Abstract
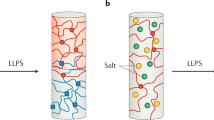
Intracellular organelles are either membrane-bound vesicles or membrane-less compartments that are made up of proteins and RNA. These organelles play key biological roles, by compartmentalizing the cell to enable spatiotemporal control of biological reactions. Recent studies suggest that membrane-less intracellular compartments are multicomponent viscous liquid droplets that form via phase separation. Proteins that have an intrinsic tendency for being conformationally heterogeneous seem to be the main drivers of liquid–liquid phase separation in the cell. These findings highlight the relevance of classical concepts from the physics of polymeric phase transitions for understanding the assembly of intracellular membrane-less compartments. However, applying these concepts is challenging, given the heteropolymeric nature of protein sequences, the complex intracellular environment, and non-equilibrium features intrinsic to cells. This provides new opportunities for adapting established theories and for the emergence of new physics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Lu, A. et al. Unified polymerization mechanism for the assembly of ASC-dependent inflammasomes. Cell 156, 1193–1206 (2014).
Li, J. et al. The RIP1/RIP3 necrosome forms a functional amyloid signaling complex required for programmed necrosis. Cell 150, 339–350 (2012).
An, S., Kumar, R., Sheets, E. D. & Benkovic, S. J. Reversible compartmentalization of de novo purine biosynthetic complexes in living cells. Science 320, 103–106 (2008).
Zwicker, D., Decker, M., Jaensch, S., Hyman, A. A. & Julicher, F. Centrosomes are autocatalytic droplets of pericentriolar material organized by centrioles. Proc. Natl Acad. Sci. USA 111, E2636–E2645 (2014).
Baron, M. K. et al. An architectural framework that may lie at the core of the postsynaptic density. Science 311, 531–535 (2006).
Ogrodnik, M. et al. Dynamic JUNQ inclusion bodies are asymmetrically inherited in mammalian cell lines through the asymmetric partitioning of vimentin. Proc. Natl Acad. Sci. USA 111, 8049–8054 (2014).
Mao, Y. S., Zhang, B. & Spector, D. L. Biogenesis and function of nuclear bodies. Trends Genet. 27, 295–306 (2011).
Ramaswami, M., Taylor, J. P. & Parker, R. Altered ribostasis: RNA-protein granules in degenerative disorders. Cell 154, 727–736 (2013).
Brangwynne, C. P. et al. Germline P granules are liquid droplets that localize by controlled dissolution/condensation. Science 324, 1729–1732 (2009).
Lee, C. F., Brangwynne, C. P., Gharakhani, J., Hyman, A. A. & Julicher, F. Spatial organization of the cell cytoplasm by position-dependent phase separation. Phys. Rev. Lett. 111, 088101 (2013).
Elbaum-Garfinkle, S. et al. The disordered P granule protein LAF-1 drives phase separation into droplets with tunable viscosity and dynamics. Proc. Natl Acad. Sci. USA 112, 7189–7194 (2015).
Hubstenberger, A., Noble, S. L., Cameron, C. & Evans, T. C. Translation repressors, an RNA helicase, and developmental cues control RNP phase transitions during early development. Dev. Cell 27, 161–173 (2013).
Brangwynne, C. P., Mitchison, T. J. & Hyman, A. A. Active liquid-like behavior of nucleoli determines their size and shape in Xenopus laevis oocytes. Proc. Natl Acad. Sci. USA 108, 4334–4339 (2011).
Feric, M. & Brangwynne, C. P. A nuclear F-actin scaffold stabilizes RNP droplets against gravity in large cells. Nature Cell Biol. 15, 1253–1259 (2013).
Weber, S. C. & Brangwynne, C. P. Inverse size scaling of the nucleolus by a concentration-dependent phase transition. Curr. Biol. 25, 641–646 (2015).
Berry, J., Weber, S. C., Vaidya, N., Haataja, M. & Brangwynne, C. P. RNA transcription modulates phase transition-driven nuclear body assembly. Proc. Natl Acad. Sci. USA 112, E5237–E5245 (2015).
Wippich, F. et al. Dual specificity kinase DYRK3 couples stress granule condensation/dissolution to mTORC1 signaling. Cell 152, 791–805 (2013).
Sear, R. P. Dishevelled: A protein that functions in living cells by phase separating. Soft Matter 3, 680–684 (2006).
Li, P. et al. Phase transitions in the assembly of multivalent signalling proteins. Nature 483, 336–340 (2012).
Singer, A. B. & Gall, J. G. An inducible nuclear body in the Drosophila germinal vesicle. Nucleus 2, 403–409 (2011).
Altmeyer, M. et al. Liquid demixing of intrinsically disordered proteins is seeded by poly(ADP-ribose). Nature Commun. 6, 8088 (2015).
Nott, T. et al. Phase transition of a disordered Nuage protein generates environmentally responsive membraneless organelles. Mol. Cell 57, 936–947 (2015).
Fromm, S. A. et al. In vitro reconstitution of a cellular phase-transition process that involves the mRNA decapping machinery. Angew. Chem. 53, 7354–7359 (2014).
Hennig, S. et al. Prion-like domains in RNA binding proteins are essential for building subnuclear paraspeckles. J. Cell Biol. 210, 529–539 (2015).
Banjade, S. & Rosen, M. K. Phase transitions of multivalent proteins can promote clustering of membrane receptors. eLife 3, e04123 (2014).
van der Lee, R. et al. Classification of intrinsically disordered regions and proteins. Chem. Rev. 114, 6589–6631 (2014).
Wright, P. E. & Dyson, H. J. Intrinsically disordered proteins in cellular signalling and regulation. Nature Rev. Mol. Cell Biol. 16, 18–29 (2015).
Uversky, V. N., Kuznetsova, I. M., Turoverov, K. K. & Zaslavsky, B. Intrinsically disordered proteins as crucial constituents of cellular aqueous two phase systems and coacervates. FEBS Lett. 589, 15–22 (2015).
Wang, J. T. et al. Regulation of RNA granule dynamics by phosphorylation of serine-rich, intrinsically disordered proteins in C. elegans. eLife 3, e04591 (2014).
Holehouse, A. S., Garai, K., Lyle, N., Vitalis, A. & Pappu, R. V. Quantitative assessments of the distinct contributions of polypeptide backbone amides versus side chain groups to chain expansion via chemical denaturation. J. Am. Chem. Soc. 137, 2984–2995 (2015).
Crick, S. L., Jayaraman, M., Frieden, C., Wetzel, R. & Pappu, R. V. Fluorescence correlation spectroscopy shows that monomeric polyglutamine molecules form collapsed structures in aqueous solutions. Proc. Natl Acad. Sci. USA 103, 16764–16769 (2006).
Mukhopadhyay, S., Krishnan, R., Lemke, E. A., Lindquist, S. & Deniz, A. A. A natively unfolded yeast prion monomer adopts an ensemble of collapsed and rapidly fluctuating structures. Proc. Natl Acad. Sci. USA 104, 2649–2654 (2007).
Tran, H. T., Mao, A. & Pappu, R. V. Role of backbone-solvent interactions in determining conformational equilibria of intrinsically disordered proteins. J. Am. Chem. Soc. 130, 7380–7392 (2008).
Crick, S. L., Ruff, K. M., Garai, K., Frieden, C. & Pappu, R. V. Unmasking the roles of N- and C-terminal flanking sequences from exon 1 of huntingtin as modulators of polyglutamine aggregation. Proc. Natl Acad. Sci. USA 110, 20075–20080 (2013).
Lai, J. et al. Intrinsically disordered proteins aggregate at fungal cell-to-cell channels and regulate intercellular connectivity. Proc. Natl Acad. Sci. USA 109, 15781–15786 (2012).
Srinivasan, N., Bhagawati, M., Ananthanarayanan, B. & Kumar, S. Stimuli-sensitive intrinsically disordered protein brushes. Nature Commun. 5, 5145 (2014).
Das, R. K. & Pappu, R. V. Conformations of intrinsically disordered proteins are influenced by linear sequence distributions of oppositely charged residues. Proc. Natl Acad. Sci. USA 110, 13392–13397 (2013).
Schmidt, H. B. & Gorlich, D. Nup98 FG domains from diverse species spontaneously phase-separate into particles with nuclear pore-like permselectivity. eLife 4, e04251 (2015).
Alberti, S., Halfmann, R., King, O., Kapila, A. & Lindquist, S. A systematic survey identifies prions and illuminates sequence features of prionogenic proteins. Cell 137, 146–158 (2009).
Kim, H. J. et al. Mutations in prion-like domains in hnRNPA2B1 and hnRNPA1 cause multisystem proteinopathy and ALS. Nature 495, 467–473 (2013).
Kato, M. et al. Cell-free formation of RNA granules: Low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Li, Y. R., King, O. D., Shorter, J. & Gitler, A. D. Stress granules as crucibles of ALS pathogenesis. J. Cell Biol. 201, 361–372 (2013).
Weber, S. C. & Brangwynne, C. P. Getting RNA and protein in phase. Cell 149, 1188–1191 (2012).
Patel, A. et al. A liquid-to-solid phase transition of the ALS ProteinFUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Molliex, A. et al. Phase separation by low-complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133 (2015).
Lin, Y., Protter, D. S. W., Rosen, M. K. & Parker, R. Formation and maturation of phase separated liquid droplets by RNA binding proteins. Mol. Cell 60, 1–12 (2015).
Zhang, H. et al. RNA controls PolyQ protein phase transitions. Mol. Cell 60, 220–230 (2015).
Galkin, O., Chen, K., Nagel, R. L., Hirsch, R. E. & Vekilov, P. G. Liquid–liquid separation in solutions of normal and sickle cell hemoglobin. Proc. Natl Acad. Sci. USA 99, 8479–8483 (2002).
Liu, C. et al. Phase separation in aqueous solutions of lens gamma-crystallins: Special role of gamma s. Proc. Natl Acad. Sci. USA 93, 377–382 (1996).
ten Wolde, P. R. & Frenkel, D. Enhancement of protein crystal nucleation by critical density fluctuations. Science 277, 1975–1978 (1997).
Kroschwald, S. et al. Promiscuous interactions and protein disaggregases determine the material state of stress-inducible RNP granules. eLife 4, e06807 (2015).
Buchan, J. R., Kolaitis, R. M., Taylor, J. P. & Parker, R. Eukaryotic stress granules are cleared by autophagy and Cdc48/VCP function. Cell 153, 1461–1474 (2013).
Sweeny, E. A. et al. The Hsp104 N-terminal domain enables disaggregase plasticity and potentiation. Mol. Cell 57, 836–849 (2015).
Parry, B. R. et al. The bacterial cytoplasm has glass-like properties and is fluidized by metabolic activity. Cell 156, 183–194 (2014).
Weber, S. C., Spakowitz, A. J. & Theriot, J. A. Nonthermal ATP-dependent fluctuations contribute to the in vivo motion of chromosomal loci. Proc. Natl Acad. Sci. USA 109, 7338–7343 (2012).
Guo, M. et al. Probing the stochastic, motor-driven properties of the cytoplasm using force spectrum microscopy. Cell 158, 822–832 (2014).
Bursac, P. et al. Cytoskeletal remodelling and slow dynamics in the living cell. Nature Mater. 4, 557–561 (2005).
Sollich, P., Lequeux, F., Hebraud, P. & Cates, M. E. Rheology of soft glassy materials. Phys. Rev. Lett. 78, 2020 (1997).
Stillinger, F. H. & Debenedetti, P. G. Glass transition thermodynamics and kinetics. Annu. Rev. Condens. Matter Phys. 4, 263–285 (2013).
Jack, R. L., Hedges, L. O., Garrahan, J. P. & Chandler, D. Preparation and relaxation of very stable glassy states of a simulated liquid. Phys. Rev. Lett. 107, 275702 (2011).
Weber, J. K., Jack, R. L., Schwantes, C. R. & Pande, V. S. Dynamical phase transitions reveal amyloid-like states on protein folding landscapes. Biophys. J. 107, 974–982 (2014).
Griffin, E. E., Odde, D. J. & Seydoux, G. Regulation of the MEX-5 gradient by a spatially segregated kinase/phosphatase cycle. Cell 146, 955–968 (2011).
Tritschler, F., Huntzinger, E. & Izaurralde, E. Role of GW182 proteins and PABPC1 in the miRNA pathway: A sense of déjà vu. Nature Rev. Mol. Cell Biol. 11, 379–384 (2010).
Huggins, M. L. Some properties of solutions of long-chain compounds. J. Phys. Chem. 46, 151–158 (1942).
Flory, P. J. Thermodynamics of high polymer solutions. J. Chem. Phys. 10, 51–61 (1942).
Das, R. K., Ruff, K. M. & Pappu, R. V. Relating sequence encoded information to form and function of intrinsically disordered proteins. Curr. Opin. Struct. Biol. 32, 102–112 (2015).
Overbeek, J. T. G. & Voorn, M. J. Phase separation in polyelectrolyte solutions. Theory of complex coacervation. J. Cell. Comp. Physiol. 49, 7–26 (1957).
Acknowledgements
We thank Y. Shin and S. E.-Garfinkle for helpful comments on the manuscript. This work was supported by grants 1DP2GM105437-01 from NIH and 1253035 from NSF to C.P.B., Odysseus grant G.0029.12 from FWO to P.T., and 5R01NS056114 from the NIH to R.V.P. We are grateful to A. Holehouse for assistance with Fig. 2.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Brangwynne, C., Tompa, P. & Pappu, R. Polymer physics of intracellular phase transitions. Nature Phys 11, 899–904 (2015). https://doi.org/10.1038/nphys3532
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nphys3532
This article is cited by
-
Crosstalk between protein post-translational modifications and phase separation
Cell Communication and Signaling (2024)
-
High-throughput and proteome-wide discovery of endogenous biomolecular condensates
Nature Chemistry (2024)
-
Present and future of synthetic cell development
Nature Reviews Molecular Cell Biology (2024)
-
Asymmetric oligomerization state and sequence patterning can tune multiphase condensate miscibility
Nature Chemistry (2024)
-
Biomolecular condensates form spatially inhomogeneous network fluids
Nature Communications (2024)