Abstract
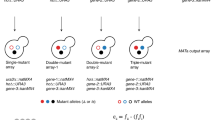
The quantitative analysis of genetic interactions between pairs of gene mutations has proven to be effective for characterizing cellular functions, but it can miss important interactions for functionally redundant genes. To address this limitation, we have developed an approach termed triple-mutant analysis (TMA). The procedure relies on a query strain that contains two deletions in a pair of redundant or otherwise related genes, which is crossed against a panel of candidate deletion strains to isolate triple mutants and measure their growth. A central feature of TMA is to interrogate mutants that are synthetically sick when two other genes are deleted but interact minimally with either single deletion. This approach has been valuable for discovering genes that restore critical functions when the principal actors are deleted. TMA has also uncovered double-mutant combinations that produce severe defects because a third protein becomes deregulated and acts in a deleterious fashion, and it has revealed functional differences between proteins presumed to act together. The protocol is optimized for Singer ROTOR pinning robots, takes 3 weeks to complete and measures interactions for up to 30 double mutants against a library of 1,536 single mutants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Tong, A.H. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Collins, S.R., Roguev, A. & Krogan, N.J. Quantitative genetic interaction mapping using the E-MAP approach. Methods Enzymol. 470, 205–231 (2010).
Schuldiner, M. et al. Exploration of the function and organization of the yeast early secretory pathway through an epistatic miniarray profile. Cell 123, 507–519 (2005).
Butland, G. et al. eSGA: E. coli. synthetic genetic array analysis. Nat. Methods 5, 789–795 (2008).
Typas, A. et al. High-throughput, quantitative analyses of genetic interactions in E. coli. Nat. Methods 5, 781–787 (2008).
Bassik, M.C. et al. A systematic mammalian genetic interaction map reveals pathways underlying ricin susceptibility. Cell 152, 909–922 (2013).
Roguev, A. et al. Quantitative genetic-interaction mapping in mammalian cells. Nat. Methods 10, 432–437 (2013).
Roguev, A., Wiren, M., Weissman, J.S. & Krogan, N.J. High-throughput genetic interaction mapping in the fission yeast Schizosaccharomyces pombe. Nat. Methods 4, 861–866 (2007).
Ryan, C.J. et al. Hierarchical modularity and the evolution of genetic interactomes across species. Mol. Cell 46, 691–704 (2012).
Pan, X. et al. A robust toolkit for functional profiling of the yeast genome. Mol. Cell 16, 487–496 (2004).
Pan, X. et al. A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell 124, 1069–1081 (2006).
Aguilar, P.S. et al. A plasma-membrane E-MAP reveals links of the eisosome with sphingolipid metabolism and endosomal trafficking. Nat. Struct. Mol. Biol. 17, 901–908 (2010).
Collins, S.R. et al. Functional dissection of protein complexes involved in yeast chromosome biology using a genetic interaction map. Nature 446, 806–810 (2007).
Fiedler, D. et al. Functional organization of the S. cerevisiae phosphorylation network. Cell 136, 952–963 (2009).
Wilmes, G.M. et al. A genetic interaction map of RNA-processing factors reveals links between Sem1/Dss1-containing complexes and mRNA export and splicing. Mol. Cell 32, 735–746 (2008).
Schuldiner, M., Collins, S.R., Weissman, J.S. & Krogan, N.J. Quantitative genetic analysis in Saccharomyces cerevisiae using epistatic miniarray profiles (E-MAPs) and its application to chromatin functions. Methods 40, 344–352 (2006).
Braberg, H. et al. From structure to systems: high-resolution, quantitative genetic analysis of RNA polymerase II. Cell 154, 775–788 (2013).
Surma, M.A. et al. A lipid E-MAP identifies Ubx2 as a critical regulator of lipid saturation and lipid bilayer stress. Mol. Cell 51, 519–530 (2013).
Beltrao, P., Cagney, G. & Krogan, N.J. Quantitative genetic interactions reveal biological modularity. Cell 141, 739–745 (2010).
Donaldson, A.D. The yeast mitotic cyclin Clb2 cannot substitute for S-phase cyclins in replication origin firing. EMBO Rep. 1, 507–512 (2000).
Haber, J.E. et al. Systematic triple-mutant analysis uncovers functional connectivity between pathways involved in chromosome regulation. Cell Rep. 3, 2168–2178 (2013).
Mai, B. & Breeden, L. CLN1 and its repression by Xbp1 are important for efficient sporulation in budding yeast. Mol. Cell Biol. 20, 478–487 (2000).
Segal, M., Clarke, D.J. & Reed, S.I. Clb5-associated kinase activity is required early in the spindle pathway for correct preanaphase nuclear positioning in Saccharomyces cerevisiae. J. Cell Biol. 143, 135–145 (1998).
Hsu, W.S. et al. S-phase cyclin-dependent kinases promote sister chromatid cohesion in budding yeast. Mol. Cell Biol. 31, 2470–2483 (2011).
Ben-Aroya, S., Koren, A., Liefshitz, B., Steinlauf, R. & Kupiec, M. ELG1, a yeast gene required for genome stability, forms a complex related to replication factor C. Proc. Natl. Acad. Sci. USA 100, 9906–9911 (2003).
Green, C.M., Erdjument-Bromage, H., Tempst, P. & Lowndes, N.F. A novel Rad24 checkpoint protein complex closely related to replication factor C. Curr. Biology 10, 39–42 (2000).
Hanna, J.S., Kroll, E.S., Lundblad, V. & Spencer, F.A. Saccharomyces cerevisiae CTF18 and CTF4 are required for sister chromatid cohesion. Mol. Cell Biol. 21, 3144–3158 (2001).
Mayer, M.L., Gygi, S.P., Aebersold, R. & Hieter, P. Identification of RFC(Ctf18p, Ctf8p, Dcc1p): an alternative RFC complex required for sister chromatid cohesion in S. cerevisiae. Mol. Cell 7, 959–970 (2001).
Naiki, T., Kondo, T., Nakada, D., Matsumoto, K. & Sugimoto, K. Chl12 (Ctf18) forms a novel replication factor C-related complex and functions redundantly with Rad24 in the DNA replication checkpoint pathway. Mol. Cell Biol. 21, 5838–5845 (2001).
Roelofs, J. et al. Chaperone-mediated pathway of proteasome regulatory particle assembly. Nature 459, 861–865 (2009).
De Koning, L., Corpet, A., Haber, J.E. & Almouzni, G. Histone chaperones: an escort network regulating histone traffic. Nat. Struct. Mol. Biol. 14, 997–1007 (2007).
Enomoto, S. & Berman, J. Chromatin assembly factor I contributes to the maintenance, but not the re-establishment, of silencing at the yeast silent mating loci. Genes Dev. 12, 219–232 (1998).
Ramey, C.J. et al. Activation of the DNA damage checkpoint in yeast lacking the histone chaperone anti-silencing function 1. Mol. Cell Biol. 24, 10313–10327 (2004).
Sugawara, N., Wang, X. & Haber, J.E. In vivo roles of Rad52, Rad54, and Rad55 proteins in Rad51-mediated recombination. Mol. Cell 12, 209–219 (2003).
Driscoll, R., Hudson, A. & Jackson, S.P. Yeast Rtt109 promotes genome stability by acetylating histone H3 on lysine 56. Science 315, 649–652 (2007).
Han, J., Zhou, H., Li, Z., Xu, R.M. & Zhang, Z. Acetylation of lysine 56 of histone H3 catalyzed by RTT109 and regulated by ASF1 is required for replisome integrity. J. Biol. Chem. 282, 28587–28596 (2007).
Blanco, M.G., Matos, J., Rass, U., Ip, S.C. & West, S.C. Functional overlap between the structure-specific nucleases Yen1 and Mus81-Mms4 for DNA-damage repair in S. cerevisiae. DNA Repair 9, 394–402 (2010).
Ho, C.K., Mazon, G., Lam, A.F. & Symington, L.S. Mus81 and Yen1 promote reciprocal exchange during mitotic recombination to maintain genome integrity in budding yeast. Mol. Cell 40, 988–1000 (2010).
Tay, Y.D. & Wu, L. Overlapping roles for Yen1 and Mus81 in cellular Holliday junction processing. J. Biol. Chem. 285, 11427–11432 (2010).
Zhao, X., Muller, E.G. & Rothstein, R. A suppressor of two essential checkpoint genes identifies a novel protein that negatively affects dNTP pools. Mol. Cell 2, 329–340 (1998).
Koranda, M., Schleiffer, A., Endler, L. & Ammerer, G. Forkhead-like transcription factors recruit Ndd1 to the chromatin of G2/M-specific promoters. Nature 406, 94–98 (2000).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Boulton, S.J. & Jackson, S.P. Identification of a Saccharomyces cerevisiae Ku80 homologue: roles in DNA double strand break rejoining and in telomeric maintenance. Nucleic Acids Res. 24, 4639–4648 (1996).
Lee, S.E., Paques, F., Sylvan, J. & Haber, J.E. Role of yeast SIR genes and mating type in directing DNA double-strand breaks to homologous and non-homologous repair paths. Curr. Biol. 9, 767–770 (1999).
Moore, J.K. & Haber, J.E. Cell cycle and genetic requirements of two pathways of nonhomologous end-joining repair of double-strand breaks in Saccharomyces cerevisiae. Mol. Cell Biol. 16, 2164–2173 (1996).
Ma, J.L., Kim, E.M., Haber, J.E. & Lee, S.E. Yeast Mre11 and Rad1 proteins define a Ku-independent mechanism to repair double-strand breaks lacking overlapping end sequences. Mol. Cell Biol. 23, 8820–8828 (2003).
Laufer, C., Fischer, B., Billmann, M., Huber, W. & Boutros, M. Mapping genetic interactions in human cancer cells with RNAi and multiparametric phenotyping. Nat. Methods 10, 427–431 (2013).
Deriano, L. & Roth, D.B. Modernizing the nonhomologous end-joining repertoire: alternative and classical NHEJ share the stage. Annu. Rev. Genet. 47, 433–455 (2013).
Collins, S.R., Schuldiner, M., Krogan, N.J. & Weissman, J.S. A strategy for extracting and analyzing large-scale quantitative epistatic interaction data. Genome Biol. 7, R63 (2006).
Acknowledgements
We thank S. Bohn for helpful discussion and comments. This work was supported by grants from the US National Institutes of Health (GM084448, GM084279, GM081879 and GM098101 to N.J.K., and GM61766, GM76020 and GM20056 to J.E.H.). N.J.K. is a Searle Scholar and a Keck Young Investigator.
Author information
Authors and Affiliations
Contributions
H.B., R.A., J.E.H. and N.J.K. designed the procedure; and H.B., R.A., M.S., J.X., K.E.F.-S., Q.W., J.E.H. and N.J.K. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Braberg, H., Alexander, R., Shales, M. et al. Quantitative analysis of triple-mutant genetic interactions. Nat Protoc 9, 1867–1881 (2014). https://doi.org/10.1038/nprot.2014.127
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.127
This article is cited by
-
Genetic interactions and functional analyses of the fission yeast gsk3 and amk2 single and double mutants defective in TORC1-dependent processes
Scientific Reports (2017)
-
Suppressive drug combinations and their potential to combat antibiotic resistance
The Journal of Antibiotics (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.