Abstract
Condensin complexes have central roles in the three-dimensional organization of chromosomes during cell divisions, but how they interact with chromatin to promote chromosome segregation is largely unknown. Previous work has suggested that condensin, in addition to encircling chromatin fibers topologically within the ring-shaped structure formed by its SMC and kleisin subunits, contacts DNA directly. Here we describe the discovery of a binding domain for double-stranded DNA formed by the two HEAT-repeat subunits of the Saccharomyces cerevisiae condensin complex. From detailed mapping data of the interfaces between the HEAT-repeat and kleisin subunits, we generated condensin complexes that lack one of the HEAT-repeat subunits and consequently fail to associate with chromosomes in yeast and human cells. The finding that DNA binding by condensin's HEAT-repeat subunits stimulates the SMC ATPase activity suggests a multistep mechanism for the loading of condensin onto chromosomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Hirano, T. Condensins: universal organizers of chromosomes with diverse functions. Genes Dev. 26, 1659–1678 (2012).
Piazza, I., Haering, C.H. & Rutkowska, A. Condensin: crafting the chromosome landscape. Chromosoma 122, 175–190 (2013).
Aragón, L., Martinez-Perez, E. & Merkenschlager, M. Condensin, cohesin and the control of chromatin states. Curr. Opin. Genet. Dev. 23, 204–211 (2013).
Wood, A.J., Severson, A.F. & Meyer, B.J. Condensin and cohesin complexity: the expanding repertoire of functions. Nat. Rev. Genet. 11, 391–404 (2010).
Anderson, D.E., Losada, A., Erickson, H.P. & Hirano, T. Condensin and cohesin display different arm conformations with characteristic hinge angles. J. Cell Biol. 156, 419–424 (2002).
Schleiffer, A. et al. Kleisins: a superfamily of bacterial and eukaryotic SMC protein partners. Mol. Cell 11, 571–575 (2003).
Onn, I., Aono, N., Hirano, M. & Hirano, T. Reconstitution and subunit geometry of human condensin complexes. EMBO J. 26, 1024–1034 (2007).
Neuwald, A.F. & Hirano, T. HEAT repeats associated with condensins, cohesins, and other complexes involved in chromosome-related functions. Genome Res. 10, 1445–1452 (2000).
Ono, T., Fang, Y., Spector, D.L. & Hirano, T. Spatial and temporal regulation of condensins I and II in mitotic chromosome assembly in human cells. Mol. Biol. Cell 15, 3296–3308 (2004).
Kimura, K. & Hirano, T. ATP-dependent positive supercoiling of DNA by 13S condensin: a biochemical implication for chromosome condensation. Cell 90, 625–634 (1997).
Kimura, K., Rybenkov, V.V., Crisona, N.J., Hirano, T. & Cozzarelli, N.R. 13S condensin actively reconfigures DNA by introducing global positive writhe: implications for chromosome condensation. Cell 98, 239–248 (1999).
Kimura, K. & Hirano, T. Dual roles of the 11S regulatory subcomplex in condensin functions. Proc. Natl. Acad. Sci. USA 97, 11972–11977 (2000).
Stray, J.E. & Lindsley, J.E. Biochemical analysis of the yeast condensin Smc2/4 complex: an ATPase that promotes knotting of circular DNA. J. Biol. Chem. 278, 26238–26248 (2003).
Stray, J.E., Crisona, N.J., Belotserkovskii, B.P., Lindsley, J.E. & Cozzarelli, N.R. The Saccharomyces cerevisiae Smc2/4 condensin compacts DNA into (+) chiral structures without net supercoiling. J. Biol. Chem. 280, 34723–34734 (2005).
Sakai, A., Hizume, K., Sutani, T., Takeyasu, K. & Yanagida, M. Condensin but not cohesin SMC heterodimer induces DNA reannealing through protein-protein assembly. EMBO J. 22, 2764–2775 (2003).
Cuylen, S., Metz, J. & Haering, C.H. Condensin structures chromosomal DNA through topological links. Nat. Struct. Mol. Biol. 18, 894–901 (2011).
Haering, C.H., Farcas, A.-M., Arumugam, P., Metson, J. & Nasmyth, K. The cohesin ring concatenates sister DNA molecules. Nature 454, 297–301 (2008).
Tada, K., Susumu, H., Sakuno, T. & Watanabe, Y. Condensin association with histone H2A shapes mitotic chromosomes. Nature 474, 477–483 (2011).
Liu, W. et al. PHF8 mediates histone H4 lysine 20 demethylation events involved in cell cycle progression. Nature 466, 508–512 (2010).
Gajiwala, K.S. & Burley, S.K. Winged helix proteins. Curr. Opin. Struct. Biol. 10, 110–116 (2000).
Bürmann, F. et al. An asymmetric SMC–kleisin bridge in prokaryotic condensin. Nat. Struct. Mol. Biol. 20, 371–379 (2013).
Haering, C.H. et al. Structure and stability of cohesin's Smc1-kleisin interaction. Mol. Cell 15, 951–964 (2004).
Woo, J.-S. et al. Structural studies of a bacterial condensin complex reveal ATP-dependent disruption of intersubunit interactions. Cell 136, 85–96 (2009).
Yoshimura, S.H. et al. Condensin architecture and interaction with DNA: regulatory non-SMC subunits bind to the head of SMC heterodimer. Curr. Biol. 12, 508–513 (2002).
Griese, J.J., Witte, G. & Hopfner, K.-P. Structure and DNA binding activity of the mouse condensin hinge domain highlight common and diverse features of SMC proteins. Nucleic Acids Res. 38, 3454–3465 (2010).
Lowary, P.T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998).
Mavrich, T.N. et al. A barrier nucleosome model for statistical positioning of nucleosomes throughout the yeast genome. Genome Res. 18, 1073–1083 (2008).
Albert, I. et al. Translational and rotational settings of H2A.Z nucleosomes across the Saccharomyces cerevisiae genome. Nature 446, 572–576 (2007).
Amlacher, S. et al. Insight into structure and assembly of the nuclear pore complex by utilizing the genome of a eukaryotic thermophile. Cell 146, 277–289 (2011).
Leitner, A. et al. Probing native protein structures by chemical cross-linking, mass spectrometry, and bioinformatics. Mol. Cell. Proteomics 9, 1634–1649 (2010).
Leitner, A. et al. Expanding the chemical cross-linking toolbox by the use of multiple proteases and enrichment by size exclusion chromatography. Mol. Cell Proteomics 11, M111.014126 (2012).
Lavoie, B.D., Hogan, E. & Koshland, D. In vivo dissection of the chromosome condensation machinery: reversibility of condensation distinguishes contributions of condensin and cohesin. J. Cell Biol. 156, 805–815 (2002).
Cuylen, S., Metz, J., Hruby, A. & Haering, C.H. Entrapment of chromosomes by condensin rings prevents their breakage during cytokinesis. Dev. Cell 27, 469–478 (2013).
Neumann, B. et al. Phenotypic profiling of the human genome by time-lapse microscopy reveals cell division genes. Nature 464, 721–727 (2010).
Arumugam, P. et al. ATP hydrolysis is required for cohesin's association with chromosomes. Curr. Biol. 13, 1941–1953 (2003).
Weitzer, S., Lehane, C. & Uhlmann, F. A model for ATP hydrolysis-dependent binding of cohesin to DNA. Curr. Biol. 13, 1930–1940 (2003).
Arumugam, P., Nishino, T., Haering, C.H., Gruber, S. & Nasmyth, K. Cohesin's ATPase activity is stimulated by the C-terminal winged-helix domain of its kleisin subunit. Curr. Biol. 16, 1998–2008 (2006).
Akai, Y. et al. Opposing role of condensin hinge against replication protein A in mitosis and interphase through promoting DNA annealing. Open Biol. 1, 110023 (2011).
Rubinson, E.H., Gowda, A.S.P., Spratt, T.E., Gold, B. & Eichman, B.F. An unprecedented nucleic acid capture mechanism for excision of DNA damage. Nature 468, 406–411 (2010).
Ouspenski, I.I., Cabello, O.A. & Brinkley, B.R. Chromosome condensation factor Brn1p is required for chromatid separation in mitosis. Mol. Biol. Cell 11, 1305–1313 (2000).
Ivanov, D. & Nasmyth, K. A topological interaction between cohesin rings and a circular minichromosome. Cell 122, 849–860 (2005).
Gruber, S. et al. Evidence that loading of cohesin onto chromosomes involves opening of its SMC hinge. Cell 127, 523–537 (2006).
Hudson, D.F. et al. Molecular and genetic analysis of condensin function in vertebrate cells. Mol. Biol. Cell 19, 3070–3079 (2008).
Murayama, Y. & Uhlmann, F. Biochemical reconstitution of topological DNA binding by the cohesin ring. Nature 505, 367–371 (2014).
Fitzgerald, D.J. et al. Protein complex expression by using multigene baculoviral vectors. Nat. Methods 3, 1021–1032 (2006).
Luger, K., Rechsteiner, T.J. & Richmond, T.J. Preparation of nucleosome core particle from recombinant histones. Methods Enzymol. 304, 3–19 (1999).
Saravanan, M. et al. Interactions between the nucleosome histone core and Arp8 in the INO80 chromatin remodeling complex. Proc. Natl. Acad. Sci. USA 109, 20883–20888 (2012).
Xu, H., Zhang, L. & Freitas, M.A. Identification and characterization of disulfide bonds in proteins and peptides from tandem MS data by use of the MassMatrix MS/MS search engine. J. Proteome Res. 7, 138–144 (2008).
Rinner, O. et al. Identification of cross-linked peptides from large sequence databases. Nat. Methods 5, 315–318 (2008).
Walzthoeni, T. et al. False discovery rate estimation for cross-linked peptides identified by mass spectrometry. Nat. Methods 9, 901–903 (2012).
Notredame, C., Higgins, D.G. & Heringa, J. T-Coffee: a novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217 (2000).
Kelley, L.A. & Sternberg, M.J.E. Protein structure prediction on the Web: a case study using the Phyre server. Nat. Protoc. 4, 363–371 (2009).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Gerlich, D., Hirota, T., Koch, B., Peters, J.-M. & Ellenberg, J. Condensin I stabilizes chromosomes mechanically through a dynamic interaction in live cells. Curr. Biol. 16, 333–344 (2006).
Carpenter, A.E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Acknowledgements
We are grateful to M. Cohen, V. Rybin, M. Saravanan and C. Tischer for assistance with yeast experiments, biophysical assays, nucleosome preparation and image segmentation, to Y. Frosi for suggesting the mutant analysis in human cells and to S. Amlacher and E. Hurt (University of Heidelberg) for providing C. thermophilum cDNA and condensin sequences. We thank I. Berger for extensive advice and training in the MultiBac technology and the EMBL Advanced Light Microscopy, Genomics and Proteomics Core Facilities for technical support. We thank V. Benes and B. Baying for discussion, technical advice and help with the preparation of genomic libraries and sequencing. We thank F. Baudin, J. Ellenberg, D. Gilmour, F. Melchior, S. Milles, A. Musacchio, C. Müller and members of the Haering laboratory for discussion and advice. This work was supported by funding from the EMBL and the German Research Foundation (DFG) grant HA 5853/2-1 (C.H.H.). A.O. was supported by postdoctoral fellowships from the Alexander von Humboldt foundation and Marie Curie Actions.
Author information
Authors and Affiliations
Contributions
I.P., A.R., A.O., M.W., J.M. and C.H.H. designed and performed the experiments; I.P., A.O. and M.B. analyzed the cross-linking MS experiments; I.P. and V.P. performed bioinformatics analysis of generated and published ChIP-seq data; and I.P. and C.H.H. conceived the project and wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Purification of S. cerevisiae non-SMC subcomplexes.
(a) Purification strategy of non-SMC subcomplexes expressed in Sf21 insect cells using the MultiBac system. (b) Anion exchange (Source 15Q) and size exclusion chromatography (Superose 6) profiles of the purification of Brn1–Ycs4–Ycg1 subcomplexes. Peak fractions were analyzed by SDS PAGE and Coomassie staining. (c) Electron micrographs of negatively stained Brn1–Ycs4–Ycg1 and Brn1ΔNC–Ycs4–Ycg1 subcomplexes. (d) Purification of Brn1ΔNC–Ycs4–Ycg1 as in b.
Supplementary Figure 2 The non-SMC subcomplex binds DNA.
(a) The electrophoretic mobility of a 6.5-kb linear dsDNA at increasing S. cerevisiae Brn1–Ycs4–Ycg1 concentrations was resolved on a 0.8% native agarose gel and detected by ethidium bromide staining. Unbound (*), major (**) and minor (***) slower migrating species are indicated. (b) Sequences of the 15 to 60-bp dsDNA substrates; green asterisks indicate the positions of the 6-FAM labels. (c) Electrophoretic mobility shift assay of a 30-bp dsDNA substrate (lanes 1-6) after addition of unlabeled 30-bp competitor DNA (lanes 7-12) or an antibody against the His6 tag on Ycs4 (lanes 13-14). Unbound (*), shifted (**) and antibody super-shifted (***) species are indicated. (d) Electrophoretic mobility of the Brn1–Ycs4–Ycg1 subcomplex during native protein gel electrophoresis with or without addition of 30-bp dsDNA at 4:1 molar DNA/protein ratio. Unbound (*) and shifted (**) species detected by post-run Coomassie staining are indicated. (e) Electrophoretic mobility shift assay of a 6.5-kb linear dsDNA with Brn1ΔNC–Ycs4–Ycg1 as in a. (f) Binding affinities of Brn1–Ycs4–Ycg1 and Brn1ΔNC–Ycs4–Ycg1 to a 30-bp 6-FAM-labeled dsDNA substrate were compared by measuring fluorescence anisotropy changes upon addition of the indicated protein concentrations. Dissociation constants (Kd) were calculated by fitting mean ΔA values for each protein concentration assuming a single-site binding model. Points and error bars indicate mean and s.d. of n = 3 independent experiments. Panels a, c and e show one representative experiment of n = 3 independent replicates.
Supplementary Figure 3 Characterization of DNA- and chromosome-binding specificities of the non-SMC subcomplex.
(a) Coomassie-stained SDS PAGE of purified S. cerevisiae the Smc2–Smc4 hinge dimer and the Brn1–Ycs4–Ycg1 subcomplex used for DNA binding experiments. (b) Electrophoretic mobility shift and fluorescence anisotropy binding assays of 6-FAM-labeled 30-bp dsDNA and ssDNA substrates at increasing Smc2–Smc4 hinge concentrations. Unbound (*) and slower migrating species (**) are indicated. Points and error bars indicate mean and s.d. of 3 independent experiments. (c) Increasing concentrations of Brn1–Ycs4–Ycg1 (0.1–1.6 μM; lanes 1–6) or of the Smc2–Smc4 hinge (0.8–25.6 μM; lanes 7–13) were incubated with a fixed concentration of 30- bp 6-FAM-labeled dsDNA (0.2 μM) and binding was analyzed by agarose gel electrophoresis. Addition of the non-SMC subcomplex to pre-formed Smc2–Smc4 hinge–DNA complexes was analyzed at the same time (lanes 14–15). (d) Purified non-SMC subcomplex and reconstituted nucleosomes (Nuc-167) used in binding assays were analyzed by SDS PAGE and Coomassie staining. (e) The budding yeast genome was sub-divided into regions enriched or depleted of histone H2A.Z (S. cerevisiae Htz1; see Online Methods) and the relative enrichment of Brn1-PK6 in each of the two categories was determined for each chromosome in a ChIP-seq experiment. Horizontal lines define the median and boxes the 25th and 75th percentiles, whiskers represent the maximum and minimum values; P < 4×10–9 by Wilcoxon two-sided test. (f) The levels of condensin and histone H2A.Z bound in every 500-bp window of the budding yeast genome were expressed as relative enrichment of Brn1-PK6 or Htz1-TAP in ChIP-seq experiments were compared; the non-parametric correlation index (Spearman) is indicated. Panels b and c show one representative experiment of n = 3 independent replicates.
Supplementary Figure 4 Purification of the C. thermophilum non-SMC subcomplex and HEAT-repeat subunits.
(a) Gelfiltration chromatography profile of Ct Brn1ΔNC–Ycs4–Ycg1ΔC. Peak fractions were analyzed by SDS PAGE and Coomassie staining. (b) Gelfiltration chromatography of the Ct Ycs4 subunit as in a. (c) Gelfiltration chromatography of the Ct Ycg1ΔC subunit as in a.
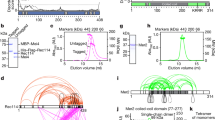
Supplementary Figure 5 Interaction interface mapping of Brn1, Ycs4 and Ycg1.
(a) The S. cerevisiae Brn1–Ycs4–Ycg1 subcomplex was incubated with the indicated concentrations of H12/D12-labeled DSS to covalently cross-link ɛ-amine groups of lysine residues within a distance of 11.4 Å. Cross-linking efficiency was analyzed by denaturing and native SDS PAGE and Coomassie staining. (b) Fragment ion spectrum of two cross-linked peptides. Ion series for both peptides are indicated, yellow peaks in the spectrum correspond to fragment ions containing the cross-linked lysine pair (cross-link peaks), blue peaks correspond to fragment ions that do not include the linked site (common peaks). All fragment ions that match a peak in the spectrum are marked with triangles. True positive hits can be separated from false positive hits (matching the decoy database) on the basis of their xQuest Linear Discriminant score (LD score).
Supplementary Figure 6 Mapping of Ycg1- and Ycs4-binding sites in Brn1.
(a) Equimolar amounts of C. thermophilum Ycs4-His6 and Ycg1-StrepII were mixed, purified over a Strep-Tactin resin, and the presence of the proteins in unbound (U) and bound (B) fractions analyzed by SDS PAGE and Coomassie staining. (b) Fragments of S. cerevisiae Brn1 of the indicated residue number range were expressed as Protein A fusion proteins in yeast and co-purification of endogenous condensin subunits on IgG beads was probed by western blotting against the PK9 tag on Ycs4, the myc9 tag on Smc4 or the HA6 tag on Ycg1 in input (IN), unbound (U), and bound (B; 10× concentrated compared to input) fractions after low (100 mM KCl) or high (300 mM KCl) salt wash steps. A band that results from binding of the anti-PK antibody by the full-length Brn1-ProtA in indicated (*). (c) Purification of a stable S. cerevisiae Brn1458–531–Ycg1 complex. The two proteins were co-expressed in insect cells, co-purified over Ni-NTA and loaded onto a Strep-Tactin resin. Input (IN), unbound (U), and bound (B) fractions of the Strep-Tactin resin were probed by SDS-PAGE and Coomassie staining.
Supplementary Figure 7 Characterization of Brn1 yeast mutants.
(a) Expression of wild type or mutant Brn1-PK6 constructs from one endogenous allele in diploid yeast strains (strains C3632, C3665, C3651, C3641, C3635, C3658, C3649 and C3634) was tested by western blotting of cell extracts against the PK epitope and tubulin as loading control. (b) Co-purification of Smc2-HA6 with wild type or mutant Brn1-PK6 from whole cell yeast extracts (strains C3632, C3665, C3651, C3641, C3635, C3658, C3649 and C3634) was probed by western blotting of input (IN), unbound (U), and bound (B, 10× concentrated compared to input) fractions with antibodies against the HA and PK epitopes. (c) Haploid yeast cells expressing wild type or mutant (M1 or M4) Brn1-PK6 from the endogenous BRN1 locus (strains C3890, C3892 and C3893) were arrested in G1 phase with mating pheromone and cell cycle progression after release at 37°C was monitored by FACScan analysis of DNA content. Segregation of chromosome V marked by the binding of TetR-GFP to tandem arrays of TetO sequences integrated proximal to the telomere of the right chromosome arm was monitored by live cell microscopy at the indicated time points. The fractions of cells from n > 200 cells per time point (sum from 3 independent experiments) are plotted. (d) Genomic binding sites of budding yeast condensin holocomplexes and condensin complexes lacking Ycg1. Relative enrichment of Brn1-PK6 wild type and M1 M2 M4 mutant at rRNA genes, tRNA genes, centromeres and coding regions of RNA pol II-transcribed genes (ORFs) is plotted. Horizontal lines define the median, boxes the 25th and 75th percentiles and whiskers the maximum and minimum values, extending up to 1.5-times the interquartile range. Outlier values are represented by circles.
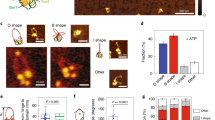
Supplementary Figure 8 Chromosome binding of mutant condensin I and II in human cells.
(a) Multi-sequence alignment of β-kleisin (condensin II) and γ-kleisin (condensin I) subunits. Boxed residues were mutated to the indicated sequences in the human CAP-H2 subunit. (b) The indicated Flag-EGFP-CAP-H2 proteins were immunoprecipitated from lysates of transiently transfected HEK 293 cells and co-precipitation of the other condensin II subunits was probed by western blotting against the condensin subunits. The slower migrating band of Flag-EGFP-CAP-H2 corresponds to a phosphorylated form that can be reverted by incubation with λ phosphatase (A.R., unpublished data). (c) Expression levels of Flag-EGFP fusion proteins of wild type and mutant CAP-H transiently expressed in HeLa H2B-mCherry cells were compared to the levels of endogenous CAP-H by western blotting of whole cell extracts with an antibody directed against CAP-H. (d) Flag-EGFP-CAPH2 proteins were transiently expressed in nocodazole-arrested HeLa cells expressing histone H2B-mCherry. Cells (yellow lines) and chromosomes (red lines) were segmented using total EGFP or mCherry signals, respectively, and mean EGFP intensities were measured in chromosome and cytoplasmic regions. (e) Ratios between chromosomal and cytoplasmic EGFP mean intensities were calculated from one representative experiment of 3 independent repeats and plotted ± s.d., n = 45 cells (wild type), 46 cells (M1), 55 cells (M2 M4) and 49 cells (M1 M2 M4).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1–5 (PDF 73750 kb)
Rights and permissions
About this article
Cite this article
Piazza, I., Rutkowska, A., Ori, A. et al. Association of condensin with chromosomes depends on DNA binding by its HEAT-repeat subunits. Nat Struct Mol Biol 21, 560–568 (2014). https://doi.org/10.1038/nsmb.2831
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2831
This article is cited by
-
The SAGA histone acetyltransferase module targets SMC5/6 to specific genes
Epigenetics & Chromatin (2023)
-
Genome folding through loop extrusion by SMC complexes
Nature Reviews Molecular Cell Biology (2021)
-
The condensin holocomplex cycles dynamically between open and collapsed states
Nature Structural & Molecular Biology (2020)
-
Tetratricopeptide repeat domain 7A is a nuclear factor that modulates transcription and chromatin structure
Cell Discovery (2018)
-
Structure of the cohesin loader Scc2
Nature Communications (2017)