Key Points
-
Bacterial genetics is based on the ability to select for rare mutants among large populations (1 in >1010) on the basis of the ability of the bacteria to survive lethal challenges, grow in the absence of supplements or take advantage of different growth substrates.
-
Bacterial genetics also uses a wide range of screens that are based on visual readouts of metabolic activity, including indicator agars (such as MacConkey and tetrazolium) and agars that contain chromogenic substrates (such as Xgal).
-
One of the most powerful genetic systems is the lactose operon of Escherichia coli. The lac genes encode an enzyme for breaking down lactose — lacZ (β-galactosidase) — and the membrane transporter for lactose — lacY (lac permease).
-
The lacZ gene can be fused to most other genes in many organisms. Transcriptional lacZ gene fusions produce wild-type β-galactosidase at levels that are determined by the promoter activity of the gene to which lacZ is fused. Translational lacZ gene fusions result in the production of hybrid β-galactosidase molecules that are encoded by both the gene to which lacZ is fused, and lacZ. The levels of production of the hybrid can depend on many post-transcriptional factors.
-
LacZ fusions allow the powerful tools that have been developed for the lac system to be applied to almost any bacterial system.
-
The properties of different LacZ fusions can provide important information about transcriptional and translational regulation as well as other cell functions, which include protein stability, protein folding, protein secretion and electron transport.
-
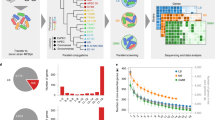
Modern 'high-tech' approaches that involve genomics and microarrays can be combined with genetic analysis to yield new insights into complex biological problems such as host–pathogen interactions.
Abstract
This article summarizes the general principles of selections and screens in Escherichia coli. The focus is on the lac operon, owing to its inherent simplicity and versatility. Examples of different strategies for mutagenesis and mutant discovery are described. In particular, the usefulness and effectiveness of simple colour-based screens are illustrated. The power of lac genetics can be applied to almost any bacterial system with gene fusions that hook any gene of interest to lacZ, which is the structural gene that encodes β-galactosidase. The diversity of biological processes that can be studied with lac genetics is remarkable and includes DNA metabolism, gene regulation and signal transduction, protein localization and folding, and even electron transport.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Beadle, G. W. & Tatum, E. L. Genetic control of biochemical reactions in Neurospora. Proc. Natl Acad. Sci. USA 27, 499–506 (1941).
Lederberg, J. Genetic recombination and linked segregation in Escherichia coli. Genetics 32, 505–525 (1947).
Luria, S. E. & Delbruck, M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics 28, 491–511 (1943).
Crick, F. H. C., Barnett, L., Brenner, S. & Watts-Tobin, R. J. General nature of the genetic code for proteins. Nature 192, 1227–1232 (1961).
De Lucia, P. & Cairns, J. Isolation of an E. coli strain with a mutation affecting DNA polymerase. Nature 224, 1164–1166 (1969). One of the first applications of a 'brute force' screen, which provided strong evidence against a role for DNA polymerase I in replication.
LaRossa, R. A. in Escherichia coli and Salmonella (ed. Neidhardt, F. C.) 2527–2587 (American Society for Microbiology, Washington, 1996).
Schreiber, S. L. The small-molecule approach to biology. Chem. Eng. News 81, 51–61 (2003).
Rickenberg, H. V., Cohen, G. N., Buttin, G. & Monod, J. La galactoside-permease d' Escherichia coli. Ann. Inst. Pasteur 91, 829–857 (1956).
Cohn, M. in The Operon (eds Miller, J. H. & Reznikoff, W. S.) 1–9 (Cold Spring Harbor Laboratory Press, New York, 1978).
Malamy, M. H. in The Lactose Operon (eds Beckwith, J. R. & Zipser, D.) 359–373 (Cold Spring Harbor Laboratory Press, New York, 1970).
Muller-Hill, B., Crapo, L. & Gilbert, W. Mutants that make more lac repressor. Proc. Natl Acad. Sci. USA 59, 1259–1264 (1968).
Silhavy, T. J., Berman, M. L. & Enquist, L. W. Experiments with gene fusions (Cold Spring Harbor Laboratory, New York, 1984).
Scaife, J. & Beckwith, J. R. Mutational alteration of the maximal level of Lac operon expression. Cold Spring Harb. Symp. Quant. Biol. 31, 403–408 (1966).
Eschenlauer, A. C. & Reznikoff, W. S. Escherichia coli catabolite gene activator protein mutants defective in positive control of lac operon transcription. J. Bacteriol. 173, 5024–5029 (1991).
Schwartz, D. & Beckwith, J. R. in The Lactose Operon (eds Beckwith, J. R. and Zipser, D.) 417–422 (Cold Spring Harbor Laboratory Press, New York, 1970).
Ullmann, A. & Perrin, D. in The Lactose Operon (eds Beckwith, J. R. and Zipser, D.) 143–172 (Cold Spring Harbor Laboratory Press, New York, 1970).
Langley, K. E., Villarejo, M. R., Fowler, A. V., Zamenhof, P. J. & Zabin, I. Molecular basis of β-galactosidase α-complementation. Proc. Natl Acad. Sci. USA 72, 1254–1257 (1975).
Beckwith, J. Restoration of operon activity by suppressors. Biochim. Biophys. Acta 76, 162–164 (1963).
Gronenborn, B. & Messing, J. Methylation of single-stranded DNA in vitro introduces new restriction endonuclease cleavage sites. Nature 272, 375–377 (1978).
Nghiem, Y., Cabrera, M., Cupples, C. G. & Miller, J. H. The mutY gene: a mutator locus in Escherichia coli that generates GC–TA transversions. Proc. Natl Acad. Sci. USA 85, 2709–2713 (1988). This paper describes a clever papillation screen that was used to identify a new mutator gene.
Bender, J. & Kleckner, N. Genetic evidence that Tn10 transposes by a nonreplicative mechanism. Cell 45, 801–815 (1986). An elegant use of sectored colonies that was applied to deduce a fundamental property of certain transposons.
Hand, N. J. & Silhavy, T. J. A practical guide to the construction and use of lac fusions in Escherichia coli. Methods Enzymol. 326, 11–35 (2000).
Debarbouille, M., Shuman, H. A., Silhavy, T. J. & Schwartz, M. Dominant constitutive mutations in malT, the positive regulator gene of the maltose regulon in Escherichia coli. J. Mol. Biol. 124, 359–371 (1978).
Danese, P. N., Snyder, W. B., Cosma, C. L., Davis, L. J. & Silhavy, T. J. The Cpx two-component signal transduction pathway of Escherichia coli regulates transcription of the gene specifying the stress-inducible periplasmic protease, DegP. Genes Dev. 9, 387–398 (1995).
Simons, R. W. & Kleckner, N. Translational control of IS10 transposition. Cell 34, 683–691 (1983).
Kennedy, D. Breakthrough of the year. Science 289, 2283 (2002).
Gottesman, S. Stealth regulation: biological circuits with small RNA switches. Genes Dev. 16, 2829–2842 (2002).
Pratt, L. A. & Silhavy, T. J. The response regulator SprE controls the stability of RpoS. Proc. Natl Acad. Sci. USA 93, 2488–2492 (1996).
Pratt, L. A. Osmotic and Growth-Phase Control of the Porin Regulon in Escherichia coli (Princeton Univ. Press, Princeton, 1996).
Muffler, A., Fischer, D., Altuvia, S., Storz, G. & Hengge-Aronis, R. The response regulator RssB controls stability of the σS subunit of RNA polymerase in Escherichia coli. EMBO J. 15, 1333–1339 (1996).
Becker, G., Klauck, E. & Hengge-Aronis, R. Regulation of RpoS proteolysis in Escherichia coli: the response regulator RssB is a recognition factor that interacts with the turnover element in RpoS. Proc. Natl Acad. Sci. USA 96, 6439–6444 (1999).
Zhou, Y., Gottesman, S., Hoskins, J. R., Maurizi, M. R. & Wickner, S. The RssB response regulator directly targets σS for degradation by ClpXP. Genes Dev. 15, 627–637 (2001).
Boos, W. & Shuman, H. Maltose/maltodextrin system of Escherichia coli: transport, metabolism, and regulation. Microbiol. Mol. Biol. Rev. 62, 204–229 (1998).
Snyder, W. B. & Silhavy, T. J. β-galactosidase is inactivated by intermolecular disulfide bonds and is toxic when secreted to the periplasm of Escherichia coli. J. Bacteriol. 177, 953–963 (1995).
Emr, S. D., Schwartz, M. & Silhavy, T. J. Mutations altering the cellular localization of the phage λ-receptor, an Escherichia coli outer membrane protein. Proc. Natl Acad. Sci. USA 75, 5802–5806 (1978). This paper describes the coupling of a selection with a screen using a chromogenic substrate for the identification of signal-sequence mutations.
Bassford, P. & Beckwith, J. Escherichia coli mutants accumulating the precursor of a secreted protein in the cytoplasm. Nature 277, 538–541 (1979).
Blobel, G. & Dobberstein, B. Transfer of proteins across membranes. I. Presence of proteolytically processed and unprocessed nascent immunoglobulin light chains on membrane-bound ribosomes of murine myeloma. J. Cell Biol. 67, 835–851 (1975).
Oliver, D. B. & Beckwith, J. E. coli mutant pleiotropically defective in the export of secreted proteins. Cell 25, 765–772 (1981). A selection on lactose tetrazolium agar was coupled with a conditional lethal screen to identify genes that specify components of the protein-secretion machinery.
Mori, H. & Ito, K. The Sec protein-translocation pathway. Trends Microbiol. 9, 494–500 (2001).
Blobel, G. Intracellular protein topogenesis. Proc. Natl Acad. Sci. USA 77, 1496–1500 (1980).
Bardwell, J. C., McGovern, K. & Beckwith, J. Identification of a protein required for disulfide bond formation in vivo. Cell 67, 581–589 (1991).
Debarbieux, L. & Beckwith, J. Electron avenue: pathways of disulfide bond formation and isomerization. Cell 99, 117–119 (1999).
Tian, H., Boyd, D. & Beckwith, J. A mutant hunt for defects in membrane protein assembly yields mutations affecting the bacterial signal recognition particle and Sec machinery. Proc. Natl Acad. Sci. USA 97, 4730–4735 (2000). This paper describes the convoluted, but ultimately fruitful, use of genetic selection to identify mutations that affect protein secretion.
Sadosky, A. B., Wiater, L. A. & Shuman, H. A. Identification of Legionella pneumophila genes required for growth within and killing of human macrophages. Infect. Immun. 61, 5361–5373 (1993).
Miller, J. H. A Short Course in Bacterial Genetics: A Laboratory Manual and Handbook for Escherichia coli and Related Bacteria (Cold Spring Harbor Laboratory Press, New York, 1992).
Kleckner, N., Bender, J. & Gottesman, S. Uses of transposons with emphasis on Tn10. Methods Enzymol. 204, 139–180 (1991).
Gibson, K. E. & Silhavy, T. J. The LysR homolog LrhA promotes RpoS degradation by modulating activity of the response regulator sprE. J. Bacteriol. 181, 563–571 (1999).
Blattner, F. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1474 (1997).
Smith, V., Botstein, D. & Brown, P. O. Genetic footprinting: a genomic strategy for determining a gene's function given its sequence. Proc. Natl Acad. Sci. USA 92, 6479–6483 (1995).
Akerley, B. J. et al. Systematic identification of essential genes by in vitro mariner mutagenesis. Proc. Natl Acad. Sci. USA 95, 8927–8932 (1998).
Hensel, M. et al. Simultaneous identification of bacterial virulence genes by negative selection. Science 269, 400–403 (1995).
Sassetti, C. M., Boyd, D. H. & Rubin, E. J. Comprehensive identification of conditionally essential genes in mycobacteria. Proc. Natl Acad. Sci. USA 98, 12712–12717 (2001).
Acknowledgements
Work in our laboratories has been supported by the National Institutes of Allergy and Infectious Disease and the National Science Foundation (H.A.S.), and by the National Institute of General Medical Sciences (T.J.S.). Thanks to R. Freudenberg for help with the photographs of the agar plates and N. Ruiz for help with figures.
Author information
Authors and Affiliations
Corresponding author
Glossary
- BACTERIOPHAGE
-
A virus that infects and replicates in bacteria. These are either lytic viruses, which always kill the host, or temperate viruses, which can either lyse the host cell or establish a stable relationship in which the bacteriophage genome is stably maintained in the host genome (see also lysogen).
- LYSOGEN
-
A bacterial strain that harbours the genome of a temperate bacteriophage.
- PLAQUE
-
A zone of lysed bacterial lawn that is formed by the growth of a bacteriophage.
- REVERSION ANALYSIS
-
The isolation and characterization of revertants, or secondary mutants, that regain a wild-type-like phenotype. This analysis can provide clues to the nature of the original mutation.
- PERMEABILIZED CELLS
-
Cells that have disrupted membranes.
- CHROMOGENIC
-
A colourless substrate that produces a coloured product following the action of one or more enzymes.
- INDUCER
-
A substance that causes a gene, or set of genes, to be expressed at a higher level.
- INSERTION SEQUENCES
-
(IS elements). Sequences that can insert themselves into target sites on DNA.
- PSEUDOREVERTANT
-
A revertant that does not re-establish a wild-type genotype.
- ENTERIC BACTERIA
-
Bacteria that normally inhabit the intestines.
- LEAKY DOWN PHENOTYPE
-
A phenotype in which there is a moderate decrease in gene function.
- CLONING VECTORS
-
Engineered plasmids with convenient cloning sites and markers.
- COMPLEMENTATION
-
The use of partial diploids to determine whether two mutations affect the same or different genes. If the mutations are in the same gene, they generally fail to complement each other and the diploid retains the mutant phenotype. By contrast, mutations in different genes usually complement one another and restore a wild-type phenotype to the diploid. However, exceptions to both cases abound.
- POLAR
-
A phenomenon in bacterial transcriptional units in which the transcription of downstream genes is decreased by a translation stop in an upstream gene.
- PAPILLATION
-
The formation of secondary colonies that grow on top of a colony that has already formed on a nutrient agar surface.
- HETERODUPLEX
-
A DNAmolecule that is formed by base pairing between strands that are derived from two DNA molecules that are not identical in sequence.
- REGULON
-
A group of transcriptional units or operons that are coordinately controlled by a regulator.
- SMALL RNAS
-
Small untranslated RNAs that function in regulation.
- STATIONARY PHASE
-
The phase of growth of a batch culture in which nutrients become limiting and growth ceases.
- PULSE-CHASE EXPERIMENT
-
An experiment in which a radioactive form of a precursor is added for a brief period ('pulse') so that it is incorporated into a macromolecule, then a non-radioactive identical chemical is added to dilute the incorporation of the radioactive species ('chase').
- SIGNAL SEQUENCE
-
An amino-acid sequence at the amino terminus of a protein that directs the protein to the secretion system for translocation across the cell membrane.
- MALTOPORIN
-
The Escherichia coli outer-membrane protein that functions as a diffusion channel for maltooligosaccharides.
- POLYTOPIC
-
A membrane-spanning protein with more than one membrane-spanning segment.
- FLUOROPHORE
-
A small molecule, or a part of a larger molecule, that can be excited by light to emit fluorescence.
Rights and permissions
About this article
Cite this article
Shuman, H., Silhavy, T. The art and design of genetic screens: Escherichia coli. Nat Rev Genet 4, 419–431 (2003). https://doi.org/10.1038/nrg1087
Issue Date:
DOI: https://doi.org/10.1038/nrg1087
This article is cited by
-
Scalable and automated CRISPR-based strain engineering using droplet microfluidics
Microsystems & Nanoengineering (2022)
-
Dual-barcoded shotgun expression library sequencing for high-throughput characterization of functional traits in bacteria
Nature Communications (2019)
-
A new recombineering system for precise genome-editing in Shewanella oneidensis strain MR-1 using single-stranded oligonucleotides
Scientific Reports (2019)
-
RpoS-independent evolution reveals the importance of attenuated cAMP/CRP regulation in high hydrostatic pressure resistance acquisition in E. coli
Scientific Reports (2017)
-
Isolating Escherichia coli strains for recombinant protein production
Cellular and Molecular Life Sciences (2017)