Abstract
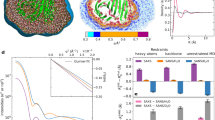
The small heat shock protein αB-crystallin (αB) contributes to cellular protection against stress. For decades, high-resolution structural studies on oligomeric αB have been confounded by its polydisperse nature. Here, we present a structural basis of oligomer assembly and activation of the chaperone using solid-state NMR and small-angle X-ray scattering (SAXS). The basic building block is a curved dimer, with an angle of ∼121° between the planes of the β-sandwich formed by α-crystallin domains. The highly conserved IXI motif covers a substrate binding site at pH 7.5. We observe a pH-dependent modulation of the interaction of the IXI motif with β4 and β8, consistent with a pH-dependent regulation of the chaperone function. N-terminal region residues Ser59-Trp60-Phe61 are involved in intermolecular interaction with β3. Intermolecular restraints from NMR and volumetric restraints from SAXS were combined to calculate a model of a 24-subunit αB oligomer with tetrahedral symmetry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bukau, B., Weissman, J. & Horwich, A. Molecular chaperones and protein quality control. Cell 125, 443–451 (2006).
Haslbeck, M., Franzmann, T., Weinfurtner, D. & Buchner, J. Some like it hot: the structure and function of small heat-shock proteins. Nat. Struct. Mol. Biol. 12, 842–846 (2005).
Ecroyd, H. & Carver, J.A. Crystallin proteins and amyloid fibrils. Cell. Mol. Life Sci. 66, 62–81 (2009).
Horwitz, J. α-crystallin can function as a molecular chaperone. Proc. Natl. Acad. Sci. USA 89, 10449–10453 (1992).
Lin, D.I. et al. Phosphorylation-dependent ubiquitination of cyclin D1 by the SCF(FBX4-αB crystallin) complex. Mol. Cell 24, 355–366 (2006).
Goldstein, L.E. et al. Cytosolic β-amyloid deposition and supranuclear cataracts in lenses from people with Alzheimer's disease. Lancet 361, 1258–1265 (2003).
Kato, K. et al. Ser-59 is the major phosphorylation site in αB-crystallin accumulated in the brains of patients with Alexander's disease. J. Neurochem. 76, 730–736 (2001).
Vicart, P. et al. A missense mutation in the αB-crystallin chaperone gene causes a desmin-related myopathy. Nat. Genet. 20, 92–95 (1998).
Rajasekaran, N.S. et al. Human α B-crystallin mutation causes oxido-reductive stress and protein aggregation cardiomyopathy in mice. Cell 130, 427–439 (2007).
Liu, Y. et al. A novel αB-crystallin mutation associated with autosomal dominant congenital lamellar cataract. Invest. Ophthalmol. Vis. Sci. 47, 1069–1075 (2006).
Selcen, D. & Engel, A.G. Myofibrillar myopathy caused by novel dominant negative α B-crystallin mutations. Ann. Neurol. 54, 804–810 (2003).
Ousman, S.S. et al. Protective and therapeutic role for αB-crystallin in autoimmune demyelination. Nature 448, 474–479 (2007).
van Noort, J.M. et al. The small heat-shock protein α B-crystallin as candidate autoantigen in multiple sclerosis. Nature 375, 798–801 (1995).
Bennardini, F., Wrzosek, A. & Chiesi, M. α B-crystallin in cardiac tissue. Association with actin and desmin filaments. Circ. Res. 71, 288–294 (1992).
Aquilina, J.A. et al. Polydispersity of a mammalian chaperone: mass spectrometry reveals the population of oligomers in αB-crystallin. Proc. Natl. Acad. Sci. USA 100, 10611–10616 (2003).
Pasta, S.Y., Raman, B., Ramakrishna, T. & Rao Ch, M. The IXI/V motif in the C-terminal extension of α-crystallins: alternative interactions and oligomeric assemblies. Mol. Vis. 10, 655–662 (2004).
Sreelakshmi, Y. & Sharma, K.K. Recognition sequence 2 (residues 60–71) plays a role in oligomerization and exchange dynamics of αB-crystallin. Biochemistry 44, 12245–12252 (2005).
Kim, K.K., Kim, R. & Kim, S.H. Crystal structure of a small heat-shock protein. Nature 394, 595–599 (1998).
van Montfort, R.L. et al. Crystal structure and assembly of a eukaryotic small heat shock protein. Nat. Struct. Biol. 8, 1025–1030 (2001).
Stamler, R., Kappe, G., Boelens, W. & Slingsby, C. Wrapping the α-crystallin domain fold in a chaperone assembly. J. Mol. Biol. 353, 68–79 (2005).
Bagneris, C. et al. Crystal structures of α-crystallin domain dimers of αB-crystallin and Hsp20. J. Mol. Biol. 392, 1242–1252 (2009).
Jehle, S. et al. αB-crystallin: a hybrid solid-state/solution-state NMR investigation reveals structural aspects of the heterogeneous oligomer. J. Mol. Biol. 385, 1481–1497 (2009).
Koteiche, H.A. & McHaourab, H.S. Folding pattern of the α-crystallin domain in αA-crystallin determined by site-directed spin labeling. J. Mol. Biol. 294, 561–577 (1999).
Haley, D.A., Horwitz, J. & Stewart, P.L. The small heat-shock protein, αB-crystallin, has a variable quaternary structure. J. Mol. Biol. 277, 27–35 (1998).
Peschek, J. et al. The eye lens chaperone α-crystallin forms defined globular assemblies. Proc. Natl. Acad. Sci. USA 106, 13272–13277 (2009).
Higman, V.A. et al. Assigning large proteins in the solid state: a MAS NMR resonance assignment strategy using selectively and extensively 13C-labelled proteins. J. Biomol. NMR 44, 245–260 (2009).
Castellani, F. et al. Structure of a protein determined by solid-state magic-angle-spinning NMR spectroscopy. Nature 420, 98–102 (2002).
Bardiaux, B. et al. Influence of different assignment conditions on the determination of symmetric homodimeric structures with ARIA. Proteins 75, 569–585 (2009).
Castellani, F. et al. Determination of solid-state NMR structures of proteins by means of three-dimensional 15N–13C–13C dipolar correlation spectroscopy and chemical shift analysis. Biochemistry 42, 11476–11483 (2003).
Lange, A. et al. Analysis of proton-proton transfer dynamics in rotating solids and their use for 3D structure determination. J. Am. Chem. Soc. 125, 12640–12648 (2003).
Gallivan, J.P. & Dougherty, D.A. Cation-pi interactions in structural biology. Proc. Natl. Acad. Sci. USA 96, 9459–9464 (1999).
Bhattacharyya, J. et al. Mini-αB-crystallin: a functional element of αB-crystallin with chaperone-like activity. Biochemistry 45, 3069–3076 (2006).
Augusteyn, R.C. α-crystallin: a review of its structure and function. Clin. Exp. Optom. 87, 356–366 (2004).
Pettersen, E.F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Goddard, T.D., Huang, C.C. & Ferrin, T.E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Schwieters, C.D., Kuszewski, J.J., Tjandra, N. & Clore, G.M. The Xplor-NIH NMR molecular structure determination package. J. Magn. Reson. 160, 65–73 (2003).
Kozin, M.B. et al. Automated matching of high- and low-resolution structural models. J. Appl. Crystallogr. 34, 33–41 (2001).
Svergun, D.I., Petoukhov, M.V. & Koch, M.H. Determination of domain structure of proteins from X-ray solution scattering. Biophys. J. 80, 2946–2953 (2001).
Carver, J.A., Aquilina, J.A., Truscott, R.J. & Ralston, G.B. Identification by 1H NMR spectroscopy of flexible C-terminal extensions in bovine lens α-crystallin. FEBS Lett. 311, 143–149 (1992).
Kundu, B., Shukla, A., Chaba, R. & Guptasarma, P. The excised heat-shock domain of αB crystallin is a folded, proteolytically susceptible trimer with significant surface hydrophobicity and a tendency to self-aggregate upon heating. Protein Expr. Purif. 36, 263–271 (2004).
Koteiche, H.A. & McHaourab, H.S. Mechanism of chaperone function in small heat-shock proteins. Phosphorylation-induced activation of two-mode binding in αB-crystallin. J. Biol. Chem. 278, 10361–10367 (2003).
Wriggers, W. & Chacón, P. Using Situs for the registration of protein structures with low-resolution bead models from X-ray solution scattering. J. Appl. Crystallogr. 34, 773–776 (2001).
Wasmer, C. et al. Amyloid fibrils of the HET-s(218–289) prion form a β solenoid with a triangular hydrophobic core. Science 319, 1523–1526 (2008).
Lewandowski, J.R., De Paepe, G. & Griffin, R.G. Proton assisted insensitive nuclei cross polarization. J. Am. Chem. Soc. 129, 728–729 (2007).
Jaroniec, C.P., Filip, C. & Griffin, R.G. 3D TEDOR NMR experiments for the simultaneous measurement of multiple carbon-nitrogen distances in uniformly (13)C,(15)N-labeled solids. J. Am. Chem. Soc. 124, 10728–10742 (2002).
Fossi, M. et al. SOLARIA: a protocol for automated cross-peak assignment and structure calculation for solid-state magic-angle spinning NMR spectroscopy. Angew. Chem. Int. Edn Engl. 44, 6151–6154 (2005).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Brunger, A.T. Version 1.2 of the Crystallography and NMR system. Nat. Protoc. 2, 2728–2733 (2007).
Linge, J.P. et al. Refinement of protein structures in explicit solvent. Proteins 50, 496–506 (2003).
Laskowski, R.A. et al. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Vriend, G. WHAT IF: a molecular modeling and drug design program. J. Mol. Graph. 8, 52–56 (1990).
Davis, I.W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Putnam, C.D., Hammel, M., Hura, G.L. & Tainer, J.A. X-ray solution scattering (SAXS) combined with crystallography and computation: defining accurate macromolecular structures, conformations and assemblies in solution. Q. Rev. Biophys. 40, 191–285 (2007).
Hura, G.L. et al. Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS). Nat. Methods 6, 606–612 (2009).
Petoukhov, M.V. & Svergun, D.I. Analysis of X-ray and neutron scattering from biomacromolecular solutions. Curr. Opin. Struct. Biol. 17, 562–571 (2007).
Nilges, M. A calculation strategy for the structure determination of symmetric dimers by 1H NMR. Proteins 17, 297–309 (1993).
Acknowledgements
The authors thank K. Rehbein and A. Diehl for sample preparation and discussions and D. Svergun for fruitful discussions concerning the SAXS experiments. This work was funded by US National Institutes of Health grant 1R01EY017370. SAXS data on αB10.1 was collected at the Stanford Synchrotron Radiation Lightsource, a national user facility operated by Stanford University on behalf of the US Department of Energy, Office of Basic Energy Sciences. The Stanford Synchrotron Radiation Lightsource Structural Molecular Biology Program is supported by the US Department of Energy, Office of Biological and Environmental Research, and by the US National Institutes of Health, National Center for Research Resources, Biomedical Technology Program. SAXS data collection for full-length αB was supported by European Community–European Molecular Biology Laboratory Hamburg Outstation; Deutsches Elektronen-Synchrotron Hamburg X33 beamline.
Author information
Authors and Affiliations
Contributions
S.J. contributed to all aspects of the manuscript; P.R. performed solution NMR experiments and helped to write the manuscript; B.B. performed structure calculations; S.M. did solid-state NMR and SAXS measurements as well as data analysis; R.K. contributed to modeling of C-terminal intermolecular interactions; J.R.S. prepared samples; V.A.H. contributed to assignment strategies, was involved in structure calculations and helped write the manuscript; R.E.K. contributed to the interpretation of results and wrote the manuscript; B.J.v.R. contributed to solid-state NMR measurements, discussed the results and helped to write the manuscript; H.O. designed experimental strategies, contributed to the interpretation of results and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1 and 2 (PDF 591 kb)
Rights and permissions
About this article
Cite this article
Jehle, S., Rajagopal, P., Bardiaux, B. et al. Solid-state NMR and SAXS studies provide a structural basis for the activation of αB-crystallin oligomers. Nat Struct Mol Biol 17, 1037–1042 (2010). https://doi.org/10.1038/nsmb.1891
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1891
This article is cited by
-
Structural basis of substrate recognition and thermal protection by a small heat shock protein
Nature Communications (2021)
-
The chaperone HSPB1 prepares protein aggregates for resolubilization by HSP70
Scientific Reports (2021)
-
O-GlcNAc modification of small heat shock proteins enhances their anti-amyloid chaperone activity
Nature Chemistry (2021)
-
Trends in HSPB5 research: a 36-year bibliometric analysis
Cell Stress and Chaperones (2021)
-
The multifaceted nature of αB-crystallin
Cell Stress and Chaperones (2020)


