Abstract
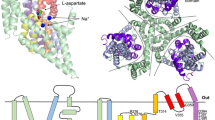
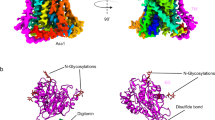
GltPh is a Pyrococcus horikoshii homotrimeric Na+-coupled aspartate transporter that belongs to the glutamate transporter family. Each protomer consists of a trimerization domain involved in subunit interaction and a transporting domain with the substrate-binding site. Here, we have studied the conformational changes underlying transport by GltPh using EPR spectroscopy. The trimerization domains form a rigid scaffold, whereas the transporting domains sample multiple conformations, consistent with large-scale movements during the transport cycle. Binding of substrates changed the occupancies of the different conformational states, but the domains remained heterogeneous. The membrane environment favored conformations different from those observed in detergent micelles, but the transporting domain remained structurally heterogeneous in both environments. We conclude that the transporting domains sample multiple conformational states with substantial occupancy regardless of the presence of substrate and coupling ions, consistent with equilibrium constants close to unity between the observed transporter conformations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Tzingounis, A.V. & Wadiche, J.I. Glutamate transporters: confining runaway excitation by shaping synaptic transmission. Nat. Rev. Neurosci. 8, 935–947 (2007).
Slotboom, D.J., Konings, W.N. & Lolkema, J.S. Structural features of the glutamate transporter family. Microbiol. Mol. Biol. Rev. 63, 293–307 (1999).
Teichman, S., Qu, S. & Kanner, B.I. The equivalent of a thallium binding residue from an archeal homolog controls cation interactions in brain glutamate transporters. Proc. Natl. Acad. Sci. USA 106, 14297–14302 (2009).
Groeneveld, M. & Slotboom, D.J. Na+:aspartate coupling stoichiometry in the glutamate transporter homologue Glt(Ph). Biochemistry 49, 3511–3513 (2010).
Zerangue, N. & Kavanaugh, M.P. Flux coupling in a neuronal glutamate transporter. Nature 383, 634–637 (1996).
Raunser, S. et al. Structure and function of prokaryotic glutamate transporters from Escherichia coli and Pyrococcus horikoshii. Biochemistry 45, 12796–12805 (2006).
Ryan, R.M., Compton, E.L. & Mindell, J.A. Functional characterization of a Na+-dependent aspartate transporter from Pyrococcus horikoshii. J. Biol. Chem. 284, 17540–17548 (2009).
Ryan, R.M. & Mindell, J.A. The uncoupled chloride conductance of a bacterial glutamate transporter homolog. Nat. Struct. Mol. Biol. 14, 365–371 (2007).
Verdon, G. & Boudker, O. Crystal structure of an asymmetric trimer of a bacterial glutamate transporter homolog. Nat. Struct. Mol. Biol. 19, 355–357 (2012).
Boudker, O., Ryan, R.M., Yernool, D., Shimamoto, K. & Gouaux, E. Coupling substrate and ion binding to extracellular gate of a sodium-dependent aspartate transporter. Nature 445, 387–393 (2007).
Yernool, D., Boudker, O., Jin, Y. & Gouaux, E. Structure of a glutamate transporter homologue from Pyrococcus horikoshii. Nature 431, 811–818 (2004).
Reyes, N., Ginter, C. & Boudker, O. Transport mechanism of a bacterial homologue of glutamate transporters. Nature 462, 880–885 (2009).
Groeneveld, M. & Slotboom, D.J. Rigidity of the subunit interfaces of the trimeric glutamate transporter GltT during translocation. J. Mol. Biol. 372, 565–570 (2007).
Steinhoff, H.J. Inter- and intra-molecular distances determined by EPR spectroscopy and site-directed spin labeling reveal protein-protein and protein-oligonucleotide interaction. Biol. Chem. 385, 913–920 (2004).
Polyhach, Y., Bordignon, E. & Jeschke, G. Rotamer libraries of spin labelled cysteines for protein studies. Phys. Chem. Chem. Phys. 13, 2356–2366 (2011).
Jeschke, G., Sajid, M., Schulte, M. & Godt, A. Three-spin correlations in double electron-electron resonance. Phys. Chem. Chem. Phys. 11, 6580–6591 (2009).
Boudker, O. & Verdon, G. Structural perspectives on secondary active transporters. Trends Pharmacol. Sci. 31, 418–426 (2010).
Focke, P.J., Moenne-Loccoz, P. & Larsson, H.P. Opposite movement of the external gate of a glutamate transporter homolog upon binding cotransported sodium compared with substrate. J. Neurosci. 31, 6255–6262 (2011).
Jiang, J., Shrivastava, I.H., Watts, S.D., Bahar, I. & Amara, S.G. Large collective motions regulate the functional properties of glutamate transporter trimers. Proc. Natl. Acad. Sci. USA 108, 15141–15146 (2011).
Radzwill, N., Gerwert, K. & Steinhoff, H.J. Time-resolved detection of transient movement of helices F and G in doubly spin-labeled bacteriorhodopsin. Biophys. J. 80, 2856–2866 (2001).
Pannier, M., Veit, S., Godt, A., Jeschke, G. & Spiess, H.W. Dead-time free measurement of dipole-dipole interactions between electron spins. J. Magn. Reson. 142, 331–340 (2000).
Hänelt, I. et al. Membrane region M2C2 in subunit KtrB of the K+ uptake system KtrAB from Vibrio alginolyticus forms a flexible gate controlling K+ flux: an electron paramagnetic resonance study. J. Biol. Chem. 285, 28210–28219 (2010).
Polyhach, Y. et al. High sensitivity and versatility of the DEER experiment on nitroxide radical pairs at Q-band frequencies. Phys. Chem. Chem. Phys. 14, 10762–10773 (2012).
Steinhoff, H.J. et al. Determination of interspin distances between spin labels attached to insulin: comparison of electron paramagnetic resonance data with the X-ray structure. Biophys. J. 73, 3287–3298 (1997).
Jeschke, G. et al. DeerAnalysis2006—a comprehensive software package for analyzing pulsed ELDOR data. Appl. Magn. Reson. 30, 473–498 (2006).
Mackerell, A.D. Jr., Feig, M. & Brooks, C.L. III. Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004).
Acknowledgements
We thank R.H. Duurkens for performing uptake experiments, C. Rickert, D. Klose and J. Klare for help with the EPR measurements and B. Poolman for constructive criticism. This work was supported by a research fellowship and by a research grant from the Deutsche Forschungsgemeinschaft (HA 6322/1-1 to I.H. and STE 640/10, SFB944 to D.W. and H.-J.S.), the Netherlands Organisation for Scientific Research (NWO Vidi and Vici grant to D.J.S.) and the European Union (EDICT program and European Research Council starting grant to D.J.S.).
Author information
Authors and Affiliations
Contributions
I.H. and D.J.S. designed the experiments. I.H., D.W. and E.B. conducted the experiments. All authors contributed to writing the manuscript and analyzing the data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figs 1–4 and Supplementary Table 1 (PDF 6524 kb)
Rights and permissions
About this article
Cite this article
Hänelt, I., Wunnicke, D., Bordignon, E. et al. Conformational heterogeneity of the aspartate transporter GltPh. Nat Struct Mol Biol 20, 210–214 (2013). https://doi.org/10.1038/nsmb.2471
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2471
This article is cited by
-
Orientational Selectivity in Pulsed-EPR Does Not Have to be Complicated
Applied Magnetic Resonance (2024)
-
Kinetic mechanism of Na+-coupled aspartate transport catalyzed by GltTk
Communications Biology (2021)
-
Millisecond dynamics of an unlabeled amino acid transporter
Nature Communications (2020)
-
Structural basis for functional interactions in dimers of SLC26 transporters
Nature Communications (2019)
-
Probing the solution structure of the E. coli multidrug transporter MdfA using DEER distance measurements with nitroxide and Gd(III) spin labels
Scientific Reports (2019)