Key Points
-
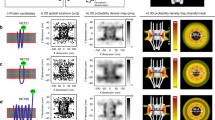
The classic nuclear protein import cycle functions as a biological molecular ratchet and is powered by the Ran GTPase that modulates interactions between carrier molecules and their cargoes. In the cytoplasm, cargo molecules carrying a classic nuclear localization signal (NLS) sequence are attached to the carrier importin-β by the importin-α adaptor. Cargoes are released in the nucleus following RanGTP binding to importin-β, after which the importins are recycled.
-
Nuclear pores facilitate the equilibration of cargo:carrier complexes between the nucleus and the cytoplasm. Transport directionality is imposed by import-complex dissociation by RanGTP in the nucleus and by RanGTP hydrolysis in the cytoplasm. Energy is used to orchestrate the binding and release of cargoes in the appropriate compartments rather than to move material directly through the pores.
-
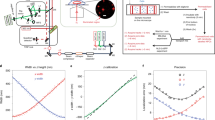
Movement of material backwards and forwards through nuclear pores is facilitated by interactions with nuclear pore proteins (nucleoporins) that contain Phe-Gly (FG) sequence repeats. These proteins also obstruct the passage of proteins that lack an NLS.
-
Nucleoporins on the nuclear face of the NPC accelerate import-complex disassembly and provide a molecular ratchet to prevent futile cycles in which cargo is returned to the cytoplasm.
-
Molecular flexibility is important in modulating interactions between importins and their partners.
Abstract
The nuclear import of proteins through nuclear pore complexes (NPCs) illustrates how a complex biological function can be generated by a spatially and temporally organized cycle of interactions between cargoes, carriers and the Ran GTPase. Recent work has given considerable insight into this process, especially about how interactions are coordinated and the basis for the molecular recognition that underlies the process. Although considerable progress has been made in identifying and characterizing the molecular interactions in the soluble phase that drive the nuclear protein import cycle, understanding the precise mechanism of translocation through NPCs remains a major challenge.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Görlich, D. & Kutay, U. Transport between the cell nucleus and the cytoplasm. Ann. Rev. Cell Dev. Biol. 15, 607–660 (1999).
Macara, I. G. Transport into and out of the nucleus. Microbiol. Mol. Biol. Rev. 65, 570–594 (2001).
Chook, Y. M. & Blobel, G. Karyopherins and nuclear import. Curr. Opin. Struct. Biol. 11, 703–715 (2001).
Conti, E. & Izaurralde, E. Nuclear transport enters the atomic age. Curr. Opin. Cell Biol. 13, 310–319 (2001).
Fahrenkrog, B. & Aebi, U. The nuclear pore complex: nucleocytoplasmic transport and beyond. Nature Rev. Mol. Cell Biol. 4, 757–766 (2003).
Mosammaparast, N. & Pemberton, L. F. Karyopherins: from nuclear-transport mediators to nuclear-function regulators. Trends Cell Biol. 14, 547–556 (2004).
Pemberton, L. F. & Paschal, B. M. Mechanisms of receptor-mediated nuclear import and nuclear export. Traffic 6, 187–198 (2005).
Madrid, A. S. & Weis, K. Nuclear transport is becoming crystal clear. Chromosoma 115, 98–109 (2006).
Conti, E., Müller, C. W. & Stewart, M. Karyopherin flexibility in nucleocytoplasmic transport. Curr. Opin. Struct. Biol. 16, 237–244 (2006).
Stewart, M. et al. Molecular mechanism of translocation through nuclear pore complexes during nuclear protein import. FEBS Lett. 498, 145–149 (2001).
Weis, K. Regulating access to the genome. Nucleocytoplasmic transport throughout the cell cycle. Cell 112, 441–451 (2003).
Feldherr, C. M., Akin, D. & Cohen, R. J. Regulation of functional nuclear pores size in fibroblasts. J. Cell Sci. 114, 4621–4627 (2001).
Rout, M. P. & Wente, S. R. Pores for thought: nuclear pore complex proteins. Trends Cell Biol. 4, 357–365 (1994).
Rout, M. P. et al. The yeast nuclear pore complex: composition, architecture, and transport mechanism. J. Cell Biol. 148, 635–651 (2000). A landmark paper that established the complete protein composition of yeast NPCs together with the location of each nucleoporin as determined by electron microscopy. The authors also proposed an entropic gating mechanism to prevent entry of inappropriate material into the NPC transport channel.
Cronshaw, J. M., Krutchinsky, A. N., Zhang, W., Chait, B. T. & Matunis, M. J. Proteomic analysis of the mammalian nuclear pore complex. J. Cell Biol. 158, 915–927 (2002).
Timney, B. L. et al. Simple kinetic relationships and nonspecific competition govern nuclear import rates in vivo. J. Cell Biol. 175, 579–593 (2006).
Mosammaparast, N. et al. Nuclear import of histone H2A and H2B is mediated by a network of karyopherins. J. Cell Biol. 153, 251–262 (2001).
Lee, B. J. et al. Rules for nuclear localization sequence recognition by karyopherinβ2. Cell 126, 543–558 (2006).
Ribbeck, K. & Görlich, D. Kinetic analysis of translocation through nuclear pore complexes. EMBO J. 20, 1320–1330 (2001). Presents a careful analysis of nuclear protein import kinetics and proposes a selective phase model based on the formation of a hydrogel formed by interactions between the hydrophobic cores of FG repeats.
Ribbeck, K., Lipowsky, G., Kent, H. M., Stewart, M. & Görlich, D. NTF2 mediates nuclear import of Ran. EMBO J. 17, 6587–6598 (1998).
Smith, A. E., Brownawell, A. & Macara, I. G. Nuclear import of Ran is mediated by the transport factor NTF2. Curr. Biol. 8, 1403–1406 (1998).
Conti, E., Uy, M., Leighton, L., Blobel, G. & Kuriyan, J. Crystallographic analysis of the recognition of a nuclear localization signal by the nuclear import factor karyopherin-α. Cell 94, 193–204 (1998). Pioneering determination of the structure of yeast importin-α and the way in which it binds NLSs.
Conti, E. & Kuriyan, J. Crystallographic analysis of the specific yet versatile recognition of distinct nuclear localization signals by karyopherin-α. Structure Fold. Des. 8, 329–338 (2000).
Fontes, M. R., The, T. & Kobe, B. Structural basis of recognition of monopartite and bipartite nuclear localization sequences by mammalian importin-α. J. Mol. Biol. 297, 1183–1194 (2000).
Hodel, M. R., Corbett, A. H. & Hodel, A. E. Dissection of a nuclear localization signal. J. Biol. Chem. 276, 1317–1325 (2001).
Catimel, B. et al. Biophysical characterization of interactions involving importin-α during nuclear import. J. Biol. Chem. 276, 34189–34198 (2001).
Rodriguez, M. et al. A cytotoxic ribonuclease variant with a discontinuous nuclear localization signal constituted by basic residues scattered over three areas of the molecule. J. Mol. Biol. 360, 548–557 (2006).
Lee, S. J. et al. The structure of importin-β bound to SREBP-2: nuclear import of a transcription factor. Science 302, 1571–1575 (2003).
Cingolani, G., Bednenko, J., Gillespie, M. T. & Gerace, L. Molecular basis for the recognition of a nonclassical nuclear localization signal by importin β. Mol. Cell 10, 1345–1353 (2002).
Görlich, D., Henklein, P., Laskey, R. A. & Hartmann, E. A 41 amino acid motif in importin-α confers binding to importin-β and hence transit into the nucleus. EMBO J. 15, 1810–1817 (1996).
Weis, K., Ryder, U. & Lamond, A. I. The conserved amino-terminal domain of hSRP1 α is essential for nuclear protein import. EMBO J. 15, 1818–1825 (1996).
Kobe, B. Autoinhibition by an internal nuclear localization signal revealed by the crystal structure of mammalian importin α. Nature Struct. Biol. 6, 388–397 (1999). Pioneering structural paper that proposed how the importin-α IBB domain could have an autoinhibitory function that facilitates cargo release in the nucleus.
Matsuura, Y. & Stewart, M. Structural basis for the assembly of a nuclear export complex. Nature 432, 872–877 (2004). Describes the structure of the yeast CAS:RanGTP:importin-α complex, showing the central role of the IBB domain in preventing futile cycles in which cargo is returned to the cytoplasm. Also proposes a spring-loaded model to account for how karyopherin flexibility can mediate the release of importin-α following GTP hydrolysis on Ran.
Matsuura, Y., Lange, A., Harreman, M. T., Corbett, A. H. & Stewart, M. Structural basis for Nup2p function in cargo release and karyopherin recycling in nuclear import. EMBO J. 22, 5358–5369 (2003).
Matsuura, Y. & Stewart, M. Nup50/Npap60 function in nuclear protein import complex disassembly and importin recycling. EMBO J. 24, 3681–3689 (2005). Demonstration of active displacement of cargo from importin-α by NUP50 and how this can generate a molecular ratchet to prevent futile cycles.
Goldfarb, D. S., Corbett, A. H., Mason, D. A., Harreman, M. T. & Adam, S. A. Importin α: a multi-purpose nuclear receptor. Trends Cell Biol. 14, 505–514 (2004).
Hodel, H. E. et al. Nuclear localization signal-receptor affinity correlates with in vivo localization in S. cerevisiae. J. Biol. Chem. 281, 23545–23556 (2006).
Yang, W., Gelles, J. & Musser, S. M. Imaging of single-molecule translocation through nuclear pore complexes. Proc. Natl Acad. Sci. USA 101, 12887–12892 (2004). Direct demonstration of the random-walk diffusion of import complexes in the central transport channel of NPCs.
Kubitscheck, U. et al. Nuclear transport of single molecules: dwell times at the nuclear pore complex. J. Cell Biol. 168, 233–243 (2005).
Wang, W. & Musser, S. M. Nuclear import time and transport efficiency depend on importin β concentration. J. Cell Biol. 174, 951–961 (2006).
Bayliss, R. et al. Interaction between NTF2 and xFxFG nucleoporins is required for the nuclear import of RanGDP. J. Mol. Biol. 293, 579–593 (1999).
Bayliss, R., Littlewood, T. & Stewart, M. Structural basis for the interaction between FxFG nucleoporin repeats and importin-β in nuclear trafficking. Cell 102, 99–108 (2000). Describes the structural basis for the interaction between importin-β and FG-nucleoporin cores.
Bayliss, R., Littlewood, T., Strawn, L. A., Wente, S. R. & Stewart, M. GLFG and FxFG nucleoporins bind to overlapping sites on importin-β. J. Biol. Chem. 277, 50597–50606 (2002).
Bayliss, R. et al. Structural basis for the interaction between NTF2 and nucleoporin FxFG repeats. EMBO J. 21, 2843–2853 (2002).
Fribourg, S. Braun, I. C., Izaurralde, E. & Conti, E. Structural basis for the recognition of a nucleoporin FG repeat by the NTF2-like domain of the TAP/p15 mRNA nuclear export factor. Mol. Cell 8, 645–656 (2001).
Grant, R. P., Neuhaus, D. N. & Stewart, M. Structural basis for the interaction between the Tap/NXF1 UBA domain and FG nucleoporins at 1Å resolution. J. Mol. Biol. 326, 849–858 (2003).
Rexach, M. & Blobel, G. Protein import into nuclei: association and dissociation reactions involving transport substrate, transport factors, and nucleoporins. Cell 83, 683–692 (1995).
Ribbeck, K. & Görlich, D. The permeability barrier of nuclear pore complexes appears to operate via hydrophobic exclusion. EMBO J. 21, 2664–2671 (2002).
Strawn, L. A., Shen, T. & Wente, S. R. The GLFG regions of Nup116p and Nup100p serve as binding sites for both Kap95p and Mex67p at the nuclear pore complex. J. Biol. Chem. 276, 6445–6452 (2001).
Strawn, L. A., Shen, T., Shulga, N., Goldfarb, D. S. & Wente, S. R. Minimal nuclear pore complexes define FG repeat domains essential for transport. Nature Cell Biol. 6, 197–206 (2004). Exploration of the importance of the FG-repeat regions of nucleoporins using an innovative method for deleting these regions in S. cerevisiae . Whereas up to half the mass of FG repeats can be deleted using particular groups of nucleoporins, deletion from several selected nucleoporins has a marked effect.
Tran, E. J. & Wente, S. R. Dynamic nuclear pore complexes: life on the edge. Cell 125, 1041–1053 (2006).
Grote, M., Kubitscheck, U., Reichelt, R. & Peters, R. Mapping of nucleoporins to the centre of the nuclear pore complex by post-embedding immunogold electron microscopy. J. Cell Sci. 108, 2963–2972 (1995).
Denning, D. P., Uversky, V., Patel, S. S., Fink, A. L. & Rexach, M. The Saccharomyces cerevisiae nucleoporin Nup2p is a natively unfolded protein. J. Biol. Chem. 277, 33447–33455 (2002).
Denning, D. P., Patel, S. S., Uversky, V., Fink, A. L. & Rexach, M. Disorder in the nuclear pore complex: the FG repeat regions of nucleoporins are natively unfolded. Proc. Natl Acad. Sci. USA 100, 2450–2455 (2003).
Paulillo, S. M. et al. Nucleoporin domain topology is linked to transport status of the nuclear pore complex. J. Mol. Biol. 351, 784–798 (2005).
Liu, S. M. & Stewart, M. Structural basis for the high-affinity binding of nucleoporin Nup1p to the Saccharomyces cerevisiae importin-β homologue, Kap95p. J. Mol. Biol. 349, 515–525 (2005).
Bednenko, J., Cingolani, G. & Gerace, L. Importin β contains a COOH-terminal nucleoporin binding region important for nuclear transport. J. Cell Biol. 162, 391–401 (2003).
Ben-Efraim, I. & Gerace, L. Gradient of increasing affinity of importin-β for nucleoporins along the pathway of nuclear import. J. Cell Biol. 152, 411–418 (2001).
Pyhtila, B. & Rexach, M. A gradient of affinity for the karyopherin Kap95p along the yeast nuclear pore complex. J. Biol. Chem. 278, 42699–42709 (2003).
Zeitler, B. & Weis, K. The FG-repeat asymmetry of the nuclear pore complex is dispensable for bulk nucleocytoplasmic transport in vivo. J. Cell Biol. 167, 583–590 (2004).
Nachury, M. V. & Weis, K. The direction of transport through the nuclear pore can be inverted. Proc. Natl Acad. Sci. USA 96, 9622–9627 (1999). Direct demonstration that the directionality of nuclear transport can be reversed by reversing the RanGTP gradient. This work shows conclusively that directionality of transport is driven by the Ran nucleotide state and not by a nucleoporin-affinity gradient for importin-β.
Lim, R. Y. et al. Flexible phenylalanine-glycine nucleoporins as entropic barriers to nucleocytoplasmic transport. Proc. Natl Acad. Sci. USA 103, 9512–9517 (2006). This paper uses atomic force microscopy to show that FG-repeats can form molecular brushes when bound to gold particles in vitro.
Rout, M. P., Aitchison, J. D., Magnasco, M. O. & Chait, B. T. Virtual gating and nuclear transport: the hole picture. Trends Cell Biol. 13, 622–628 (2003).
Tanford, C. The Physical Chemistry of Macromolecules. (Wiley, New York, 1961).
de Gennes, P. G. Conformations of polymers attached to an interface. Macromolecules 13, 1069–1075 (1980).
Frey, S., Richter, R. P. & Görlich, D. FG-rich repeats of nuclear pore proteins form a three-dimensional meshwork with hydrogel-like properties. Science 314, 815–817 (2006).
Doi, M. & Edwards, S. F. The Theory of Polymer Dynamics (Oxford University Press, Oxford, 1986).
Gilchrist, D., Mykytka, B. & Rexach, M. Accelerating the rate of disassembly of karyopherin–cargo complexes. J. Biol. Chem. 277, 18161–18172 (2002).
Franke, W. W. Structure, biochemistry and functions of the nuclear envelope. Internat. Rev. Cytol. (Suppl. 4), 71–236 (1974).
Richardson, W. D., Mills, A. D., Dilworth, S. M., Laskey, R. A. & Dingwall, C. Nuclear protein migration involves two steps: rapid binding at the nuclear envelope followed by slower translocation through nuclear pores. Cell 52, 655–664 (1988).
Walther, T. C. et al. The cytoplasmic filaments of the nuclear pore complex are dispensable for selective nuclear protein import. J. Cell Biol. 158, 63–77 (2002).
Rexach, M. & Blobel, G. Protein import into nuclei: association and dissociation reactions involving transport substrate, transport factors and nucleoporins. Cell 83, 683–692 (1995).
Vetter, I. R., Arndt, A., Kutay, U., Görlich, D. & Wittinghofer, A. Structural view of the Ran-importin β interaction at 2. 3 Å resolution. Cell 97, 635–646 (1999).
Lee, S. J., Matsuura, Y., Liu, S. M. & Stewart, M. Structural basis for nuclear import complex dissociation by RanGTP. Nature 435, 693–696 (2005). The structure of full-length yeast importin-β bound to RanGTP shows how Ran binding alters the pitch of the importin-β helicoid and facilitates release of the importin-α IBB domain.
Cingolani, G., Petosa, C., Weis, K. & Müller C. W. Structure of importin-β bound to the IBB domain of importin-α. Nature 399, 221–229 (1999). Structure of the IBB:importin-β complex showing how the 19 HEAT repeats of importin-β are arranged to form a helicoidal molecule that coils around the α-helical IBB domain. Comparison between different crystal forms indicated that importin-β might be flexible.
Görlich, D. et al. A novel class of RanGTP binding proteins. J. Cell Biol. 138, 65–80 (1997).
Chook, Y. M. & Blobel, G. Structure of the nuclear transport complex karyopherin-β2–Ran x GppNHp. Nature 399, 230–237 (1999).
Bischoff, F. R. & Görlich, D. RanBP1 is crucial for the release of RanGTP from importin-β-related nuclear transport factors. FEBS Lett. 419, 249–254 (1997).
Cook, A. et al. The structure of the nuclear export receptor Cse1 in its cytosolic state reveals a closed conformation incompatible with cargo binding. Mol. Cell 18, 355–367 (2005). Direct demonstration of the dramatic conformational change in the yeast homologue of CAS, Cse1. In the absence of RanGTP and importin-α, the Cse1 helicoid adopts a closed conformation in which the N terminus becomes bound to a region near the centre of the molecule.
Fukuhara, N., Fernandez, E., Ebert, J., Conti, E. & Svergun, D. Conformational variability of nucleo-cytoplasmic transport factors. J. Biol. Chem. 279, 2176–2181 (2004). Low-angle X-ray scattering is used to show that several β-karyopherins can adopt different structures in solution, leading to the concept of these molecules having flexible structures.
Stewart, M. Molecular recognition in nuclear trafficking. Science 302, 1513–1514 (2003).
Görlich, D., Seewald, M. J. & Ribbeck, K. Characterization of Ran-driven cargo transport and the RanGTPase system by kinetic measurements and computer simulation. EMBO J. 22, 1088–1100 (2003).
Riddick, G. & Macara, I. G. A systems analysis of importin-α:β mediated nuclear protein import. J. Cell Biol. 168, 1027–1038 (2005). A comprehensive simulation of the classic nuclear protein import cycle that gives several novel and unanticipated functional insights.
Smith, A., Slepchenko, B. M., Schaff, J. C. Loew, L. M. & Macara, I. G. Systems analysis of Ran transport. Science 295, 488–491 (2002).
Becskei, A. & Mattaj, I. W. Quantitative models of nuclear transport. Curr. Opin. Cell Biol. 17, 27–34 (2005).
Akey, C. W. Structural plasticity of the nuclear pore complex. J. Mol. Biol. 248, 273–293 (1995).
Akey, C. W. & Radermacher, M. Architecture of the Xenopus nuclear pore complex revealed by three-dimensional cryo-electron microscopy. J. Cell Biol. 122, 1–19 (1993).
Stoffler, D. et al. Cryo-electron microscopy provides novel insights into nuclear pore architecture: implications for nucleocytoplasmic transport. J. Mol. Biol. 328, 119–130 (2003). Comprehensive analysis of NPC morphology using cryo-EM and tomography.
Lim, R. Y. H. & Fahrenkrog, B. The nuclear pore complex up close. Curr. Opin. Cell Biol. 18, 342–347 (2006).
Lim, R. Y., Aebi U. & Stoffler D. From the trap to the basket: getting to the bottom of the nuclear pore complex. Chromosoma 115, 15–26 (2006).
Beck, M. et al. Nuclear pore complex structure and dynamics revealed by cryoelectron tomography. Science 306, 1387–1390 (2004).
Fahrenkrog, B. et al. Domain-specific antibodies reveal multiple-site topology of Nup153 within the nuclear pore complex. J. Struct. Biol. 140, 254–267 (2002).
Devos, D. et al. Components of coated vesicles and nuclear pore complexes share a common molecular architecture. PloS Biol. 2, e380 (2004).
Vetter, I. R., Nowak, C., Nishimoto, T., Kuhlmann, J. & Wittinghofer, A. Structure of a Ran-binding domain complexed with Ran bound to a GTP analogue: implications for nuclear transport. Nature 398, 39–46 (1999).
Stewart, M., Kent, H. M. & McCoy, A. J. Structural basis for molecular recognition between nuclear transport factor 2 (NTF2) and the GDP-bound form of the Ras-family GTPase, Ran. J. Mol. Biol. 277, 635–646 (1998).
Bullock, T. L., Clarkson, W. D., Kent, H. M. & Stewart, M. The 1.6 Å resolution crystal structure of nuclear transport factor 2 (NTF2). J. Mol. Biol. 260, 422–431 (1996).
Renault, L., Kuhlmann, J., Henkel, A. & Wittinghofer, A. Structural basis for guanine nucleotide exchange on ran by the regulator of chromosome condensation (RCC1). Cell 105, 245–255 (2001).
Hillig, R. C. et al. The crystal structure of rna1p: a new fold for a GTPase activating protein. Mol. Cell 3, 781–791 (1999).
Seewald, M. J., Korner, C., Wittinghofer, A. & Vetter, I. R. RanGAP mediates GTP hydrolysis without an arginine finger. Nature 415, 662–666 (2002).
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Supplementary information
Supplementary information S1 (movie)
Structure of importin-a bound to the nucleoplasmin nuclear localization signal (NLS). Importin-α (green) is constructed from a tandem series of Armadillo (ARM) repeats that stack to form a gently curving banana-like molecule. NLSs (blue) bind to the inner concave surface. See also Figure 2b. (PDB accession number 1EYJ). (MOV 4209 kb)
Supplementary information S2 (movie)
Structure of importin-β complexed with the importin-α IBB (importin-β binding) domain. 'Importin-β' (cyan) is constructed from 19 tandem HEAT repeats, each of which contains two α-helices (See BOX 2). The HEAT repeats stack to form a helicoidal molecule that coils around the IBB domain α-helix (red) like a snake around its prey. (PDB accession number 1QGK). (MOV 8513 kb)
Supplementary information S3 (movie)
Structure of Ran showing the structural changes between the GDP- and GTP-bound states. The larger switch I and smaller switch II loops change conformation dramatically depending on the state of the bound nucleotide (GTPbound is red; GDP-bound is blue) and these changes control the way in which Ran interacts with its partners in the nuclear protein import pathway. The remainder of the Ran chain (shown as cyan for GTP-bound and yellow for GDP-bound) is virtually unchanged by the state of the bound nucleotide. (PDB accession files 1BYU (RanGDP) and 1RRP (RanGTP). Only residues 8-177 are shown. (MOV 6825 kb)
Supplementary information S4 (movie)
Conformational changes in importin-β between different functional states. Comparison of the structures of importin-β when bound to either the IBB domain (cyan) or RanGTP (yellow) shows the large conformational change that occurs between different functional states. When bound to RanGTP, the importin-β helicoid pitch increases dramatically so that it no longer matches the IBB domain's α-helix, thus leading to import complex disassembly (see also Figure 3c). IBB domain is red. (PDB accession codes 1QGK and 2BKU). (MOV 13233 kb)
Supplementary information S5 (movie)
Structure of the complex between yeast importin-β (Kap95p) and RanGTP. Importin-β (yellow) coils around RanGTP (cyan) interacting with it at three sites (see also Figure 3 a,b). GTP is shown in space-filling format. (PDB accession code 2BKU). (MOV 8743 kb)
Supplementary information S6 (movie)
Structure of the complex between Nup50 and importin-α. The Nup50 Nterminus (blue) binds to two sites on importin-α (green): a higher affinity site at the importin-α C-terminus (at the top of the molecule in this movie) and a lower affinity site that overlaps the NLS binding area (in the central region of the importin-α chain). Nup50 binding actively displaces nuclear localization signals (NLSs) from importin-α. (PDB accession code 2C1M). (MOV 3736 kb)
Supplementary information S7 (movie)
Structure of the CAS:importin-α:RanGTP nuclear export complex. CAS (yeast Cse1, yellow) coils around both RanGTP (cyan) and importin-α (green) in the complex. The IBB domain (blue) is sandwiched between CAS and importin-α. The binding of the IBB domain to the NLS binding sites is crucial for the formation of this complex and ensures that only cargo-free importin-α is exported to the nucleus, thus preventing futile transport cycles. (PDB accession code 1WA5). (MOV 9573 kb)
Supplementary information S8 (movie)
Structure of isolated CAS. Isolated CAS (cyan), corresponding to the species present in the cytoplasm after the export complex dissociates following RanGTP hydrolysis, assumes a "closed" conformation in which its N-terminus binds to a region near the centre of the molecule. (PDB accession code 1Z3H). (MOV 7422 kb)
Supplementary information S9 (movie)
Comparison between the open and closed forms of CAS. CAS undergoes a considerable conformational change between the open form assumed in the CAS:importin-α:RanGTP export complex (yellow) and the closed form (cyan) that results from the disassembly of this complex in the cytoplasm following GTP hydrolysis on Ran. The helicoidal pitch has changed substantially between the two forms (compare with S4, movie), consistent with the molecule being flexible. (PDB accession codes 1Z3H and 1WA5). (MOV 10126 kb)
Supplementary information S10 (movie)
Illustration of the conformational change in CAS between the open and closed states. The MolMov package (http://bioinfo.mbb.yale.edu/MolMovDB) was used to simulate the transition between the "open" and "closed" forms of CAS. (PDB accession codes 1Z3H and 1WA5). (MOV 3809 kb)
Supplementary information S11 (movie)
Structure of nuclear transport factor 2 (NTF2). NTF2 is a dimer constructed from two chains (cyan and yellow). Each chain generates a hydrophobic cavity to which RanGDP binds (see S12, movie), whereas a hydrophobic patch formed between the two NTF2 chains binds the FxFG repeats from nucleoporins. (PDB accession number 1GY6). (MOV 8015 kb)
Supplementary information S12 (movie)
Complex between nuclear transport factor 2 (NTF2) and RanGDP. Two RanGDP chains (red and yellow) bind to the hyrodophobic cavities in the NTF2 dimer (blue and cyan) to facilitate its transport into the nucleus for nucleotide exchange using RanGEF. GDP is shown as space-filling format. (PDB accession number 1A2K). (MOV 7023 kb)
Supplementary information S13 (movie)
Structure of the metazoan RanGEF, RCC1. RCC1 (cyan) is based on a seven-bladed propeller structure. It accelerates the rate of nucleotide exchange on Ran by stabilizing the nucleotide-free state. (PDB accession code 1A12). (MOV 3546 kb)
Supplementary information S14 (movie)
Structure of the yeast RanGAP, Rna1. RanGAP (green) is constructed from leucine-rich repeats (LLRs each based on an α-helix and a β-strand) and accelerates the rate of the Ran GTPase by ∼105. (PDB accession code 2CA6). (MOV 4027 kb)
Related links
Glossary
- Coiled coil
-
A protein fold in which two α-helices coil around one another.
- WD propeller
-
A protein fold formed by a series of repeating WD sequence motifs that structurally resemble the blades of a propeller.
- α-helical solenoid
-
A protein fold formed by successive repeats, each of which contains a number of α-helices that form a loop that is like a coil of a spring or solenoid.
- Adaptor proteins
-
Proteins that augment cellular responses by recruiting other proteins to a complex. They usually contain several protein:protein interaction domains.
- Scanning Ala mutagenesis
-
A technique in which successive residues in a region of a sequence of a protein are mutated to Ala to define those that influence an activity, such as binding to another protein.
- Random walk
-
The path followed by taking successive steps, each in a random direction relative to the previous step.
- Entropy
-
The component of free energy due to the disorder or randomness of the system. Increases in entropy (disorder) lower the free energy, whereas increases in order (lower entropy) increase energy.
- Enthalpy
-
The heat component of free energy that, in biological systems, is derived primarily from chemical bonds.
- Le Chatelier's principle
-
Le Chatelier's principle states that if a dynamic equilibrium is disturbed by changing conditions, the position of equilibrium moves to counteract the change.
- Thermal ratchet
-
A molecular mechanism by which the thermal (Brownian) motion of a particle is biased (or rectified) so that there is net movement in a particular direction. A thermal ratchet requires an input of energy in order not to violate the second law of thermodynamics.
- Helicoidal pitch
-
The distance along the axis of a helicoid corresponding to a rotation of 360°.
- Helicoid
-
A spiral that is shaped like the shell of a snail such that its radius decreases progressively along its axis.
- Sensitivity analysis
-
A procedure to determine the sensitivity of the outcomes of a model to changes in its parameters.
Rights and permissions
About this article
Cite this article
Stewart, M. Molecular mechanism of the nuclear protein import cycle. Nat Rev Mol Cell Biol 8, 195–208 (2007). https://doi.org/10.1038/nrm2114
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrm2114
This article is cited by
-
Structural anisotropy results in mechano-directional transport of proteins across nuclear pores
Nature Physics (2024)
-
Analysis of the effects of importin α1 on the nuclear translocation of IL-1α in HeLa cells
Scientific Reports (2024)
-
Progress in the study of parvovirus entry pathway
Virology Journal (2023)
-
B-type Plexins promote the GTPase activity of Ran to affect androgen receptor nuclear translocation in prostate cancer
Cancer Gene Therapy (2023)
-
Are we ready to fight the Nipah virus pandemic? An overview of drug targets, current medications, and potential leads
Structural Chemistry (2023)