Abstract
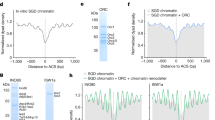
The eukaryotic DNA replication machinery must traverse every nucleosome in the genome during S phase. As nucleosomes are generally inhibitory to DNA-dependent processes, chromatin structure must undergo extensive reorganization to facilitate DNA synthesis. However, the identity of chromatin-remodeling factors involved in replication and how they affect DNA synthesis is largely unknown. Here we show that two highly conserved ATP-dependent chromatin-remodeling complexes in Saccharomyces cerevisiae, Isw2 and Ino80, function in parallel to promote replication fork progression. As a result, Isw2 and Ino80 have especially important roles for replication of late-replicating regions during periods of replication stress. Both Isw2 and Ino80 complexes are enriched at sites of replication, suggesting that these complexes act directly to promote fork progression. These findings identify ATP-dependent chromatin-remodeling complexes that promote DNA replication and define a specific stage of replication that requires remodeling for normal function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Becker, P.B. & Horz, W. ATP-dependent nucleosome remodeling. Annu. Rev. Biochem. 71, 247–273 (2002).
Smith, C.L. & Peterson, C.L. ATP-dependent chromatin remodeling. Curr. Top. Dev. Biol. 65, 115–148 (2005).
Narlikar, G.J., Fan, H.Y. & Kingston, R.E. Cooperation between complexes that regulate chromatin structure and transcription. Cell 108, 475–487 (2002).
Varga-Weisz, P. Chromatin remodeling factors and DNA replication. Prog. Mol. Subcell. Biol. 38, 1–30 (2005).
Ataian, Y. & Krebs, J.E. Five repair pathways in one context: chromatin modification during DNA repair. Biochem. Cell Biol. 84, 490–504 (2006).
Jaskelioff, M., Van Komen, S., Krebs, J.E., Sung, P. & Peterson, C.L. Rad54p is a chromatin remodeling enzyme required for heteroduplex DNA joint formation with chromatin. J. Biol. Chem. 278, 9212–9218 (2003).
van Attikum, H. & Gasser, S.M. ATP-dependent chromatin remodeling and DNA double-strand break repair. Cell Cycle 4, 1011–1014 (2005).
Eisen, J.A., Sweder, K.S. & Hanawalt, P.C. Evolution of the SNF2 family of proteins: subfamilies with distinct sequences and functions. Nucleic Acids Res. 23, 2715–2723 (1995).
Flaus, A., Martin, D.M., Barton, G.J. & Owen-Hughes, T. Identification of multiple distinct Snf2 subfamilies with conserved structural motifs. Nucleic Acids Res. 34, 2887–2905 (2006).
Fazzio, T.G. & Tsukiyama, T. Chromatin remodeling in vivo: evidence for a nucleosome sliding mechanism. Mol. Cell 12, 1333–1340 (2003).
Whitehouse, I. & Tsukiyama, T. Antagonistic forces that position nucleosomes in vivo. Nat. Struct. Mol. Biol. 13, 633–640 (2006).
Papamichos-Chronakis, M., Krebs, J.E. & Peterson, C.L. Interplay between Ino80 and Swr1 chromatin remodeling enzymes regulates cell cycle checkpoint adaptation in response to DNA damage. Genes Dev. 20, 2437–2449 (2006).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Shen, X., Mizuguchi, G., Hamiche, A. & Wu, C. A chromatin remodelling complex involved in transcription and DNA processing. Nature 406, 541–544 (2000).
Morrison, A.J. et al. INO80 and γ-H2AX interaction links ATP-dependent chromatin remodeling to DNA damage repair. Cell 119, 767–775 (2004).
Raghuraman, M.K. et al. Replication dynamics of the yeast genome. Science 294, 115–121 (2001).
McCarroll, R.M. & Fangman, W.L. Time of replication of yeast centromeres and telomeres. Cell 54, 505–513 (1988).
Alvino, G.M. et al. Replication in hydroxyurea: it's a matter of time. Mol. Cell. Biol. 27, 6396–6406 (2007).
Yabuki, N., Terashima, H. & Kitada, K. Mapping of early firing origins on a replication profile of budding yeast. Genes Cells 7, 781–789 (2002).
Feng, W. et al. Genomic mapping of single-stranded DNA in hydroxyurea-challenged yeasts identifies origins of replication. Nat. Cell Biol. 8, 148–155 (2006).
Wyrick, J.J. et al. Genome-wide distribution of ORC and MCM proteins in S. cerevisiae: high-resolution mapping of replication origins. Science 294, 2357–2360 (2001).
Nieduszynski, C.A., Knox, Y. & Donaldson, A.D. Genome-wide identification of replication origins in yeast by comparative genomics. Genes Dev. 20, 1874–1879 (2006).
Santocanale, C. & Diffley, J.F.A. Mec1- and Rad53-dependent checkpoint controls late-firing origins of DNA replication. Nature 395, 615–618 (1998).
Shirahige, K. et al. Regulation of DNA-replication origins during cell-cycle progression. Nature 395, 618–621 (1998).
Tercero, J.A. & Diffley, J.F. Regulation of DNA replication fork progression through damaged DNA by the Mec1/Rad53 checkpoint. Nature 412, 553–557 (2001).
Fazzio, T.G. et al. Widespread collaboration of Isw2 and Sin3-Rpd3 chromatin remodeling complexes in transcriptional repression. Mol. Cell. Biol. 21, 6450–6460 (2001).
Chang, M., Bellaoui, M., Boone, C. & Brown, G.W. A genome-wide screen for methyl methanesulfonate-sensitive mutants reveals genes required for S phase progression in the presence of DNA damage. Proc. Natl. Acad. Sci. USA 99, 16934–16939 (2002).
Sugino, A. Yeast DNA polymerases and their role at the replication fork. Trends Biochem. Sci. 20, 319–323 (1995).
Burgers, P.M. Eukaryotic DNA polymerases in DNA replication and DNA repair. Chromosoma 107, 218–227 (1998).
Papamichos-Chronakis, M. & Peterson, C.L. The Ino80 chromatin-remodeling enzyme regulates replisome function and stability. Nat. Struct. Mol. Biol. 15, 338–345 (2008).
Donaldson, A.D. et al. CLB5-dependent activation of late replication origins in S. cerevisiae. Mol. Cell 2, 173–182 (1998).
Falbo, K.B. & Shen, X. Chromatin remodeling in DNA replication. J. Cell. Biochem. 97, 684–689 (2006).
Ye, X. et al. Defective S phase chromatin assembly causes DNA damage, activation of the S phase checkpoint, and S phase arrest. Mol. Cell 11, 341–351 (2003).
Collins, N. et al. An ACF1-ISWI chromatin-remodeling complex is required for DNA replication through heterochromatin. Nat. Genet. 32, 627–632 (2002).
Bozhenok, L., Wade, P.A. & Varga-Weisz, P. WSTF-ISWI chromatin remodeling complex targets heterochromatic replication foci. EMBO J. 21, 2231–2241 (2002).
Poot, R.A. et al. The Williams syndrome transcription factor interacts with PCNA to target chromatin remodelling by ISWI to replication foci. Nat. Cell Biol. 6, 1236–1244 (2004).
Li, J., Santoro, R., Koberna, K. & Grummt, I. The chromatin remodeling complex NoRC controls replication timing of rRNA genes. EMBO J. 24, 120–127 (2005).
Thomas, B.J. & Rothstein, R. The genetic control of direct-repeat recombination in Saccharomyces: the effect of Rad52 and Rad1 on mitotic recombination at GAL10, a transcriptionally regulated gene. Genetics 123, 725–738 (1989).
Zhao, X., Muller, E.G. & Rothstein, R. A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol. Cell 2, 329–340 (1998).
Guldener, U., Heck, S., Fielder, T., Beinhauer, J. & Hegemann, J.H. A new efficient gene disruption cassette for repeated use in budding yeast. Nucleic Acids Res. 24, 2519–2524 (1996).
Goldstein, A.L. & McCusker, J.H. Three new dominant drug resistance cassettes for gene disruption in Saccharomyces cerevisiae. Yeast 15, 1541–1553 (1999).
Lindstrom, K.C., Vary, J.C. Jr, Parthun, M.R., Delrow, J. & Tsukiyama, T. Isw1 functions in parallel with the NuA4 and Swr1 complexes in stress-induced gene repression. Mol. Cell. Biol. 26, 6117–6129 (2006).
Gelbart, M.E., Bachman, N., Delrow, J., Boeke, J.D. & Tsukiyama, T. Genome-wide identification of Isw2 chromatin-remodeling targets by localization of a catalytically inactive mutant. Genes Dev. 19, 942–954 (2005).
Nieduszynski, C.A., Hiraga, S., Ak, P., Benham, C.J. & Donaldson, A.D. OriDB: a DNA replication origin database. Nucleic Acids Res. 35, D40–D46 (2007).
Acknowledgements
We thank G.M. Alvino, S. Biggins, B.J. Brewer, D. Collingwood, H.S. Malik, M.K. Raghuraman and members of the Tsukiyama laboratory for critical reading of the manuscript; J.F.X. Diffley (Cancer Research UK) for the YJT80 strain; members of the Biggins and Brewer-Raghuraman laboratories for helpful discussions; D. Collingwood for help with analysis of replication data; H.S. Malik for the manuscript title suggestion; C.L. Peterson for sharing data before publication; and G.M. Alvino for a DNA-labeling protocol for microarrays, technical advice on density transfer experiments and help with replication data analysis. This work was supported by grants to T.T. from the US National Institutes of Health (NIH) and the Leukemia and Lymphoma Society. J.A.V. and T.J.K. were supported in part by training grants from the NIH.
Author information
Authors and Affiliations
Contributions
J.A.V. designed and conducted the drug sensitivity, FACS, DNA microarray and DNA replication experiments, and wrote the manuscript; T.J.K. designed and conducted the ChIP experiments; T.T. supervised the project and contributed as the senior author. All authors contributed to the preparation of the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–4, Supplementary Discussion and Supplementary Methods (PDF 3821 kb)
Rights and permissions
About this article
Cite this article
Vincent, J., Kwong, T. & Tsukiyama, T. ATP-dependent chromatin remodeling shapes the DNA replication landscape. Nat Struct Mol Biol 15, 477–484 (2008). https://doi.org/10.1038/nsmb.1419
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1419
This article is cited by
-
Mechanisms of chromatin-based epigenetic inheritance
Science China Life Sciences (2022)
-
ATM-mediated phosphorylation of the chromatin remodeling enzyme BRG1 modulates DNA double-strand break repair
Oncogene (2015)
-
Stabilization and targeting of INO80 to replication forks by BAP1 during normal DNA synthesis
Nature Communications (2014)