Abstract
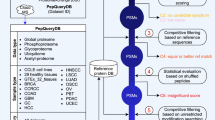
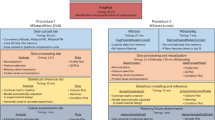
MaxQuant is a quantitative proteomics software package designed for analyzing large mass spectrometric data sets. It is specifically aimed at high-resolution mass spectrometry (MS) data. Currently, Thermo LTQ-Orbitrap and LTQ-FT-ICR instruments are supported and Mascot is used as a search engine. This protocol explains step by step how to use MaxQuant on stable isotope labeling by amino acids in cell culture (SILAC) data obtained with double or triple labeling. Complex experimental designs, such as time series and drug-response data, are supported. A standard desktop computer is sufficient to fulfill the computational requirements. The workflow has been stress tested with more than 1,000 liquid chromatography/mass spectrometry runs in a single project. In a typical SILAC proteome experiment, hundreds of thousands of peptides and thousands of proteins are automatically and reliably quantified. Additional information for identified proteins, such as Gene Ontology, domain composition and pathway membership, is provided in the output tables ready for further bioinformatics analysis. The software is freely available at the MaxQuant home page.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Aebersold, R. & Mann, M. Mass spectrometry-based proteomics. Nature 422, 198–207 (2003).
Zubarev, R. & Mann, M. On the proper use of mass accuracy in proteomics. Mol. Cell. Proteomics 6 377–381 (2007).
Mann, M., & Kelleher, N.L. Special feature: Precision proteomics: the case for high resolution and high mass accuracy. Proc. Natl Acad. Sci. USA 105 18132–18138 (2008).
Makarov, A. et al.Performance evaluation of a hybrid linear ion trap/orbitrap mass spectrometer. Anal. Chem. 78 2113–2120 (2006).
Olsen, J.V. et al. Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics 4, 2010–2021 (2005).
Cox, J., & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Ong, S.E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell. Proteomics 1, 376–386 (2002).
Mann, M. Functional and quantitative proteomics using SILAC. Nat. Rev. Mol. Cell Biol. 7 952–958 (2006).
de Godoy, L.M. et al. Comprehensive mass-spectrometry-based proteome quantification of haploid versus diploid yeast. Nature 455, 1251–1254 (2008).
Bonaldi, T. et al. Combined use of RNAi and quantitative proteomics to study gene function in Drosophila . Mol. Cell 31, 762–772 (2008).
Selbach, M. et al. Widespread changes in protein synthesis induced by microRNAs. Nature 455, 58–63 (2008).
Graumann, J. et al. Stable isotope labeling by amino acids in cell culture (SILAC) and proteome quantitation of mouse embryonic stem cells to a depth of 5,111 proteins. Mol. Cell. Proteomics 7, 672–683 (2008).
Zanivan, S. et al. Solid tumor proteome and phosphoproteome analysis by high resolution mass spectrometry. J. Proteome Res. 7, 5314–5326 (2008).
Cox, J. & Mann, M. Is proteomics the new genomics? Cell 130, 395–398 (2007).
Schimmel, J. et al. The ubiquitin-proteasome system is a key component of the SUMO-2/3 cycle. Mol. Cell. Proteomics 7, 2107–2122 (2008).
Pan, C., Kumar, C., Bohl, S., Klingmueller, U. & Mann, M. Comparative proteomic phenotyping of cell lines and primary cells to assess preservation of cell type- specific functions. Mol. Cell. Proteomics 8, 443–450 (2009).
Hubner, NC., Ren, S., & Mann, M. Peptide separation with immobilized pI strips is an attractive alternative to in-gel protein digestion for proteome analysis. Proteomics 8, 4862–4872 (2008).
Blagoev, B., Ong, S.E., Kratchmarova, I. & Mann, M. Temporal analysis of phosphotyrosine-dependent signaling networks by quantitative proteomics. Nat. Biotechnol. 22, 1139–1145 (2004).
Vermeulen, M., Hubner, N.C. & Mann, M. High confidence determination of specific protein-protein interactions using quantitative mass spectrometry. Curr. Opin. Biotechnol. 19, 331–337 (2008).
Perkins, D.N., Pappin, D.J., Creasy, D.M. & Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Kersey, P.J. et al. The International Protein Index: an integrated database for proteomics experiments. Proteomics 4, 1985–1988 (2004).
UniProt Consortium. The universal protein resource (UniProt). Nucleic Acids Res. 36, D190–D195 (2008).
Flicek, P. et al. Ensembl 2008. Nucleic Acids Res. 36, D707–D714 (2008).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Pevzner, P.A., Mulyukov, Z., Dancik, V. & Tang, C.L. Efficiency of database search for identification of mutated and modified proteins via mass spectrometry. Genome Res. 11, 290–299 (2001).
Cox, J., Hubner, N.C. & Mann, M. How much peptide sequence information is contained in ion trap tandem mass spectra? J. Am. Soc. Mass Spectrom. 19, 1813–1820 (2008).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Finn, R.D. et al. The Pfam protein families database. Nucleic Acids Res. 36, D281–D288 (2008).
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 36, D480–D484 (2008).
Acknowledgements
We thank all the other members of the Proteomics and Signal Transduction group for help with the development of MaxQuant. This work was supported by the Max-Planck Society and by the 6th Framework Program of the European Union (Interaction Proteome Grant LSHG-CT-2003-505520 and HEROIC Grant LSHG-CT-2005-018883).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Cox, J., Matic, I., Hilger, M. et al. A practical guide to the MaxQuant computational platform for SILAC-based quantitative proteomics. Nat Protoc 4, 698–705 (2009). https://doi.org/10.1038/nprot.2009.36
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2009.36
This article is cited by
-
Melatonin decreases GSDME mediated mesothelial cell pyroptosis and prevents peritoneal fibrosis and ultrafiltration failure
Science China Life Sciences (2024)
-
SeqWiz: a modularized toolkit for next-generation protein sequence database management and analysis
BMC Bioinformatics (2023)
-
Identification and partial characterization of new cell density-dependent nucleocytoplasmic shuttling proteins and open chromatin
Scientific Reports (2023)
-
A proteomics analysis of 5xFAD mouse brain regions reveals the lysosome-associated protein Arl8b as a candidate biomarker for Alzheimer’s disease
Genome Medicine (2023)
-
Proteotoxicity caused by perturbed protein complexes underlies hybrid incompatibility in yeast
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.