Abstract
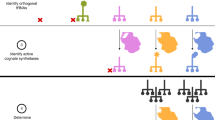
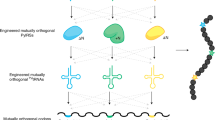
The genetic code of living organisms has been expanded to allow the site-specific incorporation of unnatural amino acids into proteins in response to the amber stop codon UAG. Numerous amino acids have been incorporated including photo-crosslinkers, chemical handles, heavy atoms and post-translational modifications, and this has created new methods for studying biology and developing protein therapeutics and other biotechnological applications. Here we describe a protocol for reprogramming the amino-acid substrate specificity of aminoacyl-tRNA synthetase enzymes that are orthogonal in eukaryotic cells. The resulting aminoacyl-tRNA synthetases aminoacylate an amber suppressor tRNA with a desired unnatural amino acid, but no natural amino acids, in eukaryotic cells. To achieve this change of enzyme specificity, a library of orthogonal aminoacyl-tRNA synthetase is generated and genetic selections are performed on the library in Saccharomyces cerevisiae. The entire protocol, including characterization of the evolved aminoacyl-tRNA synthetase in S. cerevisiae, can be completed in approximately 1 month.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rodriguez, E.A., Lester, H.A. & Dougherty, D.A. In vivo incorporation of multiple unnatural amino acids through nonsense and frameshift suppression. Proc. Natl. Acad. Sci. USA 103, 8650–8655 (2006).
Lummis, S.C. et al. Cis–trans isomerization at a proline opens the pore of a neurotransmitter-gated ion channel. Nature 438, 248–252 (2005).
Monahan, S.L., Lester, H.A. & Dougherty, D.A. Site-specific incorporation of unnatural amino acids into receptors expressed in mammalian cells. Chem. Biol. 10, 573–580 (2003).
Chin, J.W. et al. An expanded eukaryotic genetic code. Science 301, 964–967 (2003).
Cropp, T.A. & Schultz, P.G. An expanding genetic code. Trends Genet. 20, 625–630 (2004).
Wu, N., Deiters, A., Cropp, T.A., King, D. & Schultz, P.G. A genetically encoded photocaged amino acid. J. Am. Chem. Soc. 126, 14306–14307 (2004).
Liu, W., Brock, A., Chen, S., Chen, S. & Schultz, P.G. Genetic incorporation of unnatural amino acids into proteins in mammalian cells. Nat. Methods 4, 239–244 (2007).
Sakamoto, K. et al. Site-specific incorporation of an unnatural amino acid into proteins in mammalian cells. Nucleic Acids Res. 30, 4692–4699 (2002).
Chin, J.W., Cropp, T.A., Chu, S., Meggers, E. & Schultz, P.G. Progress toward an expanded eukaryotic genetic code. Chem. Biol. 10, 511–519 (2003).
Kobayashi, T. et al. Structural basis of nonnatural amino acid recognition by an engineered aminoacyl-tRNA synthetase for genetic code expansion. Proc. Natl. Acad. Sci. USA 102, 1366–1371 (2005).
Kobayashi, T. et al. Structural snapshots of the KMSKS loop rearrangement for amino acid activation by bacterial tyrosyl-tRNA synthetase. J. Mol. Biol. 346, 105–117 (2005).
Stemmer, W.P. & Morris, S.K. Enzymatic inverse PCR: a restriction site independent, single-fragment method for high-efficiency, site-directed mutagenesis. Biotechniques 13, 214–220 (1992).
Summerer, D. et al. A genetically encoded fluorescent amino acid. Proc. Natl. Acad. Sci. USA 103, 9785–9789 (2006).
Shevchenko, A., Wilm, M., Vorm, O. & Mann, M. Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal. Chem. 68, 850–858 (1996).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J.V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Acknowledgements
T.A.C. is grateful to the University of Maryland for financial support. J.W.C. is an EMBO Young Investigator and is supported by the Medical Research Council. We are grateful to P.G.S. for supporting this work.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Cropp, T., Anderson, J. & Chin, J. Reprogramming the amino-acid substrate specificity of orthogonal aminoacyl-tRNA synthetases to expand the genetic code of eukaryotic cells. Nat Protoc 2, 2590–2600 (2007). https://doi.org/10.1038/nprot.2007.378
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.378
This article is cited by
-
Expanding the limits of the second genetic code with ribozymes
Nature Communications (2019)
-
Flexizymes for genetic code reprogramming
Nature Protocols (2011)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.