Abstract
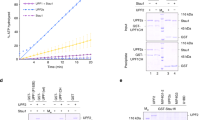
The AU-rich element (ARE)-mediated mRNA-degradation activity of the RNA binding K-homology splicing regulator protein (KSRP) is regulated by phosphorylation of a serine within its N-terminal KH domain (KH1). In the cell, phosphorylation promotes the interaction of KSRP and 14-3-3ζ protein and impairs the ability of KSRP to promote the degradation of its RNA targets. Here we examine the molecular details of this mechanism. We report that phosphorylation leads to the unfolding of the structurally atypical and unstable KH1, creating a site for 14-3-3ζ binding. Using this site, 14-3-3ζ discriminates between phosphorylated and unphosphorylated KH1, driving the nuclear localization of KSRP. 14-3-3ζ –KH1 interaction regulates the mRNA-decay activity of KSRP by sequestering the protein in a separate functional pool. This study demonstrates how an mRNA-degradation pathway is connected to extracellular signaling networks through the reversible unfolding of a protein domain.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Audic, Y. & Hartley, R.S. Post-transcriptional regulation in cancer. Biol. Cell 96, 479–498 (2004).
Kontoyiannis, D., Pasparakis, M., Pizarro, T.T., Cominelli, F. & Kollias, G. Impaired on/off regulation of TNF biosynthesis in mice lacking TNF AU-rich elements: implications for joint and gut-associated immunopathologies. Immunity 10, 387–398 (1999).
Barreau, C., Paillard, L. & Osborne, H.B. AU-rich elements and associated factors: are there unifying principles? Nucleic Acids Res. 33, 7138–7150 (2006).
Pullmann, R. et al. Analysis of turnover and translation regulatory RNA-binding protein expression through binding to cognate mRNAs. Mol. Cell. Biol. 27, 6265–6278 (2007).
Chen, C.-Y. et al. AU binding proteins recruit the exosome to degrade ARE-containing mRNAs. Cell 107, 451–464 (2001).
Gherzi, R. et al. A KH domain RNA binding protein, KSRP, promotes ARE-directed mRNA turnover by recruiting the degradation machinery. Mol. Cell 14, 571–583 (2004).
García-Mayoral, M.F. et al. The structure of the C-terminal KH domains of KSRP reveals a non canonical motif important for mRNA degradation. Structure 15, 485–498 (2007).
Briata, P. et al. p38-dependent phosphorylation of the mRNA decay-promoting factor KSRP controls the stability of select myogenic transcripts. Mol. Cell 20, 891–903 (2005).
Gherzi, R. et al. The RNA-binding protein KSRP promotes decay of β-catenin mRNA and is inactivated by PI3K-AKT signaling. PLoS Biol. 5, e5 (2006).
Ruggiero, T. et al. Identification of a set of KSRP target transcripts up-regulated by PI3K-AKT signaling. BMC Mol. Biol. 8, 28–43 (2007).
Yaffe, M.B. et al. The structural basis for 14–3-3:phospho-peptide binding specificity. Cell 91, 961–971 (1997).
Mackintosh, C. Dynamic interactions between 14–3-3 proteins and phospho-proteins regulate diverse cellular processes. Biochem. J. 381, 329–342 (2004).
Aitken, A. 14–3-3 proteins: a historic overview. Semin. Cancer Biol. 16, 162–172 (2006).
Gardino, A.K., Smerdon, S.J. & Yaffe, M.B. Structural determinants of 14–3-3 binding specificities and regulation of sub cellular localization of 14–3-3-ligand complexes: a comparison of the X-ray crystal structures of all human 14–3-3 isoforms. Semin. Cancer Biol. 16, 173–182 (2006).
Stoecklin, G. et al. MK2-induced tristetraprolin:14–3-3 complexes prevent stress granule association and ARE-mRNA decay. EMBO J. 23, 1313–1324 (2004).
Johnson, B.A., Stehn, J.R., Yaffe, M.B. & Blackwell, T.K. Cytoplasmic localization of tristetraprolin involves 14–3-3-dependent and -independent mechanisms. J. Biol. Chem. 277, 18029–18036 (2002).
Musco, G. et al. The solution structure of the first KH domain of FMR1, the protein responsible for the fragile X syndrome. Nat. Struct. Biol. 4, 712–716 (1997).
Baber, J.L., Levens, D., Libutti, D. & Tjandra, N. Chemical shift mapped DNA-binding sites and 15N relaxation analysis of the C-terminal KH domain of heterogeneous nuclear ribonucleoprotein K. Biochemistry 39, 6022–6032 (2000).
Ramos, A. et al. Role of dimerization in KH/RNA complexes: the example of Nova KH3. Biochemistry 41, 4193–4201 (2002).
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 233, 123–138 (1993).
Camproux, A.C., Gautier, R. & Tufféry, P. A hidden Markov model derived structural alphabet for proteins. J. Mol. Biol. 339, 591–605 (2004).
Pandini, A., Bonati, L., Fraternali, F. & Kleinjung, J. MinSet: a general approach to derive maximally representative database subsets by using fragment dictionaries and its application to the SCOP database. Bioinformatics 23, 515–516 (2007).
Beuth, B., Pennell, S., Arnvig, K.B., Martin, S.R. & Taylor, I.A. Structure of a Mycobacterium tuberculosis NusA-RNA complex. EMBO J. 24, 3576–3587 (2005).
Git, A. & Standart, N. The KH domains of Xenopus Vg1RBP mediate RNA binding and self-association. RNA 8, 1319–1333 (2002).
Valverde, R., Pozdnyakova, I., Kajander, T., Venkatraman, J. & Regan, L. Fragile X mental retardation syndrome: structure of the KH1–KH2 domains of fragile X mental retardation protein. Structure 15, 1090–1098 (2007).
Bernadó, P. et al. Interpretation of NMR relaxation properties of Pin1, a two-domain protein, based on Brownian dynamic simulations. J. Biomol. NMR 29, 21–35 (2004).
Lewis, H.A. et al. Sequence-specific RNA binding by a Nova KH domain: implications for paraneoplastic disease and the fragile X syndrome. Cell 100, 323–332 (2000).
Schanda, P. & Brutscher, B. Very fast two-dimensional NMR spectroscopy for real-time investigation of dynamic events in proteins on the time scale of seconds. J. Am. Chem. Soc. 127, 8014–8015 (2005).
Sreerama, N., Venyaminov, S.Y. & Woody, R.W. Estimation of protein secondary structure from circular dichroism spectra: inclusion of denatured proteins with native proteins in the analysis. Anal. Biochem. 287, 243–251 (2000).
Masters, S.C. & Fu, H. 14–3-3 proteins mediate an essential anti-apoptotic signal. J. Biol. Chem. 276, 45193–45200 (2001).
Johnson, L.N. & O'Reilly, M. Control by phosphorylation. Curr. Opin. Struct. Biol. 6, 762–769 (1996).
Messías, A.C., Harnisch, C., Ostareck-Lederer, A., Sattler, M. & Ostareck, D.H. The DICE-binding activity of KH domain 3 of hnRNP K is affected by c-src-mediated tyrosine phosphorylation. J. Mol. Biol. 361, 470–481 (2006).
García-Mayoral, M.F., Díaz-Moreno, I., Hollingworth, D. & Ramos, A. The sequence selectivity of KSRP explains its flexibility in the recognition of the RNA targets. Nucleic Acids Res. 36, 5290–5296 (2008).
Zhang, S., Xing, H. & Musli, A.J. Nuclear localization of protein kinase U-α is regulated by 14–3-3. J. Biol. Chem. 274, 24865–24872 (1999).
Faul, C., Hüttelmaier, S., Oh, J., Hachet, V., Singer, R.H. & Mundel, P. Promotion of importin α-mediated nuclear import by the phosphorylation-dependent binding of cargo protein to 14–3-3. J. Cell Biol. 169, 415–424 (2005).
Chou, C.F. et al. Tethering KSRP, a decay-promoting AU-rich element-binding protein, to mRNAs elicits mRNA decay. Mol. Cell. Biol. 26, 3695–3706 (2006).
Lin, W.J., Duffy, A. & Chen, C.-Y. Localization of AU-rich element-containing mRNA in cytoplasmic granules containing exosome subunits. J. Biol. Chem. 282, 19958–19968 (2007).
Markovtsov, V. et al. Cooperative assembly of an hnRNP complex induced by a tissue-specific homolog of poly-pyrimidine tract binding protein. Mol. Cell. Biol. 20, 7463–7479 (2000).
Blichenberg, A. et al. Identification of a cis-acting dendritic targeting element in MAP2 mRNAs. J. Neurosci. 19, 8818–8829 (1999).
Masino, L., Martin, S.R. & Bayley, P.M. Ligand binding and thermodynamic stability of a multidomain protein, calmodulin. Protein Sci. 9, 1519–1529 (2000).
Linge, J.P., O'Donoghue, S.I. & Nilges, M. Automated asignment of ambiguous nuclear overhauser effects with ARIA. Methods Enzymol. 339, 71–90 (2001).
Ye, J., Mayer, K.L., Mayer, M.R. & Stone, M.J. NMR solution structure and backbone dynamics of the CC chemokine eotaxin-3. Biochemistry 40, 7820–7831 (2001).
Cornilescu, G., Delaglio, F. & Bax, A. Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J. Biomol. NMR 13, 289–302 (1999).
Bartels, C., Xia, T.-H., Billeter, M., Güntert, P. & Wüthrich, K. The program XEASY for computer-supported NMR spectral analysis of biological macromolecules. J. Biomol. NMR 6, 1–10 (1995).
Koradi, R., Billeter, M. & Wüthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graph. 14, 51–55 (1996).
De Chiara, C. et al. The AXH domain adopts alternative folds: the solution structure of HBP1 AXH. Structure 13, 743–753 (2005).
Laskowski, R.A., Rullman, J.A., MacArthur, M.W., Kaptein, R. & Thorton, J.M. AQUA and PROCHECK-NMR: programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 8, 477–486 (1996).
Jeanmougin, F., Thompson, J.D., Gouy, M., Higgins, D.G. & Gibson, T.J. Multiple sequence alignment with Clustal X. Trends Biochem. Sci. 23, 403–405 (1998).
Acknowledgements
I.D.-M. was supported by European Molecular Biology Organization (EMBO), fellowship number 240-2005. The work of R.G. is supported by a grant from the Associazione Italiana per le Ricerca sul Cancro (AIRC) and the Istituto Superiore di Sanita' (ISS), whereas P.B. is a recipient of a Senior Scholar Consultancy Grant from American Italian Cancer Foundation (AICF). The work of R.G. and P.B. is also supported by the Italian Comitato Interministerial per le Programmazione Economica (CIPE)-2007. We would like to thank J. Kleinjung and A. Pandini for the fragment-based search SCOP40 database, C. De Chiara for advice on the ARIA protocols, A. Oreggioni for his help in recording spectra, and I. Taylor for checking the oligomerization state of protein constructs by MALLS. We would like to thank K. Rittinger (MRC National Institute for Medical Research) for the gift of the GST–14-3-3ζ expression vector and for help in recording the ITC measurements and M. Lalle (Istituto Superiore di Sanita') for the gift of the difopein expression plasmid. Finally, we would like to thank P. Fletcher (MRC National Institute for Medical Research) for the synthesis of many of the KH1 peptides. All NMR spectra were recorded at the MRC Biomedical NMR centre.
Author information
Authors and Affiliations
Contributions
Cloning and site-specific mutagenesis was performed by D.H. Protein purification was performed by I.D.-M. and D.H. NMR experiments were performed by I.D.-M., G.K., T.A.F. and A.R. NMR data analysis was performed by I.D.-M. and M.G.-M. Structure calculations were performed by I.D.-M. CD experiments and analysis of CD data were performed by A.R. and S.M. MS analysis was performed by S.H. Phosphorylation experiments were designed and carried out by D.H. ITC experiments were performed by A.R. Subcellular localization experiments were performed by P.B. and R.G. The paper was written by I.D.-M. and A.R.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Table 1 and Supplementary Methods (PDF 846 kb)
Rights and permissions
About this article
Cite this article
Díaz-Moreno, I., Hollingworth, D., Frenkiel, T. et al. Phosphorylation-mediated unfolding of a KH domain regulates KSRP localization via 14-3-3 binding. Nat Struct Mol Biol 16, 238–246 (2009). https://doi.org/10.1038/nsmb.1558
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1558
This article is cited by
-
De novo design of tyrosine and serine kinase-driven protein switches
Nature Structural & Molecular Biology (2021)
-
RNA-binding protein KHSRP promotes tumor growth and metastasis in non-small cell lung cancer
Journal of Experimental & Clinical Cancer Research (2019)
-
Phosphorylation induced cochaperone unfolding promotes kinase recruitment and client class-specific Hsp90 phosphorylation
Nature Communications (2018)
-
SUMO1 modification of KHSRP regulates tumorigenesis by preventing the TL-G-Rich miRNA biogenesis
Molecular Cancer (2017)
-
On the role of residue phosphorylation in 14-3-3 partners: AANAT as a case study
Scientific Reports (2017)