Abstract
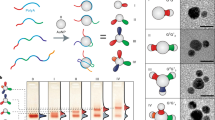
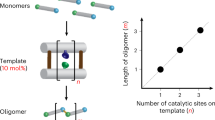
Synthetic polymers are ubiquitous in the modern world, but our ability to exert control over the molecular conformation of individual polymers is very limited. In particular, although the programmable self-assembly of oligonucleotides and proteins into artificial nanostructures has been demonstrated, we currently lack the tools to handle other types of synthetic polymers individually and thus the ability to utilize and study their single-molecule properties. Here we show that synthetic polymer wires containing short oligonucleotides that extend from each repeat can be made to assemble into arbitrary routings. The wires, which can be more than 200 nm in length, are soft and bendable, and the DNA strands allow individual polymers to self-assemble into predesigned routings on both two- and three-dimensional DNA origami templates. The polymers are conjugated and potentially conducting, and could therefore be used to create molecular-scale electronic or optical wires in arbitrary geometries.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Heeger, A. J. Semiconducting and metallic polymers: the fourth generation of polymeric materials. Synthetic Met. 125, 23–42 (2001).
Heeger, A. J. Semiconducting polymers: the third generation. Chem. Soc. Rev. 39, 2354–2371 (2010).
Facchetti, A. π-Conjugated polymers for organic electronics and photovoltaic cell applications. Chem. Mater. 23, 733–758 (2011).
Baeg, K.-J. et al. High speeds complementary integrated circuits fabricated with all-printed polymeric semiconductors. J. Polym. Sci. B 49, 62–67 (2011).
Gong, S., Yang, C. & Qin, J. Efficient phosphorescent polymer light-emitting diodes by suppressing triplet energy back transfer. Chem. Soc. Rev. 41, 4797–4807 (2012).
Zheng, H. et al. All-solution processed polymer light-emitting diode displays. Nature Commun. 4, 1–7 (2013).
Günes, S., Neugebauer, H. & Sariciftci, N. S. Conjugated polymer-based organic solar cells. Chem. Rev. 107, 1324–1338 (2007).
Thomas, S. W., Joly, G. D. & Swager, T. M. Chemical sensors based on amplifying fluorescent conjugated polymers. Chem. Rev. 107, 1339–1386 (2007).
Barbara, P. F., Gesquiere, A. J., Park, S. J. & Lee, Y. J. Single-molecule spectroscopy of conjugated polymers. Acc. Chem. Res. 38, 602–610 (2005).
Hugel, T. et al. Single-molecule optomechanical cycle. Science 296, 1103–1106 (2002).
Kawai, S. et al. Quantifying the atomic-level mechanics of single long physisorbed molecular chains. Proc. Natl Acad. Sci. USA 111, 3968–3972 (2014).
Taniguchi, M. et al. Self-organized interconnect method for molecular devices. J. Am. Chem. Soc. 128, 15062–15063 (2006).
Lafferentz, L. et al. Conductance of a single conjugated polymer as a continuous function of its length. Science 323, 1193–1197 (2009).
Shimomura, T. et al. Conductivity measurement of insulated molecular wire formed by molecular nanotube and polyaniline. Synthetic Met. 153, 497–500 (2005).
Kiriy, A. et al. Cascade of coil–globule conformational transitions of single flexible polyelectrolyte molecules in poor solvent. J. Am. Chem. Soc. 124, 13454–13462 (2002).
Shimomura, T., Akai, T., Abe, T. & Ito, K. Atomic force microscopy observation of insulated molecular wire formed by conducting polymer and molecular nanotube. J. Chem. Phys. 116, 1753–1756 (2002).
Ouchi, M., Badi, N., Lutz, J.-F. & Sawamoto, M. Single-chain technology using discrete synthetic macromolecules. Nature Chem. 3, 917–924 (2011).
Müllen, K. Evolution of graphene molecules: structural and functional complexity as driving forces behind nanoscience. ACS Nano 8, 6531–6541 (2011).
Palma, C.-A. & Samori, P. Blueprinting macromolecular electronics. Nature Chem. 3, 431–436 (2011).
Lörtscher, E. Wiring molecules into circuits. Nature Nanotech. 8, 381–384 (2013).
Peng, H., Zhang, L., Soeller, C. & Travas-Sejdic, J. Conducting polymers for electrochemical DNA sensing. Biomaterials 30, 2132–2148 (2009).
Lo, P. K. & Sleiman, H. F. Nucleobase-templated polymerization: copying the chain length and polydispersity of living polymers into conjugated polymers. J. Am. Chem. Soc. 131, 4182–4183 (2009).
Rothemund, P. W. K. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Tørring, T., Voigt, N. V., Nangreave, J., Yan, H. & Gothelf, K. V. DNA origami: a quantum leap for self-assembly of complex structures. Chem. Soc. Rev. 40, 5636–5646 (2011).
Sacca, B. & Niemeyer, C. M. Functionalization of DNA nanostructures with proteins. Chem. Soc. Rev. 40, 5910–5921 (2011).
Wang, Z. G. & Ding, B. Engineering DNA self-assemblies as templates for functional nanostructures. Acc. Chem. Res. 47, 1654–1662 (2014).
Maune, H. T. et al. Self-assembly of carbon nanotubes into two-dimensional geometries using DNA origami templates. Nature Nanotech. 5, 61–66 (2010).
Chaix, C., Minard-Basquin, C., Delair, T., Pichot, C. & Mandrand, B. Oligonucleotide synthesis on maleic anhydride copolymers covalently bound to silica spherical support and characterization of the obtained conjugates. J. Appl. Polym. Sci. 70, 2487 (1998).
Henckens, A., Duyssens, I., Lutsen, L., Vanderzande, D. & Cleij, T. J. Synthesis of poly(p-phenylene vinylene) and derivatives via a new precursor route, the dithiocarbamate route. Polymer 47, 123–131 (2006).
Vandenbergh, J. et al. Synthesis and characterization of water-soluble poly(p-phenylene vinylene) derivatives via the dithiocarbamate precursor route. Eur. Polym. J. 47, 1827–1835 (2011).
Minard-Basquin, C., Chaix, C., Pichot, C. & Mandrand, B. Oligonucleotide−polymer conjugates: effect of the method of synthesis on their structure and performance in diagnostic assays. Bioconjugate Chem. 11, 795–805 (2000).
Volcke, C. et al. Influence of DNA condensation state on transfection efficiency in DNA/polymer complexes: an AFM and DLS comparative study. J. Biotechnol. 125, 11–21 (2006).
Zhang, S. et al. Coexistence of ribbon and helical fibrils originating from hIAPP20–29 revealed by quantitative nanomechanical atomic force microscopy. Proc. Natl Acad. Sci. USA 110, 2798–2803 (2013).
Pfeffer, C. et al. Filamentous bacteria transport electrons over centimetre distances. Nature 491, 218–221 (2012).
Sinensky, A. K. & Belcher, A. M. Label-free and high-resolution protein/DNA nanoarray analysis using Kelvin probe force microscopy. Nature Nanotech. 2, 653–659 (2007).
Ke, Y., Lindsay, S., Chang, Y., Liu, Y. & Yan, H. Self-assembled water-soluble nucleic acid probe tiles for label-free RNA hybridization assays. Science 319, 180–183 (2008).
Shih, W. M. & Lin, C. Knitting complex weaves with DNA origami. Curr. Opin. Chem. Biol. 20, 276–282 (2010).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Iinuma, R. et al. Polyhedra self-assembled from DNA tripods and characterized with 3D DNA-PAINT. Science 344, 65–69 (2014).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nature Methods 11, 313 (2014).
Jungmann, R. et al. Single-molecule kinetics and super-resolution microscopy by fluorescence imaging of transient binding on DNA origami. Nano Lett. 10, 4756–4761 (2010).
Kao, H. P. & Verkman, A. S. Tracking of single fluorescent particles in three dimensions: use of cylindrical optics to encode particle position. Biophys. J. 67, 1291–1300 (1994).
Huang, B., Wang, W., Bates, M. & Zhuang, X. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319, 810–813 (2008).
Burns, P. L. et al. Chemical tuning of the electronic properties of poly(p-phenylenevinylene)-based copolymers. J. Am. Chem. Soc. 115, 10117–10124 (1993).
Kershner, R. J. et al. Placement and orientation of individual DNA shapes on lithographically patterned surfaces. Nature Nanotech. 4, 557–561 (2009).
Acknowledgements
We thank P.W.K. Rothemund for discussions. This work was funded by the Danish National Research Foundation (Centre for DNA Nanotechnology, DNRF81), Sino-Danish Centre for Education and Research, Carlsberg Foundation, Danish Research Council (V.B.) (Sapere Aude Starting Grant (A.N.Z. and V.B.), STENO grant and an individual post-doctorate grant (R.O.)), Villum Foundation (Young Investigator Program (M.D.)), and the Lundbeck Foundation (A.N.Z). R.J. acknowledges support from the Deutsche Forschungsgemeinschaft through the Emmy Noether program (DFG JU 2957/1–1) and the Max Planck Society. W.M.S. acknowledges support for the contributions to his laboratory from the National Science Foundation (CCF-1317291), Army Research Office (W911NF-12-1-0420) and the Wyss Institute for Biologically Inspired Engineering.
Author information
Authors and Affiliations
Contributions
J.B.K. and M.M. synthesized and characterized the APPV-DNA polymer. L.L., J.S., Q.L. and J.V. conducted the SPM experiments. A.L.B.K. and A.K. designed the DNA origami and conducted the AFM experiments. A.A.A.S. and A.N.Z. assisted in performing and interpreting the GPC analysis. R.O. performed XPS and assisted in the interpretation. D.G. measured the fluorescence quantum yields of the polymer. M.M. designed and conducted the FRET experiments. J.B.W., R.J., S.F.J.W., W.M.S. and K.V.G. designed the DNA-PAINT experiments. S.F.J.W. and W.M.S. designed the 3D DNA origami samples, and S.F.J.W. assembled, purified and characterized the 3D DNA origami samples. J.B.W., M.T.S. and F.S. performed the DNA-PAINT experiments and analysed the data. M.T.S. wrote the 3D image analysis and fitting software. R.J. performed the class averaging. P.Y. and R.J. supervised the DNA-PAINT study. A.Z., V.B. and F.B. supervised parts of the project. M.D. supervised and analysed the SPM experiments. K.V.G. conceived and supervised the project. J.B.K., M.D., S.F.J.W., A.L.B.K., J.B.W., M.T.S., R.J. and K.V.G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 16027 kb)
Rights and permissions
About this article
Cite this article
Knudsen, J., Liu, L., Bank Kodal, A. et al. Routing of individual polymers in designed patterns. Nature Nanotech 10, 892–898 (2015). https://doi.org/10.1038/nnano.2015.190
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2015.190
This article is cited by
-
Directly visualizing individual polyorganophosphazenes and their single-chain complexes with proteins
Communications Materials (2024)
-
Biotemplated precise assembly approach toward ultra-scaled high-performance electronics
Nature Protocols (2023)
-
Programmable multispecific DNA-origami-based T-cell engagers
Nature Nanotechnology (2023)
-
Site-directed placement of three-dimensional DNA origami
Nature Nanotechnology (2023)
-
Interfacing DNA nanotechnology and biomimetic photonic complexes: advances and prospects in energy and biomedicine
Journal of Nanobiotechnology (2022)