Abstract
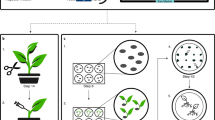
Agrobacterium-mediated transformation of rice is an important method that has been widely adopted by many laboratories. However, because current approaches rely on culture systems, routine protocols have been established only in japonica rice, especially those varieties with higher regeneration potential. Some very efficient methods have been developed for japonica varieties that enable high-throughput functional analysis in rice; however, many elite japonica, and most indica, varieties are difficult to regenerate, leading to low transformation efficiencies. Much effort has been devoted to improving transformation efficiency for all rice genotypes. Here, we describe an Agrobacterium-mediated rice transformation method that is applicable to easily cultured varieties in addition to elite japonica varieties that are more difficult to culture. Using this method, transgenic rice plants can be obtained in about 2–3 months with a transformation frequency of 30–50%, both in easily cultured varieties and recalcitrant elite japonica rice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hiei, Y., Ohta, S., Komari, T. & Kumashiro, T. Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J. 6, 271–282 (1994).
Hiei, Y., Komari, T. & Kubo, T. Transformation of rice mediated by Agrobacterium tumefaciens. Plant Mol. Biol. 35, 205–218 (1997).
Toki, S. Rapid and efficient Agrobacterium-mediated transformation in rice. Plant Mol. Biol. Rep. 15, 16–21 (1997).
Cho, S.K. et al. Efficient transformation of korean rice cultivars (Oryza sativa L.) mediated by Agrobacterium tumefaciens. J. Plant Biol. 41, 262–268 (1998).
Yara, A. et al. Production of transgenic japonica rice (Oryza sativa) cultivar, Taichung 65, by the Agrobacterium-mediated method. Plant Biotech. 18, 305–310 (2001).
Rashid, H., Yokoi, S., Toriyama, K. & Hinata, K. Transgenic plant production mediated by Agrobacterium in indica rice. Plant Cell Rep. 15, 727–730 (1996).
Aldemita, R.R. & Hodges, T.K. Agrobacterium tumefaciens-mediated tansformation of japonica and indica rice varieties. Planta 199, 612–617 (1996).
Zhang, J., Xu, R., Elliott, M.C. & Chen, D.-F. Agrobacterium-mediated transformation of elite indica and japonica rice cultivars. Mol. Biotech. 8, 223–231 (1997).
Dong, J., Teng, W., Buchholz, W.G. & Hall, T.C. Agrobacterium-mediated transformation of javanica rice. Mol. Breed. 2, 267–276 (1996).
Rachmawati, D., Hosaka, T., Inoue, E. & Anzai, H. Agrobacterium-mediated transformation of Javanica rice cv. Rojolele. Biosci. Biotechnol. Biochem., 68, 1193–1200 (2004).
Jeon, J.S. et al. T-DNA insertion mutagenesis for functional genomics in rice. Plant J. 22, 561–570 (2000).
Jeong, D.H. et al. T-DNA insertional mutagenesis for activation tagging in rice. Plant Physiol. 130, 1636–1644 (2002).
Wu, C.Y. et al. Development of enhancer trap lines for functional analysis of the rice genome. Plant J. 35, 418–427 (2003).
Sha, Y. et al. Generation and flanking sequence analysis of a rice T-DNA tagged population. Theor. Appl. Genet., 108, 306–314 (2004).
Sallaud, C. et al. Highly efficient production and characterization of T-DNA plants for trice (Oryza sativa L.) functional genomics. Theor. Appl. Genot. 106, 1396–1408 (2003).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Chu, C.-C., Wang, C.-C. & Sun, C.-S. Establishment of an efficient medium for anther culture of rice through comparative experiments on the nitrogen sources. Sci. Sin. 18, 659–668 (1975).
Nishimura, A. et al. Isolation of a rice regeneration QTL gene and its application to new transformation system. Proc. Natl. Acad. Sci. USA 102, 11940–11944 (2005).
Hoa, T.T.C. et al. Golden indica and japonica rice lines amenable to deregulation. Plant Physiol. 133, 161–169 (2003).
Kumar, K.K. et al. An improved Agrobacterium-mediated transformation protocol for recalcitrant elite indica rice cultivars. Plant Mol. Biol. Rep. 23, 67–73 (2005).
Ignacimuthu, S. & Arockiasamy, S. Agrobacterium-mediated transformation of an elite indica rice for insect resistance. Curr. Sci. 90, 829–835 (2006).
Supartana, P. et al. Development of simple and efficient in planta transformation method for rice (Oryza sativa L.) using Agrobacterium tumefaciens. J. Biosci. Bioeng. 100, 391–397 (2005).
Sentoku, N., Sato, Y. & Matsuoka, M. Overexpression of rice OSH genes induces ectopic shoots on leaf sheaths of transgenic rice plants. Dev. Biol. 220, 358–364 (2000).
Nishimura, A. et al. OsPNH1 regulates leaf development and maintenance of the shoot apical meristem in rice. Plant J. 30, 189–201 (2002).
Ashikari, M. et al. Cytokinin oxidase regulates rice grain production. Science 309, 741–745 (2005).
Hood, E.E., Helmer, G.L., Fraley, R.T. & Chilton, M.D. The hypervirulence of Agrobacterium tumefaciens A281 is encoded in a region of pTiBo542 outside of T-DNA. J. Bacteriol. 168, 1291–1301 (1986).
Hood, E.E., Gelvin, S.B., Melches, L.S. & Hoekema, A. New Agrobacterium helper plasmids for gene transfer to plants. Transgenic Res. 2, 208–218 (1993).
Ohta, S., Mita, S., Hattori, T. & Nakamura, K. Construction and expression in tobacco of a β glucuronidase (GUS) reporter gene containing an intron within the coding sequence. Plant Cell Physiol. 31, 805–813 (1990).
Zhang, W., McElroy, D. & Wu, R. Analysis of rice Act1 5′ region activity in transgenic rice plants. Plant Cell 3, 1155–1165 (1991).
Joy, K.W. & Hageman, R.H. The purification and properties of nitrite reductase from higher plants, and its dependence on ferredoxin. Biochem. J. 100, 263–273 (1966).
Sanderson, G.W. & Cocking, E.C. Enzymic assimilation of nitrate in tomato plants. I. Reduction of nitrate to nitrite. Plant Physiol. 39, 416–422 (1964).
Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976).
Misko, T.P. et al. A fluorometric assay for the measurement of nitrite in biological samples. Anal. Biochem. 214, 11–16 (1993).
Acknowledgements
We thank Sakimi Watanabe and Sachiko Nomura for their technical assistance in establishing this protocol.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Nishimura, A., Aichi, I. & Matsuoka, M. A protocol for Agrobacterium-mediated transformation in rice. Nat Protoc 1, 2796–2802 (2006). https://doi.org/10.1038/nprot.2006.469
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.469
This article is cited by
-
OsMKK1 is a novel element that positively regulates the Xa21-mediated resistance response to Xanthomonas oryzae pv. oryzae in rice
Plant Cell Reports (2024)
-
IPA1 improves drought tolerance by activating SNAC1 in rice
BMC Plant Biology (2023)
-
Hidden prevalence of deletion-inversion bi-alleles in CRISPR-mediated deletions of tandemly arrayed genes in plants
Nature Communications (2023)
-
A natural mutation in the promoter of Ms-cd1 causes dominant male sterility in Brassica oleracea
Nature Communications (2023)
-
CRISPR/Cas9-mediated editing of Wx and BADH2 genes created glutinous and aromatic two-line hybrid rice
Molecular Breeding (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.