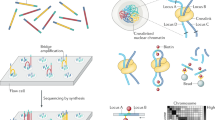
Abstract
The Wellcome Trust Sanger Institute is one of the world's largest genome centers, and a substantial amount of our sequencing is performed with 'next-generation' massively parallel sequencing technologies: in June 2008 the quantity of purity-filtered sequence data generated by our Genome Analyzer (Illumina) platforms reached 1 terabase, and our average weekly Illumina production output is currently 64 gigabases. Here we describe a set of improvements we have made to the standard Illumina protocols to make the library preparation more reliable in a high-throughput environment, to reduce bias, tighten insert size distribution and reliably obtain high yields of data.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bentley, D.R. Whole-genome re-sequencing. Curr. Opin. Genet. Dev. 16, 545–552 (2006).
Mardis, E.R. The impact of next-generation sequencing technology on genetics. Trends Genet. 24, 133–141 (2008).
Smith, D.R. et al. Rapid whole-genome mutational profiling using next-generation sequencing technologies. Genome Res. 18, 1638–1642 (2008).
Surzycki, S. DNA sequencing. in Basic Techniques in Molecular Biology 377–380 (Springer-Verlag, Berlin, 2000).
Albert, T.J. et al. Direct selection of human genomic loci by microarray hybridization. Nat. Methods 4, 903–905 (2007).
Hodges, E. et al. Genome-wide in situ exon capture for selective resequencing. Nat. Genet. 39, 1522–1527 (2007).
Sambrook, J., Fritsch, E. & Maniatis, T. Molecular Cloning: A Laboratory Manual, (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, 1989).
Smith, D. & Malek, J. Asymmetrical adapters and uses thereof. US patent 0172839 (2007).
Riley, J. et al. A novel, rapid method for the isolation of terminal sequences from yeast artificial chromosome (YAC) clones. Nucleic Acids Res. 18, 2887–2890 (1990).
Hawkins, T.L., O'Connor-Morin, T., Roy, A. & Santillan, C. DNA purification and isolation using a solid-phase. Nucleic Acids Res. 22, 4543–4544 (1994).
Meyer, M. et al. From micrograms to picograms: quantitative PCR reduces the material demands of high-throughput sequencing. Nucleic Acids Res. 36, e5 (2008).
Thomas, R. The denaturation of DNA. Gene 135, 77–79 (1993).
Mandel, M. & Marmur, J. Use of ultraviolet absorbance-temperature profile for determining the guanine plus cytosine content of DNA. Methods Enzymol. 12, 195–206 (1968).
Acknowledgements
We thank all the staff at Illumina for their support, particularly T. Ost, M. Gibbs, J. Smith, N. Gormley, V. Smith and K. Hall. We also thank C. Brown, A. Brown, R. Pettett, T. Skelly, N. Whiteford, L. Mamanova, E. Sheridan and E. Huckle for helpful discussions and assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
H.S. is an Illumina shareholder and share option holder.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1 , Supplementary Protocols 1–13 (PDF 110 kb)
Rights and permissions
About this article
Cite this article
Quail, M., Kozarewa, I., Smith, F. et al. A large genome center's improvements to the Illumina sequencing system. Nat Methods 5, 1005–1010 (2008). https://doi.org/10.1038/nmeth.1270
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1270
This article is cited by
-
Hypomethylation in MTNR1B: a novel epigenetic marker for atherosclerosis profiling using stenosis radiophenotype and blood inflammatory cells
Clinical Epigenetics (2023)
-
Knowledge Map and Global Trends in Root Exudates Research from 2012 to 2021: a Bibliometric Analysis
Journal of Soil Science and Plant Nutrition (2023)
-
Exploring cross-species genetic diversity: unveiling new insights in Megalobrama through whole genome-wide simple sequence repeats
Conservation Genetics (2023)
-
Identification and analysis of toxins in novel Bacillus thuringiensis strain Bt S3076-1 against Spodoptera frugiperda and Helicoverpa armigera (Lep.: Noctuidae)
Archives of Microbiology (2023)
-
Extensive mitogenomic heteroplasmy and its implications in the phylogeny of the fish genus Megalobrama
3 Biotech (2023)