Abstract
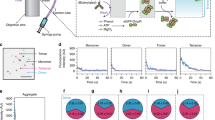
CooA is a homodimeric transcription factor that belongs to the catabolite activator protein (CAP) family. Binding of CO to the heme groups of CooA leads to the transcription of genes involved in CO oxidation in Rhodospirillum rubrum. The 2.6 Å structure of reduced (Fe2+) CooA reveals that His 77 in both subunits provides one heme ligand while the N-terminal nitrogen of Pro 2 from the opposite subunit provides the other ligand. A structural comparison of CooA in the absence of effector and DNA (off state) with that of CAP in the effector and DNA bound state (on state) leads to a plausible model for the mechanism of allosteric control in this class of proteins as well as the CO dependent activation of CooA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Busby, S. & Ebright, R.H. J. Mol. Biol. 293, 199–213 (1999).
Crasnier, M. Res. Microbiol. 147, 479–482 (1996).
McKay, D.B., Weber, I.T. & Steitz, T.A. J. Biol. Chem. 257, 9518– 9524 (1982).
Schultz, S.C., Sheilds, G.C. & Steitz, T.A. Science 253, 1001– 1007 (1991).
Vaney, M.C., Gilliland, G.L., Harman, J.G., Peterkofsky, A. & Weber, I.T. Biochemistry 28, 4567–4574 (1989).
Weber, I.T. & Steitz, T.A. J. Mol. Biol. 198, 311–326 (1987).
Craven, P.A. & DeRubertis, F.R. Biochim. Biophys. Acta 745, 310–321 (1983).
Ignarro, L.J., Degnan, J.N., Baricos, W.H., Kadowitz, P.J. & Wolin, M.S. Biochem. Biophys. Acta 718, 49–59 (1982).
Yu, A.E., Hu, S.Z., Spiro, T.G. & Burstyn, J.N. J. Am. Chem. Soc. 116, 4117–4118 ( 1994).
Gilles-Gonzalez, M.A., Gonzalez, G. & Perutz, M.P. Biochemistry 34, 232– 236 (1995).
Gilles-Gonzalez, M.A., Ditta, G.S. & Helinski, D.R. Nature 350, 170– 172 (1991).
Rodgers, K.R., Lukat-Rodgers, G.S. & Barron, J.A. Biochemistry 35, 9539– 9548 (1996).
Aono, S. H. N., Saito, K. & Okada, M. Biochem. Biophys. Res. Commun. 228, 752–756 (1996).
Shelver, D., Kerby, R.L., He, Y. & Roberts, G.P. Proc. Natl. Acad. Sci. USA 94, 11216–11220 (1997).
Parkinson, G., et al. J. Mol. Biol. 260, 395– 408 (1996).
Bell, C.E. & Lewis, M. Nature Struct. Biol. 7 , 209–214 (2000).
Cheng, X., Kovac, L. & Lee, J.C. Biochemistry 34, 10816– 10826 (1995).
Leu, S.F., Baker, C.H., Lee, E.J. & Harman, J.G. Biochemistry 38, 6222–6230 ( 1999).
Cheng, X. & Lee, J.C. Biochemistry 37, 51–60 (1998).
Shaw, D.J., Rice, D.W. & Guest, J.R. J. Mol. Biol. 166, 241– 247 (1983).
Kim, J., Adhya, S. & Garges, S. Proc. Natl. Acad. Sci. 89, 9700 –9704 (1992).
Ryu, S., Kim, J., Adhya, S. & Garges, S. Proc. Natl. Acad. Sci. 90, 75–79 ( 1993).
Garges, S. & Adhya, S. Cell 41, 745 –751 (1985).
Aiba, H., Nakamura, T., Mitani, H. & Mori, H. EMBO J. 4, 3329–3332 (1985).
Reynolds, M.F., et al. Biochemistry 39, 388– 396 (2000).
Thorsteinsson, M.V., Kerby, R.L. & Roberts, G.P. Biochemistry 39, 8284– 8290 (2000).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 ( 1997).
Collaborative Computational Project, Number 4. Acta Crystallogr. D 50, 760–763 (1994).
Fortelle, E. & Bricogne, G. Methods Enzymol. 276 , 472–494 (1997).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905– 921 (1998).
Acknowledgements
The authors would like to thank M. Sundaramoorthy and H. Li for help in the initial collection and processing of data. W.N.L. would also like to thank A.L. Pearcy for providing housing in Corona del Mar/Newport Beach during the completion of this work. The authors were supported by the United States Department of Agriculture (W.N.L.), National Science Foundation (T.L.P.), and National Institutes of Health (M.V.T. and G.P.R.). This work is based upon research conducted at the SSRL, which is funded by the Department of Energy (BES, BER) and the National Institutes of Health (NCRR, NIGMS).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lanzilotta, W., Schuller, D., Thorsteinsson, M. et al. Structure of the CO sensing transcription activator CooA. Nat Struct Mol Biol 7, 876–880 (2000). https://doi.org/10.1038/82820
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/82820
This article is cited by
-
Structural insights into glutathione-mediated activation of the master regulator PrfA in Listeria monocytogenes
Protein & Cell (2017)
-
Met104 is the CO-replaceable ligand at Fe(II) heme in the CO-sensing transcription factor BxRcoM-1
JBIC Journal of Biological Inorganic Chemistry (2016)
-
Structural and functional insights into the heme-binding domain of the human soluble guanylate cyclase α2 subunit and heterodimeric α2β1
JBIC Journal of Biological Inorganic Chemistry (2012)
-
Identification of Cys94 as the distal ligand to the Fe(III) heme in the transcriptional regulator RcoM-2 from Burkholderia xenovorans
JBIC Journal of Biological Inorganic Chemistry (2012)
-
Spectroscopic and computational investigation of three Cys-to-Ser mutants of nickel superoxide dismutase: insight into the roles played by the Cys2 and Cys6 active-site residues
JBIC Journal of Biological Inorganic Chemistry (2010)