Abstract
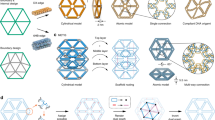
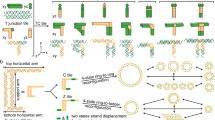
Structural DNA nanotechnology1,2,3,4 and the DNA origami technique5, in particular, have provided a range of spatially addressable two- and three-dimensional nanostructures6,7,8,9,10. These structures are, however, typically formed of tightly packed parallel helices5,6,7,8,9. The development of wireframe structures10,11 should allow the creation of novel designs with unique functionalities, but engineering complex wireframe architectures with arbitrarily designed connections between selected vertices in three-dimensional space remains a challenge. Here, we report a design strategy for fabricating finite-size wireframe DNA nanostructures with high complexity and programmability. In our approach, the vertices are represented by n × 4 multi-arm junctions (n = 2–10) with controlled angles, and the lines are represented by antiparallel DNA crossover tiles12 of variable lengths. Scaffold strands are used to integrate the vertices and lines into fully assembled structures displaying intricate architectures. To demonstrate the versatility of the technique, a series of two-dimensional designs including quasi-crystalline patterns and curvilinear arrays or variable curvatures, and three-dimensional designs including a complex snub cube and a reconfigurable Archimedean solid were constructed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Aldaye, F. A., Palmer, A. L. & Sleiman, H. F. Assembling materials with DNA as the guide. Science 321, 1795–1799 (2008).
Seeman, N. C. Nanomaterials based on DNA. Annu. Rev. Biochem. 79, 65–87 (2010).
Pinheiro, A. V., Han, D. R., Shih, W. M. & Yan, H. Challenges and opportunities for structural DNA nanotechnology. Nature Nanotech. 6, 763–772 (2011).
Zhang, F., Nangreave, J., Liu, Y. & Yan, H. Structural DNA nanotechnology: state of the art and future perspective. J. Am. Chem. Soc. 136, 11198–11211 (2014).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Andersen, E. S. et al. Self-assembly of a nanoscale DNA box with a controllable lid. Nature 459, 73–75 (2009).
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009).
Han, D. R. et al. DNA origami with complex curvatures in three-dimensional space. Science 332, 342–346 (2011).
Han, D. R. et al. DNA gridiron nanostructures based on four-arm junctions. Science 339, 1412–1415 (2013).
He, Y. et al. Hierarchical self-assembly of DNA into symmetric supramolecular polyhedra. Nature 452, 198–201 (2008).
Winfree, E., Liu, F. R., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
West, D. B. Introduction to Graph Theory (Prentice Hall, 1996).
Gibbons, A. Algorithmic Graph Theory (Cambridge Univ. Press, 1985).
He, Y., Chen, Y., Liu, H., Ribbe, A. E. & Mao, C. Self-assembly of hexagonal DNA two-dimensional (2D) arrays. J. Am. Chem. Soc. 127, 12202–12203 (2005).
Yan, H., Park, S. H., Finkelstein, G., Reif, J. H. & LaBean, T. H. DNA templated self-assembly of protein arrays and highly conductive nanowires. Science 301, 1882–1884 (2003).
He, Y., Tian, Y., Ribbe, A. E. & Mao, C. D. Highly connected two-dimensional crystals of DNA six-point-stars. J. Am. Chem. Soc. 128, 15978–15979 (2006).
Liu, Y., Ke, Y. G. & Yan, H. Self-assembly of symmetric finite size DNA nanoarrays. J. Am. Chem. Soc. 127, 17140–17141 (2005).
Sommerville, D. M. Y. Introduction to the Geometry of N Dimensions (E. P. Dutton, 1929).
Wells, A. F. Three-Dimensional Nets and Polyhedra (Wiley, 1977).
Chen, J. & Seeman, N. C. Synthesis from DNA of a molecule with the connectivity of a cube. Nature 350, 631–633 (1991).
Zhang, Y. & Seeman, N. C. Consruction of a DNA-truncated octahedron. J. Am. Chem. Soc. 116, 1661–1669 (1994).
Yan, H., Zhang, X., Shen, Z. & Seeman, N. C. A robust DNA mechanical device controlled by hybridization topology. Nature 415, 62–65 (2002).
Kato, T., Goodman, R. P., Erben, C. M., Turberfield, A. J. & Namba, K. High-resolution structural analysis of a DNA nanostructure by cryoEM. Nano Lett. 9, 2747–2750 (2009).
Shen, Z., Yan, H., Wang, T. & Seeman, N. C. Paranemic crossover DNA: a generalized Holliday structure with applications in nanotechnology. J. Am. Chem. Soc. 126, 1666–1674 (2004).
Geary, C., Rothemund, P. W. K. & Andersen, E. S. A single-stranded architecture for cotranscriptional folding of RNA nanostructures. Science 345, 799–804 (2014).
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Goddard, T. D., Huang, C. C. & Ferrin, T. E. Visualizing density maps with UCSF Chimera. J. Struct. Biol. 157, 281–287 (2007).
Acknowledgements
This research was partly supported by grants to H.Y. and Y.L. from the National Science Foundation (nos. 1360635 and 1334109), the Army Research Office (no. W911NF-12-1-0420) and the National Institutes of Health (no. R01GM104960). H.Y. was supported by the Presidential Strategic Initiative Fund from Arizona State University. The authors thank M. Madjidi for proofreading.
Author information
Authors and Affiliations
Contributions
H.Y., Y.L. and F.Z. conceived and designed the experiment. F.Z., S.J., S.W. and Y.L. performed the experiments. F.Z., S.J., S.W. and Y.L. analysed the data. All authors discussed the results. All authors contributed to the writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Information (PDF 26533 kb)
Rights and permissions
About this article
Cite this article
Zhang, F., Jiang, S., Wu, S. et al. Complex wireframe DNA origami nanostructures with multi-arm junction vertices. Nature Nanotech 10, 779–784 (2015). https://doi.org/10.1038/nnano.2015.162
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2015.162
This article is cited by
-
Exploring the diverse biomedical applications of programmable and multifunctional DNA nanomaterials
Journal of Nanobiotechnology (2023)
-
Fabricating higher-order functional DNA origami structures to reveal biological processes at multiple scales
NPG Asia Materials (2023)
-
Interfacing DNA nanotechnology and biomimetic photonic complexes: advances and prospects in energy and biomedicine
Journal of Nanobiotechnology (2022)
-
Prospects and challenges of dynamic DNA nanostructures in biomedical applications
Bone Research (2022)
-
DNA origami single crystals with Wulff shapes
Nature Communications (2021)