Voolstra
-
Article
| Open Access Shape shifter: redirection of prolate phage capsid assembly by staphylococcal pathogenicity islands
Shape shifter: redirection of prolate phage capsid assembly by staphylococcal pathogenicity islands
Phage-inducible chromosomal islands (PICIs) are a group of mobile genetic elements that hijack the replication and assembly machinery of helper bacteriophages. Here the authors describe a mechanism by which a group of PICIs from Staphylococcus aureus re-direct the assembly pathway of their helpers using a capsid protein homolog.
- N’Toia C. Hawkins
- , James L. Kizziah
- & Terje Dokland
-
Article
| Open Access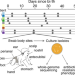 Rapid methicillin resistance diversification in Staphylococcus epidermidis colonizing human neonates
Rapid methicillin resistance diversification in Staphylococcus epidermidis colonizing human neonates
Staphylococcus epidermidis is a widespread early colonizer in the neonatal skin and a cause of hospital-acquired infections. Here, using whole-genome sequencing of 632 cultured S. epidermidis isolates derived from premature infants, the authors characterize the spatiotemporally strain-level genomic variability, finding patient-specific colonization signatures and a fast gain and loss of the antibiotic resistance gene mecA via the evolution of genotypically diverse structural variants.
- Manoshi S. Datta
- , Idan Yelin
- & Roy Kishony
-
Article
| Open Access Staphylococcal phages and pathogenicity islands drive plasmid evolution
Staphylococcal phages and pathogenicity islands drive plasmid evolution
Many plasmids can be transferred between bacterial cells via conjugation; however, the mechanisms underlying the transfer of non-conjugative plasmids are less clear. Here, Humphrey et al. show that staphylococcal phages and a family of pathogenicity islands (PICIs) can mediate intra- and inter-species plasmid transfer via generalised transduction.
- Suzanne Humphrey
- , Álvaro San Millán
- & José R. Penadés
-
Article
| Open Access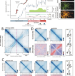 Spatial rearrangement of the Streptomyces venezuelae linear chromosome during sporogenic development
Spatial rearrangement of the Streptomyces venezuelae linear chromosome during sporogenic development
Streptomyces bacteria have a linear chromosome and a complex life cycle, including development of multi-genomic hyphae that differentiate into mono-genomic exospores. Here, Szafran et al. show that the chromosome of Streptomyces venezuelae undergoes substantial remodelling during sporulation, from an ‘open’ to a ‘closed’ conformation.
- Marcin J. Szafran
- , Tomasz Małecki
- & Dagmara Jakimowicz
-
Article
| Open Access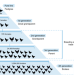 Global spread of Salmonella Enteritidis via centralized sourcing and international trade of poultry breeding stocks
Global spread of Salmonella Enteritidis via centralized sourcing and international trade of poultry breeding stocks
Salmonella enterica serotype Enteritidis is a pathogen of poultry that can cause outbreaks in humans. Here the authors use genomic and trade data to investigate a pandemic in the 1980s, finding evidence that international trade of breeding stocks led to global spread of the pathogen.
- Shaoting Li
- , Yingshu He
- & Xiangyu Deng
-
Article
| Open Access Evolutionary dynamics of multidrug resistant Salmonella enterica serovar 4,[5],12:i:- in Australia
Evolutionary dynamics of multidrug resistant Salmonella enterica serovar 4,[5],12:i:- in Australia
Salmonella enterica serovar 4,[5],12:i:- (Salmonella 4,[5],12:i:-) is a major pathogen of humans and animals with a reported incidence in Australia three times higher than the UK and USA. Here, the authors report the circulation, antimicrobial resistance signatures, and effects on host cells, of three Salmonella4,[5],12:i:- lineages within Australia.
- Danielle J. Ingle
- , Rebecca L. Ambrose
- & Deborah A. Williamson
-
Article
| Open Access Reversible gene silencing through frameshift indels and frameshift scars provide adaptive plasticity for Mycobacterium tuberculosis
Reversible gene silencing through frameshift indels and frameshift scars provide adaptive plasticity for Mycobacterium tuberculosis
Bacterial adaptation through frame-shifting insertions and deletions (indels) could be reversed by secondary introduction of a frame-restoring indel. Here, the authors develop ScarTrek, a program that scans genomic data for different indels, and analyze 5977 clinical M. tuberculosis isolates for indel frequency.
- Aditi Gupta
- & David Alland
-
Article
| Open Access In vivo evolution of an emerging zoonotic bacterial pathogen in an immunocompromised human host
In vivo evolution of an emerging zoonotic bacterial pathogen in an immunocompromised human host
Bordetella hinzii is an emerging pathogen with zoonotic risk to humans, known to be able to cause respiratory tract infection, bacteremia and endocarditis. Here, applying whole genome sequencing to bacterial isolates, the authors characterize the mechanisms driving adaptive evolution in B. hinzii in a patient with interleukin-12 receptor β1 deficiency, suggesting a role for host immune phenotype in shaping within-host pathogen evolution following zoonotic infection.
- A. Launay
- , C.-J. Wu
- & J. P. Dekker
-
Article
| Open Access Small RNA mediated gradual control of lipopolysaccharide biosynthesis affects antibiotic resistance in Helicobacter pylori
Small RNA mediated gradual control of lipopolysaccharide biosynthesis affects antibiotic resistance in Helicobacter pylori
The small RNA RepG modulates expression of chemotaxis receptor TlpB in Helicobacter pylori by targeting a length-variable G-repeat in the tlpB mRNA. Here, Pernitzsch et al. show that RepG also gradually controls lipopolysaccharide biosynthesis, antibiotic susceptibility, and in-vivo colonization of the stomach, by regulating a gene that is co-transcribed with tlpB.
- Sandy R. Pernitzsch
- , Mona Alzheimer
- & Cynthia M. Sharma
-
Article
| Open Access A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex
A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex
Klebsiella pneumoniae is a pathogen of increasing public health concern and antimicrobial resistance is becoming more prevalent. Here, the authors describe a K. pneumoniae genotyping tool, Kleborate, that can be used to identify lineages and detect antimicrobial resistance and virulence loci.
- Margaret M. C. Lam
- , Ryan R. Wick
- & Kathryn E. Holt
-
Article
| Open Access AsCas12a ultra nuclease facilitates the rapid generation of therapeutic cell medicines
AsCas12a ultra nuclease facilitates the rapid generation of therapeutic cell medicines
The utility of AsCas12a can be limited to poor editing efficiency. Here the authors identify a variant, “AsCas12a Ultra”, that has high on-target specificity demonstrated through editing of clinically relevant T cell genes.
- Liyang Zhang
- , John A. Zuris
- & Christopher A. Vakulskas
-
Article
| Open Access Phylogenomic analysis reveals persistence of gonococcal strains with reduced-susceptibility to extended-spectrum cephalosporins and mosaic penA-34
Phylogenomic analysis reveals persistence of gonococcal strains with reduced-susceptibility to extended-spectrum cephalosporins and mosaic penA-34
Resistance of Neisseria gonorrhoeae to extended spectrum cephalosporins is an increasing concern. Here, the authors conduct whole genome sequencing of isolates from the United States and find that most resistant isolates were associated with a persistent circulating lineage.
- Jesse C. Thomas IV
- , Sandeep J. Joseph
- & Zach Perry
-
Article
| Open Access The extracellular contractile injection system is enriched in environmental microbes and associates with numerous toxins
The extracellular contractile injection system is enriched in environmental microbes and associates with numerous toxins
The extracellular Contractile Injection System (eCIS) is a toxin-delivery particle that mediates interactions between bacteria and their invertebrate hosts. Here, the authors catalogue eCIS loci from 1,249 prokaryotic genomes, showing enrichment in non-pathogenic environmental microbes, and identifying eCIS-associated toxins that inhibit the growth of bacteria and/or yeast.
- Alexander Martin Geller
- , Inbal Pollin
- & Asaf Levy
-
Article
| Open Access Acinetobacter baylyi regulates type IV pilus synthesis by employing two extension motors and a motor protein inhibitor
Acinetobacter baylyi regulates type IV pilus synthesis by employing two extension motors and a motor protein inhibitor
Type IV pili (T4P) are retractile appendages used by bacteria for DNA uptake and other purposes. T4P extension is thought to occur through the action of a single motor protein, PilB. Here, Ellison et al. show that T4P synthesis in Acinetobacter baylyi depends not only on PilB but also on an additional, distinct motor, TfpB.
- Courtney K. Ellison
- , Triana N. Dalia
- & Ankur B. Dalia
-
Article
| Open Access Kin discrimination promotes horizontal gene transfer between unrelated strains in Bacillus subtilis
Kin discrimination promotes horizontal gene transfer between unrelated strains in Bacillus subtilis
Genetically distinct swarms of the bacterium Bacillus subtilis attack each other, forming a boundary upon encounter. Here, the authors show that these swarm antagonisms promote transformation-mediated horizontal gene transfer between strains of low relatedness.
- Polonca Stefanic
- , Katarina Belcijan
- & Ines Mandic-Mulec
-
Article
| Open Access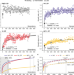 Compensatory evolution of Pseudomonas aeruginosa’s slow growth phenotype suggests mechanisms of adaptation in cystic fibrosis
Compensatory evolution of Pseudomonas aeruginosa’s slow growth phenotype suggests mechanisms of adaptation in cystic fibrosis
Long-term infection of cystic fibrosis patients with Pseudomonas aeruginosa is often accompanied by a reduction in bacterial growth rate. Here, La Rosa et al. use adaptive laboratory evolution to increase the growth rate of clinical isolates, and identify mechanisms and evolutionary trajectories that, in reverse direction, may help the pathogen to adapt to the patients’ airways.
- Ruggero La Rosa
- , Elio Rossi
- & Søren Molin
-
Article
| Open Access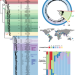 Global population structure and genotyping framework for genomic surveillance of the major dysentery pathogen, Shigella sonnei
Global population structure and genotyping framework for genomic surveillance of the major dysentery pathogen, Shigella sonnei
Whole genome sequencing is increasingly being adopted for Shigella sonnei outbreak investigation and surveillance, but there is no global classification standard. Here, the authors develop and validate a genomic framework implemented using open-source software, and demonstrate its application using surveillance data.
- Jane Hawkey
- , Kalani Paranagama
- & Kathryn E. Holt
-
Article
| Open Access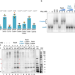 RNA binding of Hfq monomers promotes RelA-mediated hexamerization in a limiting Hfq environment
RNA binding of Hfq monomers promotes RelA-mediated hexamerization in a limiting Hfq environment
RelA stimulates RyhB small RNA–target mRNA interaction by promoting assembly of Hfq monomers into hexamers. Here the authors show that RelA-mediated Hfq hexamerization requires an initial binding of RNA to Hfq monomers.
- Pallabi Basu
- , Maya Elgrably-Weiss
- & Shoshy Altuvia
-
Article
| Open Access Bacterial cyclic diguanylate signaling networks sense temperature
Bacterial cyclic diguanylate signaling networks sense temperature
Many bacteria use the second messenger cyclic diguanylate (c-di-GMP) to control motility, biofilm production and virulence. Here, the authors identify a thermosensitive enzyme that synthesizes c-di-GMP and modulates temperature-dependent motility, biofilm development and virulence in the opportunistic pathogen Pseudomonas aeruginosa.
- Henrik Almblad
- , Trevor E. Randall
- & Joe Jonathan Harrison
-
Article
| Open Access Pili allow dominant marine cyanobacteria to avoid sinking and evade predation
Pili allow dominant marine cyanobacteria to avoid sinking and evade predation
It was thought that marine cyanobacteria drifted randomly in the water column. Here the authors show that one in four picocyanobacteria encode a type IV pilus which allows these organisms to increase drag and retain optimal positions in the water column, as well as evade predation by grazers.
- Maria del Mar Aguilo-Ferretjans
- , Rafael Bosch
- & Joseph A. Christie-Oleza
-
Article
| Open Access Control of a programmed cell death pathway in Pseudomonas aeruginosa by an antiterminator
Control of a programmed cell death pathway in Pseudomonas aeruginosa by an antiterminator
In Pseudomonas aeruginosa, the protein AlpA activates the expression of the alp locus in response to DNA damage, leading to lysis in a subset of cells and enhancing virulence of other, surviving cells. Here, the authors show that AlpA acts as an antiterminator rather than a transcriptional activator.
- Jennifer M. Peña
- , Samantha M. Prezioso
- & Simon L. Dove
-
Article
| Open Access A meet-up of two second messengers: the c-di-AMP receptor DarB controls (p)ppGpp synthesis in Bacillus subtilis
A meet-up of two second messengers: the c-di-AMP receptor DarB controls (p)ppGpp synthesis in Bacillus subtilis
In several bacteria, cyclic di-AMP mediates potassium (K+) and osmotic homeostasis. Here, the authors show that DarB, a Bacillus subtilis protein previously reported to bind cyclic di-AMP, interacts with the (p)ppGpp synthetase/hydrolase Rel in a K+-dependent manner in turn leading to Rel-dependent accumulation of pppGpp under conditions of K+ starvation.
- Larissa Krüger
- , Christina Herzberg
- & Jörg Stülke
-
Article
| Open Access Multiplexed characterization of rationally designed promoter architectures deconstructs combinatorial logic for IPTG-inducible systems
Multiplexed characterization of rationally designed promoter architectures deconstructs combinatorial logic for IPTG-inducible systems
Precisely tuning the genetic response to environmental stimuli is a key step in engineering synthetic biology systems. Here, the authors profile 8269 IPTG-induced promoters to deconstruct the relationship between sequence architecture and gene expression.
- Timothy C. Yu
- , Winnie L. Liu
- & Guillaume Urtecho

 Structure-function studies reveal ComEA contains an oligomerization domain essential for transformation in gram-positive bacteria
Structure-function studies reveal ComEA contains an oligomerization domain essential for transformation in gram-positive bacteria