Abstract
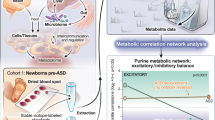
We here describe the MetaNetwork protocol to reconstruct metabolic networks using metabolite abundance data from segregating populations. MetaNetwork maps metabolite quantitative trait loci (mQTLs) underlying variation in metabolite abundance in individuals of a segregating population using a two-part model to account for the often observed spike in the distribution of metabolite abundance data. MetaNetwork predicts and visualizes potential associations between metabolites using correlations of mQTL profiles, rather than of abundance profiles. Simulation and permutation procedures are used to assess statistical significance. Analysis of about 20 metabolite mass peaks from a mass spectrometer takes a few minutes on a desktop computer. Analysis of 2,000 mass peaks will take up to 4 days. In addition, MetaNetwork is able to integrate high-throughput data from subsequent metabolomics, transcriptomics and proteomics experiments in conjunction with traditional phenotypic data. This way MetaNetwork will contribute to a better integration of such data into systems biology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Wink, M. Plant breeding: importance of plant secondary metabolites for protection against pathogens and herbivores. Theor. Appl. Genet. 75, 225–233 (1988).
Baxter, J.D. & Webb, P. Metabolism: bile acids heat things up. Nature 439, 402–403 (2006).
Keurentjes, J.J.B. et al. The genetics of plant metabolism. Nat. Genet. 38, 842–849 (2006).
Jansen, R.C. & Nap, J.P. Genetical genomics: the added value from segregation. Trends Genet. 17, 388–391 (2001).
Brem, R.B., Yvert, G., Clinton, R. & Kruglyak, L. Genetic dissection of transcriptional regulation in budding yeast. Science 296, 752–755 (2002).
Bystrykh, L. et al. Uncovering regulatory pathways that affect hematopoietic stem cell function using “genetical genomics”. Nat. Genet. 37, 225–232 (2005).
Chesler, E.J. et al. Complex trait analysis of gene expression uncovers polygenic and pleiotropic networks that modulate nervous system function. Nat. Genet. 37, 233–242 (2005).
Cheung, V.G. et al. Mapping determinants of human gene expression by regional and genome-wide association. Nature 437, 1365–1369 (2005).
DeCook, R., Lall, S., Nettleton, D. & Howell, S.H. Genetic regulation of gene expression during shoot development in Arabidopsis . Genetics 172, 1155–1164 (2006).
Hubner, N. et al. Integrated transcriptional profiling and linkage analysis for identification of genes underlying disease. Nat. Genet. 37, 243–253 (2005).
Keurentjes, J.J.B. et al. Regulatory network construction in Arabidopsis using genome-wide expression QTLs. Proc. Natl. Acad. Sci. USA 104, 1708–1713 (2007).
Morley, M. et al. Genetic analysis of genome-wide variation in human gene expression. Nature 430, 743–747 (2004).
Schadt, E.E. et al. Genetics of gene expression surveyed in maize, mouse and man. Nature 422, 297–302 (2003).
Yvert, G. et al. Trans-acting regulatory variation in Saccharomyces cerevisiae and the role of transcription factors. Nat. Genet. 35, 57–64 (2003).
Bing, N. & Hoeschele, I. Genetical genomics analysis of a yeast segregant population for transcription network inference. Genetics 170, 533–542 (2005).
Lan, H. et al. Combined expression trait correlations and expression quantitative trait locus mapping. PLoS. Genet. 2, e6 (2006).
Zhu, J. et al. An integrative genomics approach to the reconstruction of gene networks in segregating populations. Cytogenet. Genome Res. 105, 363–374 (2004).
Broman, K.W. Mapping quantitative trait loci in the case of a spike in the phenotype distribution. Genetics 163, 1169–1175 (2003).
Sabatti, C., Service, S. & Freimer, N. False discovery rate in linkage and association genome screens for complex disorders. Genetics 164, 829–833 (2003).
Storey, J.D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Broman, K.W. Mapping expression in randomized rodent genomes. Nat. Genet. 37, 209–210 (2005).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Zhang, P. et al. MetaCyc and AraCyc. Metabolic pathway databases for plant research. Plant Physiol. 138, 27–37 (2005).
Mueller, L.A., Zhang, P. & Rhee, S.Y. AraCyc: a biochemical pathway database for Arabidopsis . Plant Physiol. 132, 453–460 (2003).
Dijkstra, M., Vonk, R.J. & Jansen, R.C. SELDI-TOF mass spectra: a view on sources of variation. J. Chromatogr. B 847, 12–23 (2007).
Tikunov, Y. et al. A novel approach for nontargeted data analysis for metabolomics. Large-scale profiling of tomato fruit volatiles. Plant Physiol. 139, 1125–1137 (2005).
de Vos, C.H. et al. Untargeted large-scale plant metabolomics using liquid chromatography coupled to mass spectrometry. Nat. Protoc. DOI 10.1038/nprot.2007.95 (2007).
Lisec, J., Schauer, N., Kopka, J., Willmitzer, L. & Fernie, A.R. Gas chromatography mass spectrometry-based metabolite profiling in plants. Nat. Protoc. 1, 387–396 (2006).
Kliebenstein, D.J., Gershenzon, J. & Mitchell-Olds, T. Comparative quantitative trait loci mapping of aliphatic, indolic and benzylic glucosinolate production in Arabidopsis thaliana leaves and seeds. Genetics 159, 359–370 (2001).
Hoheisel, J.D. Microarray technology: beyond transcript profiling and genotype analysis. Nat. Rev. Genet. 7, 200–210 (2006).
Ihaka, R. & Gentleman, R. R: a language for data analysis and graphics. J. Comput. Graph. Stat. 5, 299–314 (1996).
Butte, A.J. & Kohane, I.S. Mutual information relevance networks: functional genomic clustering using pairwise entropy measurements. Pac. Symp. Biocomput. 5, 418–429 (2000).
Jansen, R.C. Interval mapping of multiple quantitative trait loci. Genetics 135, 205–211 (1993).
Jansen, R.C. Quantitative trait loci in inbred lines. in Handbook of Statistical Genetics (eds. Balding, D.J., Bishop, M. & Cannings, C.) 445–476 (John Wiley & Sons, New York, 2003).
Swertz, M.A. & Jansen, R.C. Beyond standardization: dynamics software infrastructures for systems biology. Nat. Rev. Genet. 8, 235–243 (2007).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Kliebenstein, D.J., Lambrix, V.M., Reichelt, M., Gershenzon, J. & Mitchell-Olds, T. Gene duplication in the diversification of secondary metabolism: tandem 2-oxoglutarate-dependent dioxygenases control glucosinolate biosynthesis in Arabidopsis . Plant Cell 13, 681–693 (2001).
Kliebenstein, D.J. et al. Genetic control of natural variation in Arabidopsis glucosinolate accumulation. Plant Physiol. 126, 811–825 (2001).
Kroymann, J. et al. A gene controlling variation in Arabidopsis glucosinolate composition is part of the methionine chain elongation pathway. Plant Physiol. 127, 1077–1088 (2001).
de la Fuente, A., Bing, N., Hoeschele, I. & Mendes, P. Discovery of meaningful associations in genomic data using partial correlation coefficients. Bioinformatics 20, 3565–3574 (2004).
Acknowledgements
We thank Dr. Jan-Peter Nap for constructive comments on an earlier version of this paper, Bruno Tesson, Gonzalo Vera and Richard Scheltema for helping to develop the R-package, and Martijn Dijkstra and Rainer Breitling for helping to predict multiple peaks belonging to the same metabolite. This work was supported by grants from the Netherlands Organization for Scientific Research Program Genomics (050-10-029).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Manual
Package 'MetaNetwork' (PDF 143 kb)
Rights and permissions
About this article
Cite this article
Fu, J., Swertz, M., Keurentjes, J. et al. MetaNetwork: a computational protocol for the genetic study of metabolic networks. Nat Protoc 2, 685–694 (2007). https://doi.org/10.1038/nprot.2007.96
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.96
This article is cited by
-
Integration of multi-omics data for prediction of phenotypic traits using random forest
BMC Bioinformatics (2016)
-
Cross-platform comparative analyses of genetic variation in amino acid content in potato tubers
Metabolomics (2014)
-
Organ specificity and transcriptional control of metabolic routes revealed by expression QTL profiling of source-sink tissues in a segregating potato population
BMC Plant Biology (2012)
-
XGAP: a uniform and extensible data model and software platform for genotype and phenotype experiments
Genome Biology (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.