Abstract
RNA-binding proteins (RBPs) regulate numerous aspects of gene expression; thus, identification of their endogenous targets is important for understanding their cellular functions. Here we identified transcriptome-wide targets of Rbfox3 in neuronally differentiated P19 cells and mouse brain by using photoactivatable ribonucleoside–enhanced cross-linking and immunoprecipitation (PAR-CLIP). Although Rbfox3 is known to regulate pre-mRNA splicing through binding the UGCAUG motif, PAR-CLIP analysis revealed diverse Rbfox3 targets including primary microRNAs (pri-miRNAs) that lack the UGCAUG motif. Induced expression and depletion of Rbfox3 led to changes in the expression levels of a subset of PAR-CLIP-detected miRNAs. In vitro analyses revealed that Rbfox3 functions as a positive and a negative regulator at the stage of pri-miRNA processing to precursor miRNA (pre-miRNA). Rbfox3 binds directly to pri-miRNAs and regulates the recruitment of the microprocessor complex to pri-miRNAs. Our study proposes a new function for Rbfox3 in miRNA biogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Jin, Y. et al. A vertebrate RNA-binding protein Fox-1 regulates tissue-specific splicing via the pentanucleotide GCAUG. EMBO J. 22, 905–912 (2003).
Ponthier, J.L. et al. Fox-2 splicing factor binds to a conserved intron motif to promote inclusion of protein 4.1R alternative exon 16. J. Biol. Chem. 281, 12468–12474 (2006).
Huh, G.S. & Hynes, R.O. Regulation of alternative pre-mRNA splicing by a novel repeated hexanucleotide element. Genes Dev. 8, 1561–1574 (1994).
Kawamoto, S. Neuron-specific alternative splicing of nonmuscle myosin II heavy chain-B pre-mRNA requires a cis-acting intron sequence. J. Biol. Chem. 271, 17613–17616 (1996).
Modafferi, E.F. & Black, D.L. A complex intronic splicing enhancer from the c-src pre-mRNA activates inclusion of a heterologous exon. Mol. Cell. Biol. 17, 6537–6545 (1997).
Brudno, M. et al. Computational analysis of candidate intron regulatory elements for tissue-specific alternative pre-mRNA splicing. Nucleic Acids Res. 29, 2338–2348 (2001).
Nakahata, S. & Kawamoto, S. Tissue-dependent isoforms of mammalian Fox-1 homologs are associated with tissue-specific splicing activities. Nucleic Acids Res. 33, 2078–2089 (2005).
Underwood, J.G., Boutz, P.L., Dougherty, J.D., Stoilov, P. & Black, D.L. Homologues of the Caenorhabditis elegans Fox-1 protein are neuronal splicing regulators in mammals. Mol. Cell. Biol. 25, 10005–10016 (2005).
Baraniak, A.P., Chen, J.R. & Garcia-Blanco, M.A. Fox-2 mediates epithelial cell-specific fibroblast growth factor receptor 2 exon choice. Mol. Cell. Biol. 26, 1209–1222 (2006).
Kuroyanagi, H., Ohno, G., Mitani, S. & Hagiwara, M. The Fox-1 family and SUP-12 coordinately regulate tissue-specific alternative splicing in vivo. Mol. Cell. Biol. 27, 8612–8621 (2007).
Zhou, H.L., Baraniak, A.P. & Lou, H. Role for Fox-1/Fox-2 in mediating the neuronal pathway of calcitonin/calcitonin gene-related peptide alternative RNA processing. Mol. Cell. Biol. 27, 830–841 (2007).
Zhang, C. et al. Defining the regulatory network of the tissue-specific splicing factors Fox-1 and Fox-2. Genes Dev. 22, 2550–2563 (2008).
Yeo, G.W. et al. An RNA code for the FOX2 splicing regulator revealed by mapping RNA-protein interactions in stem cells. Nat. Struct. Mol. Biol. 16, 130–137 (2009).
Kim, K.K., Kim, Y.C., Adelstein, R.S. & Kawamoto, S. Fox-3 and PSF interact to activate neural cell-specific alternative splicing. Nucleic Acids Res. 39, 3064–3078 (2011).
Kim, K.K., Adelstein, R.S. & Kawamoto, S. Identification of neuronal nuclei (NeuN) as Fox-3, a new member of the Fox-1 gene family of splicing factors. J. Biol. Chem. 284, 31052–31061 (2009).
Lee, J.A., Tang, Z.Z. & Black, D.L. An inducible change in Fox-1/A2BP1 splicing modulates the alternative splicing of downstream neuronal target exons. Genes Dev. 23, 2284–2293 (2009).
Gallagher, T.L. et al. Rbfox-regulated alternative splicing is critical for zebrafish cardiac and skeletal muscle functions. Dev. Biol. 359, 251–261 (2011).
Gehman, L.T. et al. The splicing regulator Rbfox1 (A2BP1) controls neuronal excitation in the mammalian brain. Nat. Genet. 43, 706–711 (2011).
Fogel, B.L. et al. RBFOX1 regulates both splicing and transcriptional networks in human neuronal development. Hum. Mol. Genet. 21, 4171–4186 (2012).
Gehman, L.T. et al. The splicing regulator Rbfox2 is required for both cerebellar development and mature motor function. Genes Dev. 26, 445–460 (2012).
Kim, K.K., Nam, J., Mukouyama, Y.S. & Kawamoto, S. Rbfox3-regulated alternative splicing of Numb promotes neuronal differentiation during development. J. Cell Biol. 200, 443–458 (2013).
Pistoni, M. et al. Rbfox1 downregulation and altered calpain 3 splicing by FRG1 in a mouse model of Facioscapulohumeral muscular dystrophy (FSHD). PLoS Genet. 9, e1003186 (2013).
Licatalosi, D.D. et al. HITS-CLIP yields genome-wide insights into brain alternative RNA processing. Nature 456, 464–469 (2008).
Xue, Y. et al. Genome-wide analysis of PTB-RNA interactions reveals a strategy used by the general splicing repressor to modulate exon inclusion or skipping. Mol. Cell 36, 996–1006 (2009).
Yeo, G.W. et al. An RNA code for the FOX2 splicing regulator revealed by mapping RNA-protein interactions in stem cells. Nat. Struct. Mol. Biol. 16, 130–137 (2009).
Macias, S. et al. DGCR8 HITS-CLIP reveals novel functions for the Microprocessor. Nat. Struct. Mol. Biol. 19, 760–766 (2012).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Jungkamp, A.C. et al. In vivo and transcriptome-wide identification of RNA binding protein target sites. Mol. Cell 44, 828–840 (2011).
Kim, V.N., Han, J. & Siomi, M.C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 10, 126–139 (2009).
Winter, J., Jung, S., Keller, S., Gregory, R.I. & Diederichs, S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat. Cell Biol. 11, 228–234 (2009).
Chang, T.C. & Mendell, J.T. microRNAs in vertebrate physiology and human disease. Annu. Rev. Genomics Hum. Genet. 8, 215–239 (2007).
Sayed, D. & Abdellatif, M. MicroRNAs in development and disease. Physiol. Rev. 91, 827–887 (2011).
Fukuda, T. et al. DEAD-box RNA helicase subunits of the Drosha complex are required for processing of rRNA and a subset of microRNAs. Nat. Cell Biol. 9, 604–611 (2007).
Guil, S. & Caceres, J.F. The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nat. Struct. Mol. Biol. 14, 591–596 (2007).
Trabucchi, M. et al. The RNA-binding protein KSRP promotes the biogenesis of a subset of microRNAs. Nature 459, 1010–1014 (2009).
Davis, B.N., Hilyard, A.C., Nguyen, P.H., Lagna, G. & Hata, A. Smad proteins bind a conserved RNA sequence to promote microRNA maturation by Drosha. Mol. Cell 39, 373–384 (2010).
Michlewski, G. & Caceres, J.F. Antagonistic role of hnRNP A1 and KSRP in the regulation of let-7a biogenesis. Nat. Struct. Mol. Biol. 17, 1011–1018 (2010).
Wu, H. et al. A splicing-independent function of SF2/ASF in microRNA processing. Mol. Cell 38, 67–77 (2010).
Rau, F. et al. Misregulation of miR-1 processing is associated with heart defects in myotonic dystrophy. Nat. Struct. Mol. Biol. 18, 840–845 (2011).
Kawai, S. & Amano, A. BRCA1 regulates microRNA biogenesis via the DROSHA microprocessor complex. J. Cell Biol. 197, 201–208 (2012).
Morlando, M. et al. FUS stimulates microRNA biogenesis by facilitating co-transcriptional Drosha recruitment. EMBO J. 31, 4502–4510 (2012).
Wang, Y., Vogel, G., Yu, Z. & Richard, S. The QKI-5 and QKI-6 RNA binding proteins regulate the expression of microRNA 7 in glial cells. Mol. Cell. Biol. 33, 1233–1243 (2013).
Maroney, P.A., Chamnongpol, S., Souret, F. & Nilsen, T.W. A rapid, quantitative assay for direct detection of microRNAs and other small RNAs using splinted ligation. RNA 13, 930–936 (2007).
Gregory, R.I. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004).
Han, J. et al. The Drosha-DGCR8 complex in primary microRNA processing. Genes Dev. 18, 3016–3027 (2004).
Auweter, S.D. et al. Molecular basis of RNA recognition by the human alternative splicing factor Fox-1. EMBO J. 25, 163–173 (2006).
Jangi, M., Boutz, P.L., Paul, P. & Sharp, P.A. Rbfox2 controls autoregulation in RNA-binding protein networks. Genes Dev. 28, 637–651 (2014).
Weyn-Vanhentenryck, S.M. et al. HITS-CLIP and integrative modeling define the Rbfox splicing-regulatory network linked to brain development and autism. Cell Reports 6, 1139–1152 (2014).
Cléry, A., Blatter, M. & Allain, F.H. RNA recognition motifs: boring? Not quite. Curr. Opin. Struct. Biol. 18, 290–298 (2008).
Michlewski, G., Guil, S., Semple, C.A. & Caceres, J.F. Posttranscriptional regulation of miRNAs harboring conserved terminal loops. Mol. Cell 32, 383–393 (2008).
Auyeung, V.C., Ulitsky, I., McGeary, S.E. & Bartel, D.P. Beyond secondary structure: primary-sequence determinants license pri-miRNA hairpins for processing. Cell 152, 844–858 (2013).
Caputi, M., Mayeda, A., Krainer, A.R. & Zahler, A.M. hnRNP A/B proteins are required for inhibition of HIV-1 pre-mRNA splicing. EMBO J. 18, 4060–4067 (1999).
Corcoran, D.L. et al. PARalyzer: definition of RNA binding sites from PAR-CLIP short-read sequence data. Genome Biol. 12, R79 (2011).
Hulsen, T., de Vlieg, J. & Alkema, W. BioVenn: a web application for the comparison and visualization of biological lists using area-proportional Venn diagrams. BMC Genomics 9, 488 (2008).
Andrus, A. & Kuimelis, R.G. Base composition analysis of nucleosides using HPLC. Curr. Protoc. Nucleic Acid Chem. 1, 10.6 (2001).
Acknowledgements
We thank U. Ohler, D.L. Corcoran and N. Mukherjee (Duke University) for help in PARalyzer analysis, H. Wu (Lawrence Berkeley National Laboratory) for advice on preparation of a sequence library for PAR-CLIP, D.-Y. Lee (Biochemistry Core Facility, US National Heart, Lung, and Blood Institute (NHLBI)) for HPLC analysis of ribonucleosides and C.A. Combs (Light Microscope Core Facility, NHLBI) for advice on confocal microscopy. We also thank D.E. Saunders and A.F. Smith for technical assistance, S. Nakahata for reagents and M.A. Conti for critical reading of the manuscript. This work was supported by the Division of Intramural Research, NHLBI, US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
K.K.K. and S.K. designed the study. K.K.K. performed the biological experiments, and K.K.K., R.S.A. and S.K. interpreted the data. Y.Y. and J.Z. performed bioinformatics analyses. K.K.K., R.S.A. and S.K. wrote the manuscript with input from Y.Y. and J.Z.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 In vivo PAR-CLIP for Rbfox3 in the mouse CNS.
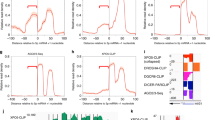
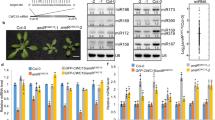
(a) Schematic outline of the strategy for in vivo mouse PAR-CLIP. Intraperitoneal injection of 4SU into adult mice and irradiation of dissociated cells from tissues with 365 nm UV followed by common PAR-CLIP protocol are illustrated. (b) Autoradiogram of radioactively phosphorylated RNA–protein complexes following LDS-PAGE. Samples are T4 polynucleotide kinase-phosphorylated anti-Rbfox3- or IgG-immunoprecipitates prepared from UV-irradiated brain cells of 4SU injected mice. The area boxed with broken red lines was processed for further analysis. IP, immunoprecipitation. (c) Immunoblot detecting Rbfox3. Samples are immunoprecipitates with mouse anti-Rbfox3 (m) or IgG from mouse brain extracts. Rabbit anti-Rbfox3 (r) was used for immunoblotting. Input contains 5% extract. Uncropped image is provided in Supplementary Figure 5. (d) Genomic distribution of Rbfox3 binding clusters. Numbers in parenthesis indicate cluster numbers. A full list of PAR-CLIP clusters is provided in Supplementary Data Set 2.
Supplementary Figure 2 Effect of Rbfox3 on pri-miRNA levels in Drosha-knockdown P19 cells.
Transfected siRNA or expression plasmid is indicated. siDrosha, siRNA against Drosha; control, non-targeting siRNA and the empty vector. (a) Quantification of pri-miRNAs by qRT-PCR. n = 3 (biological replicates, cell cultures), average ± s.e.m., * P < 0.001 (ANOVA, Bonferroni’s multiple comparisons test). ns, no statistically significant difference. (b) RT-PCR analysis of pri-miRNAs. PCR products with (+) or without (−) RT following agarose gel electrophoresis is shown. (c) Immunoblot analysis of Drosha and myc-Rbfox3. Gapdh serves as a loading control. Uncropped images are provided in Supplementary Figure 5.
Supplementary Figure 3 Nucleotide sequences of the stem-loop region of wild-type and mutant pri-miRNAs.
Mature miR-5p and 3p are indicated in red. Mutant nucleotides are indicated in cyan and dots indicate deletion. The underlined region corresponds to the Rbfox3-binding cluster detected by PAR-CLIP. Wt, wild-type; Mt, mutant.
Supplementary Figure 4 Determination of 4SU residues on pri-miRNAs cross-linked with Rbfox3 by in vitro PAR-CLIP.
The “u” residues in red were converted to “c”, indicating the crosslinked residues. (a,b) The nucleotides with underlines were the cluster sequences determined in RA-treated P19 cells. (c) The underlined sequences are the canonical Rbfox binding sequence, UGCAUG, in the IDDE.
Supplementary Figure 5 Uncropped images of immunoblots and autoradiograms.
The areas boxed with broken red lines are used for the figures.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 3670 kb)
Supplementary Data Set 1
Rbfox3-binding clusters in neuronally differentiated P19 cells determined by PAR-CLIP (XLSX 418 kb)
Supplementary Data Set 2
Rbfox3-binding clusters in the mouse brain and spinal cord determined by PAR-CLIP (XLSX 97 kb)
Supplementary Data Set 3
Comparative miRNA microarray analysis of untreated P19-GFP, RA-treated P19-GFP and RA-treated P19-T2 cells (XLSX 24 kb)
Supplementary Data Set 4
PAR-CLIP read numbers at each processing step from P19 cells and mouse brain (XLSX 9 kb)
Rights and permissions
About this article
Cite this article
Kim, K., Yang, Y., Zhu, J. et al. Rbfox3 controls the biogenesis of a subset of microRNAs. Nat Struct Mol Biol 21, 901–910 (2014). https://doi.org/10.1038/nsmb.2892
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2892
This article is cited by
-
Functional identification of microRNA-centered complexes in C. elegans
Scientific Reports (2022)
-
Regulation of microRNA biogenesis and its crosstalk with other cellular pathways
Nature Reviews Molecular Cell Biology (2019)
-
CLIP-related methodologies and their application to retrovirology
Retrovirology (2018)
-
Neuron loss and degeneration in the progression of TDP-43 in frontotemporal lobar degeneration
Acta Neuropathologica Communications (2017)
-
Stress Granules Contain Rbfox2 with Cell Cycle-related mRNAs
Scientific Reports (2017)