Abstract
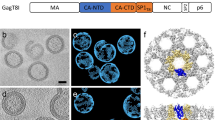
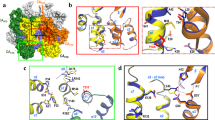
The capsid protein (CA) of the mature human immunodeficiency virus (HIV) contains an N-terminal β-hairpin that is essential for formation of the capsid core particle. CA is generated by proteolytic cleavage of the Gag precursor polyprotein during viral maturation. We have determined the NMR structure of a 283-residue N-terminal fragment of immature HIV-1 Gag (Gag283), which includes the intact matrix (MA) and N-terminal capsid (CAN) domains. The β-hairpin is unfolded in Gag283, consistent with the proposal that hairpin formation occurs subsequent to proteolytic cleavage of Gag, triggering capsid assembly. Comparison of the immature and mature CAN structures reveals that β-hairpin formation induces a ∼2 Å displacement of helix 6 and a concomitant displacement of the cyclophylin-A (CypA)-binding loop, suggesting a possible allosteric mechanism for CypA-mediated destabilization of the capsid particle during infectivity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Coffin, J.M., Hughes, S.H. & Varmus, H.E. Retroviruses (Cold Spring Harbor Laboratory Press, Plainview, New York; 1997).
Vogt, V.M. & Simon, M.N. Mass determination of Rous sarcoma virus virions by scaning transmission electron microscopy. J. Virol. 73, 7050–7055 (1999).
Turner, B.G. & Summers, M.F. Structural biology of HIV. J. Mol. Biol. 285, 1–32 (1999).
Gitti, R.K. et al. Structure of the amino-terminal core domain of the HIV-1 capsid protein. Science 273, 231–235 (1996).
Gamble, T.R. et al. Crystal structure of human cyclophilin A bound to the amino-terminal domain of HIV-1 capsid. Cell 87, 1285–1294 (1996).
Momany, C. et al. Crystal structure of dimeric HIV-1 capsid protein. Nature Struct. Biol. 9, 763–770 (1996).
Gamble, T.R. et al. Structure of the carboxyl-terminal dimerization domain of the HIV-1 capsid protein. Science 278, 849–853 (1997).
Worthylake, D.K., Wang, H., Yoo, S., Sundquist, W.I. & Hill, C.P. Structures of the HIV-1 capsid protein dimerization domain at 2.6 Å resolution. Acta Crystallogr. D 55, 85–92 (1999).
Berthet-Colominas, C. et al. Head-to-tail dimers and interdomain flexibility revealed by the crystal structure of HIV-1 capsid protein (p24) complexed with a monoclonal antibody Fab. EMBO J. 18, 1124–1136 (1999).
Kingston, R.L. et al. Structure and self-association of the Rous sarcoma virus capsid protein. Structure 8, 617–628 (2000).
Cornilescu, C.C., Bouamr, F., Yao, X., Carter, C. & Tjandra, N. Structural analysis of the N-terminal domain of the human T-cell leukemia virus capsid protein. J. Mol. Biol. 306, 783–797 (2001).
von Schwedler, U.K. et al. Proteolytic refolding of the HIV-1 capsid protein amino-terminus facilitates viral core assembly. EMBO J. 17, 1555–1568 (1998).
Gross, I., Hohenberg, H., Juckhagel, C. & Krausslich, H.-G. N-terminal extension of human immunodeficiency virus capsid protein converts the in vitro assembly phenotype from tubular to spherical particles. J. Virol. 72, 4798–4810 (1998).
Braaten, D., Franke, E.K. & Luban, J. Cyclophilin A is required for an early step in the life cycle of human immunodeficiency virus type-1 before the initiation of reverse transcription. J. Virol. 70, 3551–3560 (1996).
Franke, E.K., Yuan, H.E. & Luban, J. Specific incorporation of cyclophilin A into HIV-1 virions. Nature 24, 359–362 (1994).
Luban, J., Franke, E.K. & Yuan, H.E. HIV-1 uses host-cell proteins to form fully infectious virions. Nature 7, 37–42 (1994).
Luban, J., Bossolt, K.L., Franke, E.K., Kalpana, G.V. & Goff, S.P. Human immunodeficiency virus type 1 gag protein binds to cyclophilins A and B. Cell 73, 1067–1078 (1993).
Thali, M. et al. Functional association of cyclophilin A with HIV-1 virions. Nature 372, 363–365 (1994).
Ackerson, B., Rey, O., Canon, J. & Krogstad, P. Cells with high cyclophilin A content support replication of Human Immunodeficiency Virus Type-1 Gag mutants with decreased ability to incorporate Cyclophilin A. J. Virol. 72, 303–308 (1998).
Rothman, J.E. & Schmid, S.L. Enzymatic recycling of clathrin from coated vesicles. Cell 46, 5–9 (1986).
Adachi, A. et al. Production of acquired immunodeficiency syndrome-associated retrovirus in human and nonhuman cells transfected with an infectious molecular clone. J. Virol. 59, 284–291 (1986).
Massiah, M.A. et al. Three-dimensional structure of the human immunodeficiency virus type 1 matrix protein. J. Mol. Biol. 244, 198–223 (1994).
Hill, C.P., Worthylake, D., Bancroft, D.P., Christensen, A.M. & Sundquist, W.I. Crystal structures of the trimeric HIV-1 matrix protein: implications for membrane association. Proc. Natl. Acad. Sci. USA 93, 3099–3104 (1996).
Massiah, M.A. et al. Comparison of the NMR and X-ray structures of the HIV-1 matrix protein: evidence for conformational changes during viral assembly. Protein Sci. 5, 2391–2398 (1996).
Nermut, M.V., Frank, H. & Schafer, W. Properties of mouse leukemia viruses. III. Electron microscopic appearance as revealed after conventional preparation techniques as well as freezing-drying and freezing-etching. Virology 49, 345–358 (1972).
Bolognesi, D.P., Luftig, R. & Shaper, J.H. Localization of RNA tumor virus polypeptides I. Isolation of futher virus substructures. Virology 56, 549–564 (1973).
Fuller, S.D., Wilk, T., Gowen, B.E., Krausslich, H.-G. & Vogt, V.M. Cryo-electron microscopy reveals ordered domains in the immature HIV-1 particle. Curr. Biol. 7, 729–738 (1997).
Wilk, T. et al. Organization of immature Human Immunodeficiency Virus Type 1. J. Virol. 75, 759–771 (2001).
Nermut, M.V. et al. Further evidence for hexagonal organization of HIV gag protein in prebudding assemblies and immature virus-like particles. J. Struct. Biol. 123, 143–149 (1998).
Fitzon, T. et al. Proline residues in the HIV-1 NH2-terminal capsid domain: structure determinants for proper core assembly and subsequent steps of early replication. Virology 268, 294–307 (2000).
Tang, S. et al. Human immunodeficiency virus type-1 N-terminal capsid mutants that show aberrant core morphology and are blocked in initiation of reverse transcription in infected cells. J. Virol. 75, 9357–9366 (2001).
Kong, L.B. et al. Cryoelectron microscopy examination of human immunodeficiency virus type-1 virions with mutations in the cyclophilin A binding loop. J. Virol. 72, 4403–4407 (1998).
Yin, L., Braaten, D. & Luban, J. Human Immunodeficiency Virus Type-1 replication is modulated by host cyclophilin A expression levels. J. Virol. 72, 6430–6436 (1998).
Campos-Olivas, R. & Summers, M.F. Backbone dynamics of the N-terminal domain of the HIV-1 capsid protein and comparison with the G94D mutant conferring cyclosporin resistance/dependence. Biochemistry 38, 10262–10271 (1999).
Bosco, D.A., Eisenmesser, E.Z., Pochapsky, S., Sundquist, W.I. & Kern, D. Catalysis of cis/trans isomerization in native HIV-1 capsid by human cyclophilin A. Proc. Natl. Acad. Sci. USA 99, 5247–5252 (2002).
Bistrow, R. et al. Human cyclophilin has a significantly higher affinity for HIV-1 recombinant p55 than p24. J. Acquir. Immune Defic. Syndr. Hum. Retrovirol. 20, 334–336 (1999).
Yoo, S. et al. Molecular recognition in the HIV-1 capsid/cyclophilin A complex. J. Mol. Biol. 269, 780–795 (1997).
Rossmann, M.G. Viral cell recognition and entry. Protein Sci. 3, 1712–1725 (1994).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, Cold Spring Harbor; 1989).
Grzesiek, S. & Bax, A. The importance of not saturating H2O in protein NMR. Application to sensitivity enhancement and NOE measurement. J. Am. Chem. Soc. 115, 12593–12593 (1993).
Grzesiek, S. & Bax, A. Improved 3D triple-resonance NMR techniques applied to a 31 kDa protein. J. Magn. Res. 96, 432–440 (1992).
Delaglio, F. et al. NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Johnson, B.A. & Blevins, R.A. NMRView: a computer program for the visualization and analysis for NMR data. J. Biomol. NMR 4, 603–614 (1994).
Wüthrich, K. NMR of Proteins and Nucleic Acids (John Wiley & Sons, New York; 1986).
Clore, G.M. & Gronenborn, A.M. Structures of larger proteins in solution: three- and four-dimensional heteronuclear NMR spectroscopy. Science 252, 1390–1399 (1991).
Bax, A. & Grzesiek, S. Methodological advances in protein NMR. Acc. Chem. Res. 26, 131–138 (1993).
Wishart, D.S., Sykes, B.D. & Richards, F.M. The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31, 1647–1651 (1992).
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR. 13, 289–302 (1999).
Güntert, P., Mumenthaler, C. & Wüthrich, K. Torsion angle dynamics for protein structure calculations with a new program, DYANA. J. Mol. Biol. 273, 283–298 (1997).
Laskowski, R.A., Rullmann, J.A.C., MacArthur, M.W., Kaptein, R. & Thornton, J.M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Ferrin, T.E., Huang, C.C., Jarvis, L.E. & Langridge, R. The MIDAS display system. J. Mol. Graph. 6, 13–27 (1988).
Kraulis, P.J. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Cryst. 24, 946–950 (1991).
Merritt, E.A. & Bacon, D.J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997).
Kay, L.E., Torchia, D.A. & Bax, A. Backbone dynamics of proteins as studied by 15N inverse detected heteronuclear NMR spectroscopy: application to staphylococcal nuclease. Biochemistry 28, 8972–8979 (1989).
Tjandra, N., Kuboniwa, H., Ren, H. & Bax, A. Rotational dynamics of calcium-free calmodulin studied by 15N-NMR relaxation measurements. Eur. J. Biochem. 230, 1014–1024 (1995).
Lee, L.K., Rance, M., Chazin, W.J. & Palmer, AG. I. Rotational diffusion anisotropy of proteins from simultaneous analysis of 15N and 13Cα nuclear spin relaxation. J. Biomol. NMR 9, 287–298 (1997).
Acknowledgements
Support from the NIAID-NIH is gratefully acknowledged. We thank D. King (U.C. Berkeley) for mass spectral measurements and R. Edwards (HHMI, UMBC) for technical assistance. Y.N. is a Meyerhoff undergraduate scholar.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Tang, C., Ndassa, Y. & Summers, M. Structure of the N-terminal 283-residue fragment of the immature HIV-1 Gag polyprotein. Nat Struct Mol Biol 9, 537–543 (2002). https://doi.org/10.1038/nsb806
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb806
This article is cited by
-
Paediatric non-progression following grandmother-to-child HIV transmission
Retrovirology (2016)