Abstract
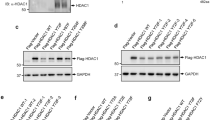
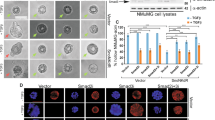
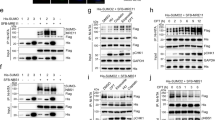
ING2 (inhibitor of growth 2) is a candidate tumor-suppressor gene involved in cell cycle control, apoptosis and senescence. Although the functions of ING2 within the chromatin remodeling complex Sin3A/histone deacetylase (HDAC) and in the p53 pathway have been described, how ING2 itself is regulated remains unknown. In this study we report for the first time that ING2 can be sumoylated by small ubiquitin-like modifier 1 (SUMO1) on lysine 195 both in vitro and in vivo. Strikingly, ING2 sumoylation enhances its association with Sin3a. We provide evidences that ING2 can bind to the promoter of genes to mediate their expression and that sumoylation of ING2 is required for this binding to some of these genes. Among them, we identified the gene TMEM71 (transmembrane protein 71), whose expression is regulated by ING2 sumoylation. ING2 must be sumoylated to bind to the promoter of TMEM71 and to recruit the Sin3A chromatin-modifying complex to this promoter, in order to regulate TMEM71 transcription. Hence, sumoylation of ING2 enhances its binding to the Sin3A/HDAC complex and is required to regulate gene transcriptions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bernier-Villamor V, Sampson DA, Matunis MJ, Lima CD . (2002). Structural basis for E2-mediated SUMO conjugation revealed by a complex between ubiquitin-conjugating enzyme Ubc9 and RanGAP1. Cell 108: 345–356.
Bienz M . (2006). The PHD finger, a nuclear protein-interaction domain. Trends Biochem Sci 31: 35–40.
Borkosky SS, Gunduz M, Nagatsuka H, Beder LB, Gunduz E, Ali MA et al. (2009). Frequent deletion of ING2 locus at 4q35.1 associates with advanced tumor stage in head and neck squamous cell carcinoma. J Cancer Res Clin Oncol 135: 703–713.
Dannenberg JH, David G, Zhong S, van der Torre J, Wong WH, Depinho RA . (2005). mSin3A corepressor regulates diverse transcriptional networks governing normal and neoplastic growth and survival. Genes Dev 19: 1581–1595.
Doyon Y, Cayrou C, Ullah M, Landry AJ, Cote V, Selleck W et al. (2006). ING tumor suppressor proteins are critical regulators of chromatin acetylation required for genome expression and perpetuation. Mol Cell 21: 51–64.
Farhana L, Dawson MI, Dannenberg JH, Xu L, Fontana JA . (2009). SHP and Sin3A expression are essential for adamantyl-substituted retinoid-related molecule-mediated nuclear factor-kappaB activation, c-Fos/c-Jun expression, and cellular apoptosis. Mol Cancer Ther 8: 1625–1635.
Feng X, Bonni S, Riabowol K . (2006). HSP70 induction by ING proteins sensitizes cells to tumor necrosis factor alpha receptor-mediated apoptosis. Mol Cell Biol 26: 9244–9255.
Garkavtsev I, Kazarov A, Gudkov A, Riabowol K . (1996). Suppression of the novel growth inhibitor p33ING1 promotes neoplastic transformation. Nat Genet 14: 415–420.
Gill G . (2005). Something about SUMO inhibits transcription. Curr Opin Genet Dev 15: 536–541.
Gong W, Russell M, Suzuki K, Riabowol K . (2006). Subcellular targeting of p33ING1b by phosphorylation-dependent 14-3-3 bbinding regulates p21WAF1 expression. Mol Cell Biol 26: 2947–2954.
Gozani O, Karuman P, Jones DR, Ivanov D, Cha J, Lugovskoy AA et al. (2003). The PHD finger of the chromatin-associated protein ING2 functions as a nuclear phosphoinositide receptor. Cell 114: 99–111.
Hay RT . (2005). SUMO: a history of modification. Mol Cell 18: 1–12.
Hietakangas V, Anckar J, Blomster HA, Fujimoto M, Palvimo JJ, Nakai A et al. (2006). PDSM, a motif for phosphorylation-dependent SUMO modification. Proc Natl Acad Sci USA 103: 45–50.
Hong Y, Rogers R, Matunis MJ, Mayhew CN, Goodson ML, Park-Sarge OK et al. (2001). Regulation of heat shock transcription factor 1 by stress-induced SUMO-1 modification. J Biol Chem 276: 40263–40267.
Larrieu D, Ythier D, Binet R, Brambilla C, Brambilla E, Sengupta S et al. (2009). ING2 controls the progression of DNA replication forks to maintain genome stability. EMBO Rep 10: 1168–1174.
Larrieu D, Pedeux R . (2009). SharING out the roles in replicatING DNA. Cell Cycle 8: 3623–3624.
Lu F, Dai DL, Martinka M, Ho V, Li G . (2006). Nuclear ING2 expression is reduced in human cutaneous melanomas. Br J Cancer 95: 80–86.
Matunis MJ, Coutavas E, Blobel G . (1996). A novel ubiquitin-like modification modulates the partitioning of the Ran-GTPase-activating protein RanGAP1 between the cytosol and the nuclear pore complex. J Cell Biol 135: 1457–1470.
Mellor J . (2006). It takes a PHD to read the histone code. Cell 126: 22–24.
Morimoto I, Sasaki Y, Ishida S, Imai K, Tokino T . (2002). Identification of the osteopontin gene as a direct target of TP53. Genes Chromosomes Cancer 33: 270–278.
Nagashima M, Shiseki M, Miura K, Hagiwara K, Linke SP, Pedeux R et al. (2001). DNA damage-inducible gene p33ING2 negatively regulates cell proliferation through acetylation of p53. Proc Natl Acad Sci USA 98: 9671–9676.
Nagashima M, Shiseki M, Pedeux RM, Okamura S, Kitahama-Shiseki M, Miura K et al. (2003). A novel PHD-finger motif protein, p47ING3, modulates p53-mediated transcription, cell cycle control, and apoptosis. Oncogene 22: 343–350.
Pedeux R, Sengupta S, Shen JC, Demidov ON, Saito S, Onogi H et al. (2005). ING2 regulates the onset of replicative senescence by induction of p300-dependent p53 acetylation. Mol Cell Biol 25: 6639–6648.
Rodriguez MS, Dargemont C, Hay RT . (2001). SUMO-1 conjugation in vivo requires both a consensus modification motif and nuclear targeting. J Biol Chem 276: 12654–12659.
Sarker KP, Kataoka H, Chan A, Netherton SJ, Pot I, Huynh MA et al. (2008). ING2 as a novel mediator of transforming growth factor-beta-dependent responses in epithelial cells. J Biol Chem 283: 13269–13279.
Sengupta S, Linke SP, Pedeux R, Yang Q, Farnsworth J, Garfield SH et al. (2003). BLM helicase-dependent transport of p53 to sites of stalled DNA replication forks modulates homologous recombination. EMBOJ 22: 1210–1222.
Shi X, Hong T, Walter KL, Ewalt M, Michishita E, Hung T et al. (2006). ING2 PHD domain links histone H3 lysine 4 methylation to active gene repression. Nature 442: 96–99.
Shiseki M, Nagashima M, Pedeux RM, Kitahama-Shiseki M, Miura K, Okamura S et al. (2003). p29ING4 and p28ING5 bind to p53 and p300, and enhance p53 activity. Cancer Res 63: 2373–2378.
Skowyra D, Zeremski M, Neznanov N, Li M, Choi Y, Uesugi M et al. (2001). Differential association of products of alternative transcripts of the candidate tumor suppressor ING1 with the mSin3/HDAC1 transcriptional corepressor complex. J Biol Chem 276: 8734–8739.
Terui Y, Saad N, Jia S, McKeon F, Yuan J . (2004). Dual role of sumoylation in the nuclear localization and transcriptional activation of NFAT1. J Biol Chem 279: 28257–28265.
Ythier D, Larrieu D, Brambilla C, Brambilla E, Pedeux R . (2008). The new tumor suppressor genes ING: genomic structure and status in cancer. Int J Cancer 123: 1483–1490.
Ythier D, Brambilla E, Binet R, Nissou D, Vesin A, de Fraipont F et al. (2009). Expression of candidate tumor suppressor gene ING2 is lost in non-small cell lung carcinoma. Lung Cancer 69: 180–186.
Zhang HK, Pan K, Wang H, Weng DS, Song HF, Zhou J et al. (2008). Decreased expression of ING2 gene and its clinicopathological significance in hepatocellular carcinoma. Cancer Lett 261: 183–192.
Acknowledgements
We thank D Nissou for technical assisstance; Dr O Gozani and C Harris for antibodies, plasmids and scientific discussion, and Dr Marc Piechaczyk and Dr Guillaume Bossis for scientific discussion and advising. RP was supported by ARC, the IASLC, an ‘Agir à dom’ grant and a Marie Curie International Reintegration Grant (MIRG-CT-2006-042148). DY, RB and DL were funded by the INCa, ARC, the FRM (Prix Mariane Josso) and the French Ministry of Education and Research, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Ythier, D., Larrieu, D., Binet, R. et al. Sumoylation of ING2 regulates the transcription mediated by Sin3A. Oncogene 29, 5946–5956 (2010). https://doi.org/10.1038/onc.2010.325
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2010.325
Keywords
This article is cited by
-
Identification and validation of an m6A-related gene signature to predict prognosis and evaluate immune features of breast cancer
Human Cell (2022)
-
ING2 tumor suppressive protein translocates into mitochondria and is involved in cellular metabolism homeostasis
Oncogene (2021)
-
INGs are potential drug targets for cancer
Journal of Cancer Research and Clinical Oncology (2017)
-
A novel crosstalk between the tumor suppressors ING1 and ING2 regulates androgen receptor signaling
Journal of Molecular Medicine (2016)
-
Epigenetic basis of opiate suppression of Bdnf gene expression in the ventral tegmental area
Nature Neuroscience (2015)