Key Points
-
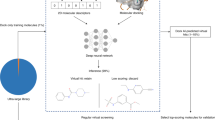
A practical and cost-effective embodiment of a chemogenomics approach to drug discovery involves the following steps:
-
Gene sequences for targets that have been identified by genomics approaches are cloned and expressed as target proteins that are suitable for screening with a probe library of small, drug-like chemical compounds.
-
These compounds are screened to find active hits using a quantitative universal binding assay.
-
Initial hits or quantitative structure–activity data emerging from the binding assay are analysed and used to formulate a selection strategy for the synthesis of additional compounds with improved properties.
-
These compounds are selected from a computer database of synthetically accessible analogues of the initial probe library, constructed using verified synthetic protocols and characterized by an extensive set of computed drug-related molecular properties.
-
The selected compounds are synthesized by parallel-synthesis methods and are subsequently tested to elaborate the structure–activity profile of the target under investigation, and refine the selection criteria for additional rounds of chemical synthesis and biological testing.
-
In each iteration, priority is assigned to the synthetic candidates using a multiobjective optimization process designed to assure that compounds are not only optimized for target binding affinity, but also have drug-like characteristics that will allow them to be used directly as tool compounds in appropriate cellular or biological model systems.
-
The potential for improved performance using such a strategy lies in the ability to rapidly follow up on initial hits through intelligent selection of related compounds from a computer database of synthetically accessible analogues with predefined synthesis recipes and predicted property profiles.
-
To address the full spectrum of targets emerging from genomics-based efforts, it will be necessary to physically screen probe libraries that span a wide range of chemotypes and contain hundreds of thousands of compounds.
-
These libraries will derive from synthetic strategies that could, in theory, produce billions of related analogues, which far exceeds the capabilities of conventional chemical-database management systems and data-modelling tools.
-
Thus, the following key questions need to be addressed:
-
How can huge combinatorial libraries be generated, represented, accessed, searched and manipulated?
-
What are the most appropriate chemical-property spaces, and how can they best be computed, sampled, visualized and validated?
-
What are the most effective ways to design, execute and analyse a combinatorial-chemistry experiment?
-
Successful deployment of such a system requires a new generation of computational tools that work effectively on a massive scale.
Abstract
The multitude of potential drug targets emerging from genome sequencing demands new approaches to drug discovery. A chemogenomics strategy, which involves the generation of small-molecule compounds that can be used both as tools to probe biological mechanisms and as leads for drug-property optimization, provides a highly parallel, industrialized solution. Key to the success of this strategy is an integrated suite of chemi-informatics applications that can allow the rapid and directed optimization of chemical compounds with drug-like properties using 'just-in-time' combinatorial chemical synthesis. An effective embodiment of this process requires new computational and data-mining tools that cover all aspects of library generation, compound selection and experimental design, and work effectively on a massive scale.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
International Human Genome Sequencing Consortium. Initial Sequencing and Analysis of the Human Genome. Nature 409, 860–921 (2001).
Agrafiotis, D. K., Bone, R. F., Salemme, F. R. & Soll, R. M. System and method for automatically generating chemical compounds with desired properties. US Patent 5,463,564 (1995).
Agrafiotis, D. K., Bone, R. F., Salemme, F. R. & Soll, R. M. System and method for automatically generating chemical compounds with desired properties. US Patent 5,574,656 (1996).
Agrafiotis, D. K., Bone, R. F., Salemme, F. R. & Soll, R. M. System, method and computer program for at least partially automatically generating chemical compounds having desired properties. US Patent 5,684,711 (1997).
Agrafiotis, D. K., Bone, R. F., Salemme, F. R. & Soll, R. M. System, method and computer program for at least partially automatically generating chemical compounds with desired properties from a list of potential chemical compounds to synthesize. US Patent 5,901,069 (1999).
Pantoliano, M. P. et al. High density miniaturized thermal shift assay as a general strategy for drug discovery. J. Biomol. Screen. 6, 492–440 (2001).This article describes the use of a high-throughput, fluorescence-based method for detecting thermal phase transitions in proteins as a means to evaluate their stability and the effects of bound ligands.
Martin, E. J., Spellmeyer, D. C., Critchlow, R. E. Jr & Blaney, J. M. in Reviews in Computational Chemistry Vol. 10 (eds Lipkowitz, K. B. & Boyd, D. B.) 75–100 (VCH, Weinheim, 1997).
Agrafiotis, D. K. in The Encyclopedia of Computational Chemistry (eds Schleyer, P. V. R. et al.) 742–761 (John Wiley and Sons, Chichester, 1998).
Bures, M. G. & Martin, Y. C. Computational methods in molecular diversity and combinatorial chemistry. Curr. Opin. Chem. Biol. 2, 376–380 (1998).
Agrafiotis, D. K., Myslik, J. C. & Salemme, F. R. Advances in diversity profiling and combinatorial series design. Mol. Divers. 4, 1–22 (1999).An in-depth review of computational methods that are used in diversity analysis and combinatorial-library design.
Drewry, D. H. & Young, S. S. Approaches to the design of combinatorial libraries. Chemometr. Intell. Lab. Syst. 48, 1–20 (1999).
Leach, A. R. & Hann, M. M. The in silico world of virtual libraries. Drug Discov. Today 5, 326–336 (2000).
Leland, B. A. et al. Managing the combinatorial explosion. J. Chem. Inf. Comput. Sci. 37, 62–70 (1997).
Leach, A. R., Bradshaw, J., Green, D. V. S., Hann, M. M. & Delany, J. J. Implementation of a system for reagent selection and library enumeration, profiling & design. J. Chem. Inf. Comput. Sci. 39, 1161–1172 (1999).
Lobanov, V. S. & Agrafiotis, D. K. Scalable methods for the construction and analysis of virtual combinatorial libraries. Combin. Chem. High-Throughput Screen. 5, 167–178 (2002).
Walters, W. P., Stahl, M. T. & Murcko, M. A. Virtual screening — an overview. Drug Discov. Today 3, 160–178 (1998).
Agrafiotis, D. K., Lobanov, V. S., Rassokhin, D. N. & Izrailev, S. in Virtual Screening for Bioactive Molecules (eds Böhm, H.-J. & Schneider, G.) 265–300 (Wiley–VCH, Weinheim, 2000).
Johnson, M. A. & Maggiora, G. M. Concepts and Applications of Molecular Similarity (Wiley, New York, 1990).An authoritative overview of the theoretical and practical aspects of molecular similarity as it applies to chemical and biological research.
Livingston, D. J. The characterization of molecular structures using molecular properties. A survey. J. Chem. Inf. Comput. Sci. 40, 195–209 (2000).
Hall, L. H. & Kier, L. B. in Reviews of Computational Chemistry (eds Boyd, D. B. & Lipkowitz, K. B.) 367–422 (VCH, Weinheim, 1991).Describes a class of important molecular-connectivity indices and their use in predicting molecular properties.
James, C. A., Weininger, D. & Delaney, J. Daylight Theory Manual. Daylight Chemical Information Systems [online] (cited 12 Mar 02) 〈http://www.daylight.com/〉.
Sadowski, J. & Kubinyi, H. A scoring scheme for discriminating between drugs and nondrugs. J. Med. Chem. 41, 3325–3329 (1998).Describes the application of neural networks for discriminating drugs from non-drugs by using simple atom-type descriptors.
Schneider, G., Neidhart, W., Giller, T. & Schmid, G. Scaffold-hopping by topological pharmacophore search: a contribution to virtual screening. Angew. Chem. Int. Edn Engl. 38, 2894–2896 (1999).
Carhart, R. E., Smith, D. H. & Venkataraghavan, R. Atom pairs as molecular features in structure–activity studies: definition and application. J. Chem. Inf. Comput. Sci. 25, 64–73 (1985).
Nilakantan, R., Bauman, N., Dixon, J. S. & Venkataraghavan, R. Topological torsions: a new molecular descriptor for SAR applications. Comparison with other descriptors. J. Chem. Inf. Comput. Sci. 27, 82–85 (1987).
Kearsley, S. K. et al. Chemical similarity using physicochemical property descriptors. J. Chem. Inf. Comput. Sci. 36, 118–127 (1996).
Moreau, G. & Broto, P. The autocorrelation of a topological structure: a new molecular descriptor. Nouv. J. Chim. 4, 359–360 (1980).
Bauknecht, H. et al. Locating biologically active compounds in medium-sized heterogeneous datasets by topological autocorrelation vectors: dopamine and benzodiazepine agonists. J. Chem. Inf. Comput. Sci. 36, 1205–1213 (1996).
Labute, P. A widely applicable set of descriptors. J. Mol. Graph. Model. 18, 464–467 (2000).
Kubinyi, H. in Methods and Principles in Medicinal Chemistry Vol. 1 (eds Manhold, R., Krogsgaard-Larsen, P. & Timmermann, H.) 21–36 (VCH, Weinheim, 1993).
Burden, F. R. Molecular identification number for substructure searches. J. Chem. Inf. Comput. Sci. 29, 225–227 (1989).
Sheridan, R. P., Miller, M. D., Underwood, D. J. & Kearsley, S. K. Chemical similarity using geometric atom pair descriptors. J. Chem. Inf. Comput. Sci. 36, 128–136 (1996).
Wagener, M., Sadowski, J. & Gasteiger, J. Autocorrelation of molecular surface properties for modeling corticosteroid binding globulin and cytosolic Ah receptor activity by neural networks. J. Am. Chem. Soc. 117, 7769–7775 (1995).
Todeschini, R., Lasagni, M. & Marengo, E. New molecular descriptors for 2D and 3D structures. Theory. J. Chemom. 8, 263–272 (1994).
Ghuloum, A. M., Sage, C. R. & Jain, A. N. Molecular hashkeys: a novel method for molecular characterization and its application for predicting important pharmaceutical properties of molecules. J. Med. Chem. 42, 1739–1748 (1999).
Pearlman, R. S. & Smith, K. M. Metric validation and the receptor-relevant subspace concept. J. Chem. Inf. Comput. Sci. 9, 28–35 (1999).
Sheridan, R. P. et al. 3Dsearch; a system for three-dimensional substructure searching. J. Chem. Inf. Comput. Sci. 29, 255–260 (1989).
Murrall, N. W. & Davies, E. K. Conformational freedom in 3-D databases. 1. Techniques. J. Chem. Inf. Comput. Sci. 30, 312–316 (1990).
Guner, O. F. Pharmacophore Perception, Development and Use in Drug Design (International Univ. Line, La Jolla, 2000).A collection of articles that describe the use of pharmacophore modelling in drug design.
Mason, J. S. et al. New 4-point pharmacophore method for molecular similarity and diversity applications: overview of the method and applications, including a novel approach to the design of combinatorial libraries containing priviledged substructures. J. Med. Chem. 42, 3251–3264 (1999).
Leach, A. R., Green, D. V. S., Hann, M. M., Judd, D. B. & Good, A. C. Where are the GaPs? A rational approach to monomer acquisition and selection. J. Chem. Inf. Comput. Sci. 40, 1262–1269 (2000).
Martin, E. J. & Hoeffel, T. J. Oriented substituent pharmacophore property space (OSPPREYS): A substituent-based calculation that describes combinatorial library products better than the corresponding product-based selection. J. Mol. Graph. Model. 18, 383–403 (2000).This paper describes the use of substituent-based pharmacophore descriptors to encode conformation-dependent properties of combinatorial products.
Cramer, R. D., Clark, R. D., Patterson, D. E. & Ferguson, A. M. Bioisosterism as a molecular diversity descriptor: steric fields of single topomeric conformers. J. Med. Chem. 39, 3060–3069 (1996).
Matter, H. & Potter, T. Comparing 3D pharmacophore triplets and 2D fingerprints for selecting diverse compound subsets. J. Chem. Inf. Comput. Sci. 39, 1211–1225 (1999).
Salemme, F. R., Spurlino, J. & Bone, R. Serendipity meets precision: the integration of structure based drug design and combinatorial chemistry for efficient drug discovery. Structure 5, 319–324 (1997).
Graybill, T. L. et al. in Molecular Diversity and Combinatorial Chemistry (eds Chaiken, I. M. & Janda, K. D.) 16–26 (ACS, Washington DC, 1996).
Jones, G., Willett, P., Glen, R. C., Leach, A. R. & Taylor, R. Further development of a genetic algorithm for ligand docking and its application to screening combinatorial libraries. ACS Symp. Ser. 719, 271–291 (1999).
Waszkowycz, B., Perkins, T. D. J., Sykes, R. A. & Li, J. Large-scale virtual screening for discovering leads in the post-genomics era. IBM Syst. J. 40, 360–376 (2001).
Sun, Y., Ewing, T. J. A., Skillman, A. G. & Kuntz, I. D. CombiDock: structure-based combinatorial docking and library design. J. Comput. Aided. Mol. Des. 12, 597–604 (1998).
Waller, C. L. & Bradley, M. P. Development and validation of a novel variable selection technique with application to multidimensional quantitative structure-activity relationship studies. J. Chem. Inf. Comput. Sci. 39, 345–355 (1999).
Rose, V. S. & Wood, J. Generalized cluster significance analysis with conditional probabilities. Quant. Struct. Activ. Rel. 17, 348–356 (1998).
Godden, J. W. & Bajorath, J. Differential Shannon entropy as a sensitive measure of differences in database variability of molecular descriptors. J. Chem. Inf. Comput. Sci. 41, 1060–1066 (2001).
Cooley, W. & Lohnes, P. Multivariate Data Analysis (Wiley, New York, 1971).
Xie, D., Tropsha, A. & Schlick, T. An efficient projection protocol for chemical databases: singular value decomposition combined with truncated Newton minimization. J. Chem. Inf. Comput. Sci. 40, 167–177 (2000).
Hull, R. D. et al. Latent semantic structure indexing (LASSI) for defining chemical similarity. J. Med. Chem. 44, 1177–1184 (2001).
Cummins, D. J., Andrews, C. W., Bentley, J. A. & Cory, M. Molecular diversity in chemical databases: comparison of medicinal chemistry knowledge bases and databases of commercially available compounds. J. Chem. Inf. Comput. Sci. 36, 750–763 (1996).
Kruskal, J. B. Non-metric multidimensional scaling: a numerical method. Phychometrika 29, 115–129 (1964).
Sammon, J. W. A nonlinear mapping for data structure analysis. IEEE Trans. Comput. C18, 401–409 (1969).
Agrafiotis, D. K. & Lobanov, V. S. Nonlinear mapping networks. J. Chem. Inf. Comput. Sci. 40, 1356–1362 (2000).
Rassokhin, D. N., Lobanov, V. S. & Agrafiotis, D. K. Nonlinear mapping of massive data sets by fuzzy clustering and neural networks. J. Comput. Chem. 22, 373–386 (2001).
Agrafiotis, D. K., Rassokhin, D. N. & Lobanov, V. S. Multidimensional scaling and visualization of large molecular similarity tables. J. Comput. Chem. 22, 488–500 (2001).
Agrafiotis, D. K. & Lobanov, V. S. Multidimensional scaling of combinatorial libraries without explicit enumeration. J. Comput. Chem. 22, 1712–1722 (2001).
Jamois, E. A., Hassan, M. & Waldman, M. Evaluation of reagent-based and product-based strategies in the design of combinatorial library subsets. J. Chem. Inf. Comput. Sci. 40, 63–70 (2000).
Agrafiotis, D. K. & Rassokhin, D. N. A fractal approach for selecting an appropriate bin size for cell-based diversity estimation. J. Chem. Inf. Comput. Sci. 42, 117–122 (2002).
Montgomery, D. C. Design and Analysis of Experiments 4th edn (John Wiley and Sons, New York, 1996).
Martin, E. J. et al. Measuring diversity: Experimental design of combinatorial libraries for drug discovery. J. Med. Chem. 38, 1431–1436 (1995).This paper describes the use of statistical experimental-design techniques to select building blocks for combinatorial libraries using a rich set of molecular descriptors.
Hassan, M., Bielawski, J. P., Hempel, J. C. & Waldman, M. Optimization and visualization of molecular diversity of combinatorial libraries. Mol. Divers. 2, 64–74 (1996).
Kennard, R. W. & Stone, L. A. Computer-aided design of experiments. Technometrics 11, 137–148 (1969).
Higgs, R. E., Bemis, K. G., Watson, I. A. & Wikel, J. H. Experimental designs for selecting molecules from large chemical databases. J. Chem. Inf. Comput. Sci. 37, 861–870 (1997).
Snarey, M., Terrett, N. K., Willett, P. & Wilton, D. J. Comparison of algorithms for dissimilarity-based compound selection. J. Mol. Graph. Model. 15, 372–385 (1997).
Mount, J., Ruppert, J., Welch, W. & Jain, A. N. IcePick: a flexible surface-based system for molecular diversity. J. Med. Chem. 42, 60–66 (1999).
Agrafiotis, D. K. & Lobanov, V. S. An efficient implementation of distance-based diversity metrics based on k-d trees. J. Chem. Inf. Comput. Sci. 39, 51–58 (1999).
Agrafiotis, D. K. A constant time algorithm for estimating the diversity of large chemical libraries. J. Chem. Inf. Comput. Sci. 41, 159–167 (2001).
Downs, G. M. & Willett, P. Similarity searching and clustering of chemical-structure databases using molecular property data. J. Chem. Inf. Comput. Sci. 34, 1094–1102 (1994).
Brown, R. D. & Martin, Y. C. Use of structure–activity data to compare structure-based clustering methods and descriptors for use in compound selection. J. Chem. Inf. Comput. Sci. 36, 572–584 (1996).A comparison of several two-dimensional and three-dimensional descriptors, which is based on their ability to discriminate active from inactive compounds.
Brown, R. D. & Martin, Y. C. The information content of 2D and 3D structural descriptors relevant to ligand–receptor binding. J. Chem. Inf. Comput. Sci. 37, 1–9 (1997).
Patterson, D. E., Cramer, R. D., Ferguson, A. M., Clark, R. D. & Weinberger, L. E. Neighborhood behavior: a useful concept for validation of molecular diversity descriptors. J. Med. Chem. 39, 3049–3059 (1996).
Matter, H. Selecting optimally diverse compounds from structure databases: a validation study of two-dimensional and three-dimensional molecular descriptors. J. Med. Chem. 40, 1219–1229 (1997).
Martin, Y. C., Bures, M. G. & Brown, R. D. Validated descriptors for diversity measurements and optimization. Pharm. Pharmacol. Commun. 4, 147–152 (1998).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeny, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 23, 3–25 (1997).A discussion of the importance of ADME screening in early-stage drug discovery.
Oprea, T. I. Property distribution of drug-related chemical databases. J. Comput. Aided Mol. Des. 14, 251–264 (2000).
Muegge, I., Heald, S. L. & Brittelli, D. Simple selection criteria for drug-like chemical matter. J. Med. Chem. 44, 1841–1846 (2001).
Sheridan, R. P. The most common chemical replacements in drug-like compounds. J. Chem. Inf. Comput. Sci. 2, 103–108 (2002).
Wang, J. & Ramnarayan, K. Toward designing drug-like libraries: a novel computational approach for prediction of drug feasibility of compounds. J. Combin. Chem. 1, 524–533 (1999).
Ajay, A., Walters, W. P. & Murcko, M. A. Can we learn to distinguish between drug-like and nondrug-like molecules? J. Med. Chem. 41, 3314–3324 (1998).
Wagener, M. & van Geerestein, V. J. Potential drugs and nondrugs: prediction and identification of important structural features. J. Chem. Inf. Comput. Sci. 40, 280–292 (2000).
Yu, L. X., Lipka, E., Crison, J. R. & Amidon, G. L. Transport approach to the biopharmaceutical design of oral drug delivery systems: prediction of intestinal absorption. Adv. Drug Deliv. Rev. 19, 359–376 (1996).
Teague, S. J., Davis, A. M., Leeson, P. D. & Oprea, T. I. The design of leadlike combinatorial libraries. Angew. Chem. Int. Edn Engl. 38, 3743–3748 (1999).Based on an analysis of 18 lead-drug pairs, the authors point out that traditional medicinal chemistry optimization tends to drive initial high-throughput screening (HTS) hits outside the “rule-of-five” range, and suggest that combinatorial libraries should have lower molecular masses and lower log P profiles than those originally proposed by Lipinski.
Koehler, R. T., Dixon, S. L. & Villar, O. H. LASSOO: a generalized directed diversity approach to the design and enrichment of chemical libraries. J. Med. Chem. 42, 4695–4704 (1999).
Gillet, V. J., Willet, P., Bradshaw, J. & Green, D. V. S. Selecting combinatorial libraries to optimize diversity and physical properties. J. Chem. Inf. Comput. Sci. 39, 169–177 (1999).
Rassokhin, D. N. & Agrafiotis, D. K. Kolmogorov–Smirnov statistic and its applications in library design. J. Mol. Graph. Model. 18, 370–384 (2000).
Brown, R. D., Hassan, M. & Waldman, M. Combinatorial library design for diversity, cost efficiency and drug-like character. J. Mol. Graph. Model. 18, 427–437 (2000).
Shi, S., Peng, Z., Kostrowicki, J., Paderes, J. & Kuki, A. Efficient combinatorial filtering for desired molecular properties of reaction products. J. Mol. Graph. Model. 18, 478–496 (2000).
Martin, E. & Wong, A. Sensitivity analysis and other improvements to tailored combinatorial library design. J. Chem. Inf. Comput. Sci. 40, 215–220 (2000).
Gillet, V. J., Willett, P. & Bradshaw, J. The effectiveness of reactant pools for generating structurally-diverse combinatorial libraries. J. Chem. Inf. Comput. Sci. 37, 731–740 (1997).
Jamois, E. A., Hassan, M. & Waldman, M. Evaluation of reagent-based and product-based strategies in the design of combinatorial library subsets. J. Chem. Inf. Comput. Sci. 40, 63–70 (2000).
Graham, E. T., Jacober, S. P. & Cardoso, M. G. A novel frequency distribution selection method for efficient plate layout of a diverse combinatorial library. J. Chem. Inf. Comput. Sci. 41, 1508–1516 (2001).
Bayada, D. M., Hamersma, H. & van Geerestein, V. J. Molecular diversity and representativity in chemical databases. J. Chem. Inf. Comput. Sci. 39, 1–10 (1999).
Agrafiotis, D. K. & Lobanov, V. S. Ultrafast algorithm for designing focused combinatorial arrays. J. Chem. Inf. Comput. Sci. 40, 1030–1038 (2000).
Stanton, R. V. et al. Combinatorial library design: maximizing model fitting compounds with matrix synthesis constraints. J. Chem. Inf. Comput. Sci. 40, 701–705 (2000).
Agrafiotis, D. K. Stochastic algorithms for maximizing molecular diversity. J. Chem. Inf. Comput. Sci. 37, 841–851 (1997).
Hassan, M., Bielawski, J. P., Hempel, J. C. & Waldman, M. Optimization and visualization of molecular diversity of combinatorial libraries. Mol. Diversity 2, 64–74 (1996).
Good, A. C. & Lewis, R. A. New methodology for profiling combinatorial libraries and screening sets: cleaning up the design process with HARPcik. J. Med. Chem. 40, 3926–3236 (1997).
Zheng, W., Cho, S. J. & Tropsha, A. Rational combinatorial library design: 1) Focus-2D: a new approach to the design of targeted combinatorial chemical libraries. J. Chem. Inf. Comput. Sci. 38, 251–258 (1998).
Waldman, M., Li, H. & Hassan, M. Novel algorithms for the optimization of molecular diversity of combinatorial libraries. J. Mol. Graph. Model. 18, 412–426 (2000).
Agrafiotis, D. K. Multiobjective optimization of combinatorial libraries. IBM J. Res. Develop. 45, 545–566 (2001).
Sheridan, R. P. & Kearsley, S. K. Using a genetic algorithm to suggest combinatorial libraries. J. Chem. Inf. Comput. Sci. 35, 310–3201 (1995).
Weber, L., Wallbaum, S., Broger, C. & Gubernator, K. Optimization of the biological activity of combinatorial compound libraries by a genetic algorithm. Angew. Chem. Int. Edn Engl. 34, 2280–2282 (1995).
Singh, J. et al. Application of genetic algorithms to combinatorial synthesis: a computational approach for lead identification and lead optimization. J. Am. Chem. Soc. 118, 1669–1676 (1996).A description of the use of a genetic algorithm to optimize peptide-based collagenase substrates using direct experimental feedback, without constructing any intermediate models of biological activity.
Brown, R. D. & Martin, Y. C. Designing combinatorial library mixtures using genetic algorithms. J. Med. Chem. 40, 2304–2313 (1997).
Sheridan, R. P., SanFeliciano, S. G. & Kearsley, S. K. Designing targeted libraries with genetic algorithms. J. Mol. Graph. Model. 18, 320–334 (2000).
Farnum, M. & Agrafiotis, D. K. Combinatorial Swarms (CombiChem, London, 2001).
Lobarov, V. S. & Agrafiotis, D. K. Stochastic similarity selections from large combinatorial libraries. J. Chem. Inf. Comput. Sci. 40, 460–470 (2000).
Downs, G. M. & Barnard, J. M. Techniques for generating descriptive fingerprints in combinatorial libraries. J. Chem. Inf. Comput. Sci. 37, 59–61 (1997).
Cramer, R. D., Patterson, D. E., Clark, R. D., Soltanshahi, F. & Lawless, M. S. Virtual compound libraries: a new approach to decision making in molecular discovery research. J. Chem. Inf. Comput. Sci. 38, 1010–1023 (1998).
Ivanciuc, O. & Klein, D. J. Computing Weiner-type indices for virtual combinatorial libraries generated from heteroatom-containing building blocks. J. Chem. Inf. Comput. Sci. 42, 8–22 (2002).
Lobanov, V. S. & Agrafiotis, D. K. Combinatorial networks. J. Mol. Graph. Model. 19, 571–578 (2001).Describes the use of neural networks for predicting properties of combinatorial products from properties of their respective building blocks. This method allows product-based virtual screening of massive combinatorial libraries in a way that circumvents their virtual synthesis.
Author information
Authors and Affiliations
Corresponding author
Related links
Related links
FURTHER INFORMATION
Chemical Informatics Societies and Professional Organizations
LINKS
Glossary
- PROBE LIBRARY
-
A collection of diverse compounds that is aimed at discovering hits across a wide variety of biological targets.
- MULTIOBJECTIVE OPTIMIZATION
-
The solution to a problem that involves the simultaneous optimization of multiple design objectives.
- VIRTUAL LIBRARY
-
A computer representation of a collection of chemical compounds.
- COMBINATORIAL LIBRARY
-
A collection of compounds that are derived from the systematic application of a synthetic sequence on a prescribed set of building blocks.
- ENUMERATION
-
The process of constructing the connection tables of the combinatorial products from their respective building blocks, as prescribed by the reaction sequence.
- CLIPPED REAGENT
-
The (potentially modified) part of a reagent that becomes part of the final product.
- CONNECTION TABLE
-
A computer representation of the atoms and bonds that comprise a molecule. This is the computer equivalent of a chemical sketch of a molecule.
- LAZY ENUMERATION
-
The on-demand virtual synthesis of combinatorial products.
- MOLECULAR PERCEPTION
-
The computational detection of important structural features, such as rings, aromaticity, stereochemistry and topological symmetry, from the molecule's connection table.
- MOLECULAR DIVERSITY
-
The chemical-information content of a collection of compounds. The concept is often context dependent.
- GRAPH THEORY
-
Formally, a connection table for a molecule records its chemical structure as a graph — a set of vertices (the atoms) linked by edges (the bonds). This allows mathematical analyses to be used to classify the structure or calculate molecular properties.
- FINGERPRINT
-
A set of binary numbers (1s and 0s) that are used to characterize a molecular structure. Each bit signifies the presence (1) or absence (0) of one or more structural features in the target molecule.
- PHARMACOPHORE
-
The ensemble of steric and electronic features that are necessary to ensure optimal supramolecular interactions with a specific biological target and to trigger (or block) its biological function. Only molecules that interact at the same receptor site in the same way share a common pharmacophore.
- DRUG LIKENESS
-
The thesis that drugs have certain common properties that differentiate them from other ordinary chemicals.
- LOG P
-
The octanol/water partition coefficient is the ratio of the compound's solubility in octanol to its solubility in water. The logarithm of this partition coefficient is called log P. It provides an estimate of the compound's ability to pass through a cell membrane.
- OUTLIER
-
A point that, because of observation noise, does not follow the characteristics of the input (or desired response) data.
- CURSE OF DIMENSIONALITY
-
The sparsity of data in higher dimensions.
- QUADRATIC COMPLEXITY
-
Quadratic complexity means that if the size of the problem doubles, the computational time that is required by the algorithm to solve it quadruples. The complexity (or order) of an algorithm is an important criterion for comparing algorithms that involve the analysis of large data sets.
- LIPINSKI RULE OF 5
-
For compounds that are not substrates of biological transporters, poor absorption and permeation are more likely to occur when there are more than 5 hydrogen-bond donors, more than 10 hydrogen-bond acceptors, the molecular mass is greater than 500 Da, or the log P is greater than 5.
- BIOISOSTERISM
-
The idea that a chemical group in a biologically active molecule can be replaced by another chemical group without loss of activity.
- COMBINATORIAL OPTIMIZATION
-
The number of different combinations of k objects out of a set of n objects is given by the binomial coefficient Cnk = n!/(n − k)!k!. This can be used to calculate the number of distinct k1×k2×...×kd combinatorial arrays in a n1×n2×...×nd combinatorial library. For example, there are approximately 1040 different 10×10×10 arrays in a 100×100×100 library.
- COMBINATORIAL NEURAL NETWORK
-
(CNN). A neural network that is trained to predict molecular properties of combinatorial products from pertinent features of their respective building blocks.
- FEATURE SELECTION
-
A computational technique that attempts to identify a small subset of features that are most relevant to a particular machine learning task.
Rights and permissions
About this article
Cite this article
Agrafiotis, D., Lobanov, V. & Salemme, F. Combinatorial informatics in the post-genomics era. Nat Rev Drug Discov 1, 337–346 (2002). https://doi.org/10.1038/nrd791
Issue Date:
DOI: https://doi.org/10.1038/nrd791
This article is cited by
-
Natural products in modern life science
Phytochemistry Reviews (2010)
-
Smart drug discovery leveraging innovative technologies and predictive knowledge
Nature Chemical Biology (2006)
-
Molecular similarity and diversity in chemoinformatics: From theory to applications
Molecular Diversity (2006)