Abstract
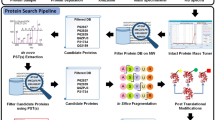
We describe a protocol for selective extraction of the amino (N)-terminal-most peptide of a protein or a mixture of proteins after proteolysis. The first stage of the protocol blocks the free amino groups α and ε (the latter being lysyl residues) on the intact proteins by acetylation. In the second stage, proteolysis of the acetylated proteins yields a mixture of N-terminally acetylated (true N-terminal) and non-acetylated (internal and carboxy-terminal) peptides. Affinity capture of peptides bearing free amino groups using an immobilized amine-reactive reagent removes internal peptides from the mixture. The unbound fraction is highly enriched in N-terminal peptides, which can be analyzed without further treatment. This method is compatible with a range of proteolytic enzymes and fragmentation methods, and should take 2 d to complete. The N-terminal peptides can then be analyzed by mass spectrometry. This low cost, rapid method is readily adopted using off the shelf reagents.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bogdanov, B. & Smith, R.D. Proteomics by FTICR mass spectrometry: top down and bottom up. Mass Spectrom. Rev. 24, 168–200 (2005).
Wu, C.C. & MacCoss, M.J. Shotgun proteomics: tools for the analysis of complex biological systems. Curr. Opin. Mol. Ther. 4, 242–250 (2002).
Liu, H., Lin, D. & Yates, J.R. Multidimensional separations for protein/peptide analysis in the post-genomic era. Biotechniques 4, 898–902 (2002).
Mirzaei, H. & Regnier, F. Structure specific chromatographic selection in targeted proteomics. J. Chromatogr. B 817, 23–34 (2005).
Smolka, M.B., Zhou, H., Purkayastha, S. & Aebersold, R. Optimization of the isotope-coded affinity tag-labeling procedure for quantitative proteome analysis. Anal Biochem. 297, 25–31 (2001).
Raggiaschi, R., Gotta, S. & Terstappen, G. Phosphoproteome analysis. Biosci. Rep. 25, 33–44 (2005).
Kweon, H.K. & Hakansson, K. Selective zirconium dioxide-based enrichment of phosphorylated peptides for mass spectrometric analysis. Anal. Chem. 78, 1743–1749 (2006).
West, I. & Goldring, O. Lectin affinity chromatography. Methods Mol. Biol. 59, 177–185 (1996).
Gevaert, K., Van Damme, P., Martens, L. & Vandekerckhove, J. Diagonal reverse-phase chromatography applications in peptide-centric proteomics: ahead of catalogue-omics? Anal. Biochem. 345, 18–29 (2005).
Martens, L. et al. The human platelet proteome mapped by peptide-centric proteomics: a functional protein profile. Proteomics 5, 3193–3204 (2005).
Gevaert, K. et al. Exploring proteomes and analyzing protein processing by mass spectrometric identification of sorted N-terminal peptides. Nat. Biotechnol. 21, 566–569 (2003).
Kasai, K.-I. Trypsin and affinity chromatography. J Chromatogr. A 597, 3–18 (1992).
Sechi, S. & Chait, B.T. A method to define the carboxyl terminal of proteins. Anal. Chem. 72, 3374–3378 (2000).
Yamaguchi, M. et al. High-throughput method for N-terminal sequencing of proteins by MALDI mass spectrometry. Anal. Chem. 77, 645–651 (2005).
Chelius, D. & Shaler, T.A. Capture of peptides with N-terminal serine and threonine: a sequence-specific chemical method for peptide mixture simplification. Bioconjugate Chem. 14, 205–211 (2003).
McDonald, L., Robertson, D.H., Hurst, J.L. & Beynon, R.J. Positional proteomics: selective recovery and analysis of N-terminal proteolytic peptides. Nat. Methods 2, 955–957 (2005).
Van Sommeren, A.P.G., Machielsen, P.A.G.M. & Gribnau,, T.C.J. Comparison of three activated agaroses for use in affinity chromatography: effects on coupling performance and ligand leakage. J. Chromatogr. A 639, 23–31 (1993).
Hayter, J.R., Robertson, D.H.L., Gaskell, S.J. & Beynon, R.J. Proteome analysis of intact proteins in complex mixtures. Mol. Cell Proteomics 2, 85–95 (2003).
Doherty, M.K. et al. The proteome of chicken skeletal muscle: changes in soluble protein expression during growth in a layer strain. Proteomics 4, 2082–2093 (2004).
Veenstra, T.D., Conrads, T.P. & Issaq, H.J. What to do with “one-hit wonders”? Electrophoresis 25, 1278–1279 (2004).
Acknowledgements
This work was supported by the Engineering and Physical Sciences Research Council.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
McDonald, L., Beynon, R. Positional proteomics: preparation of amino-terminal peptides as a strategy for proteome simplification and characterization. Nat Protoc 1, 1790–1798 (2006). https://doi.org/10.1038/nprot.2006.317
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.317
This article is cited by
-
Characterising proteolysis during SARS-CoV-2 infection identifies viral cleavage sites and cellular targets with therapeutic potential
Nature Communications (2021)
-
C-Terminal sequencing of protein by MALDI mass spectrometry through the specific derivatization of the α-carboxyl group with 3-aminopropyltris-(2,4,6-trimethoxyphenyl)phosphonium bromide
Analytical and Bioanalytical Chemistry (2012)
-
Selecting protein N-terminal peptides by combined fractional diagonal chromatography
Nature Protocols (2011)
-
Identifying and quantifying proteolytic events and the natural N terminome by terminal amine isotopic labeling of substrates
Nature Protocols (2011)
-
Isotopic labeling of terminal amines in complex samples identifies protein N-termini and protease cleavage products
Nature Biotechnology (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.