Abstract
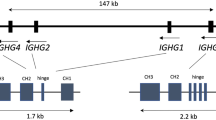
The only immunoglobulin heavy-chain classes known so far in teleosts have been μ and δ. We identify here a previously unknown class, immunoglobulin ζ, expressed in zebrafish and other teleosts. In the zebrafish heavy-chain locus, variable (V) gene segments lie upstream of two tandem diversity, joining and constant (DJC) clusters, resembling the mouse T cell receptor α (Tcra) and δ (Tcrd) locus. V genes rearrange to (DJC)ζ or to (DJC)μ without evidence of switch rearrangement. The zebrafish immunoglobulin ζ gene (ighz) and mouse Tcrd, which are proximal to the V gene array, are expressed earlier in development. In adults, ighz was expressed only in kidney and thymus, which are primary lymphoid organs in teleosts. This additional class adds complexity to the immunoglobulin repertoire and raises questions concerning the evolution of immunoglobulins and the regulation of the differential expression of ighz and ighm.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Accessions
GenBank/EMBL/DDBJ
References
Schatz, D.G.V. (D)J recombination. Immunol. Rev. 200, 5–11 (2004).
Cannon, J.P., Haire, R.N., Rast, J.P. & Litman, G.W. The phylogenetic origins of the antigen-binding receptors and somatic diversification mechanisms. Immunol. Rev. 200, 12–22 (2004).
Amemiya, C.T. & Litman, G.W. Complete nucleotide sequence of an immunoglobulin heavy-chain gene and analysis of immunoglobulin gene organization in a primitive teleost species. Proc. Natl. Acad. Sci. USA 87, 811–815 (1990).
Ghaffari, S.H. & Lobb, C.J. Nucleotide sequence of channel catfish heavy chain cDNA and genomic blot analyses. Implications for the phylogeny of Ig heavy chains. J. Immunol. 143, 2730–2739 (1989).
Daggfeldt, A., Bengten, E. & Pilstrom, L. A cluster type organization of the loci of the immunoglobulin light chain in Atlantic cod (Gadus morhua L.) and rainbow trout (Oncorhynchus mykiss Walbaum) indicated by nucleotide sequences of cDNAs and hybridization analysis. Immunogenetics 38, 199–209 (1993).
Danilova, N. & Steiner, L.A. B cells develop in the zebrafish pancreas. Proc. Natl. Acad. Sci. USA 99, 13711–13716 (2002).
Danilova, N., Hohman, V.S., Kim, E.H. & Steiner, L.A. Immunoglobulin variable-region diversity in the zebrafish. Immunogenetics 52, 81–91 (2000).
Ghaffari, S.H. & Lobb, C.J. Structure and genomic organization of a second cluster of immunoglobulin heavy chain gene segments in the channel catfish. J. Immunol. 162, 1519–1529 (1999).
Bengten, E. et al. The IgH locus of the channel catfish, Ictalurus punctatus, contains multiple constant region gene sequences: different genes encode heavy chains of membrane and secreted IgD. J. Immunol. 169, 2488–2497 (2002).
Hordvik, I. The impact of ancestral tetraploidy on antibody heterogeneity in salmonid fishes. Immunol. Rev. 166, 153–157 (1998).
Brodeur, P.H. & Riblet, R. The immunoglobulin heavy chain variable region (Igh-V) locus in the mouse. I. One hundred Igh-V genes comprise seven families of homologous genes. Eur. J. Immunol. 14, 922–930 (1984).
Ota, T. & Nei, M. Divergent evolution and evolution by the birth-and-death process in the immunoglobulin VH gene family. Mol. Biol. Evol. 11, 469–482 (1994).
Andersson, E. & Matsunaga, T. Jaw, adaptive immunity and phylogeny of vertebrate antibody VH gene family. Res. Immunol. 147, 233–240 (1996).
Wilson, M.R. et al. The immunoglobulin M heavy chain constant region gene of the channel catfish, Ictalurus punctatus: an unusual mRNA splice pattern produces the membrane form of the molecule. Nucleic Acids Res. 18, 5227–5233 (1990).
Bengten, E., Leanderson, T. & Pilstrom, L. Immunoglobulin heavy chain cDNA from the teleost Atlantic cod (Gadus morhua L.): nucleotide sequences of secretory and membrane form show an unusual splicing pattern. Eur. J. Immunol. 21, 3027–3033 (1991).
Hordvik, I., Voie, A.M., Glette, J., Male, R. & Endresen, C. Cloning and sequence analysis of two isotypic IgM heavy chain genes from Atlantic salmon, Salmo salar L. Eur. J. Immunol. 22, 2957–2962 (1992).
Lee, M.A., Bengten, E., Daggfeldt, A., Rytting, A.S. & Pilstrom, L. Characterization of rainbow trout cDNAs encoding a secreted and membrane-bound Ig heavy chain and the genomic intron upstream of the first constant exon. Mol. Immunol. 30, 641–648 (1993).
Kabat, E.A., Wu, T.T., Perry, H.M., Gottesman, K.S. & Foeller, C. Sequences of Proteins of Immunological Interest (US Department of Health and Human Services, National Institutes of Health, Washington, 1991).
Wiersma, E.J. et al. Analysis of IgM structures involved in J chain incorporation. J. Immunol. 158, 1719–1726 (1997).
Wilson, M. et al. A novel chimeric Ig heavy chain from a teleost fish shares similarities to IgD. Proc. Natl. Acad. Sci. USA 94, 4593–4597 (1997).
Stenvik, J. & Jorgensen, T.O. Immunoglobulin D (IgD) of Atlantic cod has a unique structure. Immunogenetics 51, 452–461 (2000).
Hordvik, I. Identification of a novel immunoglobulin delta transcript and comparative analysis of the genes encoding IgD in Atlantic salmon and Atlantic halibut. Mol. Immunol. 39, 85–91 (2002).
Srisapoome, P., Ohira, T., Hirono, I. & Aoki, T. Genes of the constant regions of functional immunoglobulin heavy chain of Japanese flounder, Paralichthys olivaceus. Immunogenetics 56, 292–300 (2004).
Saha, N.R., Suetake, H., Kikuchi, K. & Suzuki, Y. Fugu immunoglobulin D: a highly unusual gene with unprecedented duplications in its constant region. Immunogenetics 56, 438–447 (2004).
Hordvik, I., Thevarajan, J., Samdal, I., Bastani, N. & Krossoy, B. Molecular cloning and phylogenetic analysis of the Atlantic salmon immunoglobulin D gene. Scand. J. Immunol. 50, 202–210 (1999).
Hirono, I., Nam, B.H., Enomoto, J., Uchino, K. & Aoki, T. Cloning and characterisation of a cDNA encoding Japanese flounder Paralichthys olivaceus IgD. Fish Shellfish Immunol. 15, 63–70 (2003).
Gillies, S.D., Morrison, S.L., Oi, V.T. & Tonegawa, S. A tissue-specific transcription enhancer element is located in the major intron of a rearranged immunoglobulin heavy chain gene. Cell 33, 717–728 (1983).
Sakai, E., Bottaro, A., Davidson, L., Sleckman, B.P. & Alt, F.W. Recombination and transcription of the endogenous Ig heavy chain locus is effected by the Ig heavy chain intronic enhancer core region in the absence of the matrix attachment regions. Proc. Natl. Acad. Sci. USA 96, 1526–1531 (1999).
Magor, B.G. et al. An Ig heavy chain enhancer of the channel catfish Ictalurus punctatus: evolutionary conservation of function but not structure. J. Immunol. 153, 5556–5563 (1994).
Cioffi, C.C. et al. An IgH enhancer that drives transcription through basic helix-loop-helix and Oct transcription factor binding motifs. Functional analysis of the Eμ3′ enhancer of the catfish. J. Biol. Chem. 276, 27825–27830 (2001).
Magor, B.G., Ross, D.A., Pilstrom, L. & Warr, G.W. Transcriptional enhancers and the evolution of the IgH locus. Immunol. Today 20, 13–17 (1999).
Ellestad, K.K. & Magor, B.G. Evolution of transcriptional enhancers in the immunoglobulin heavy chain gene:functional characteristics of the zebrafish E(mu)3′ enhancer. Immunogenetics (in the press).
Aparicio, S. et al. Whole-genome shotgun assembly and analysis of the genome of Fugu rubripes. Science 297, 1301–1310 (2002).
Krangel, M.S., Carabana, J., Abbarategui, I., Schlimgen, R. & Hawwari, A. Enforcing order within a complex locus: current perspectives on the control of V(D)J recombination at the murine T-cell receptor α/δ locus. Immunol. Rev. 200, 224–232 (2004).
Desiderio, S.V. et al. Insertion of N regions into heavy-chain genes is correlated with expression of terminal deoxytransferase in B cells. Nature 311, 752–755 (1984).
Willett, C.E., Zapata, A.G., Hopkins, N. & Steiner, L.A. Expression of zebrafish rag genes during early development identifies the thymus. Dev. Biol. 182, 331–341 (1997).
Zapata, A. & Amemiya, C.T. Phylogeny of lower vertebrates and their immunological structures. Curr. Top. Microbiol. Immunol. 248, 67–107 (2000).
Ventura-Holman, T., Jones, J.C., Ghaffari, S.H. & Lobb, C.J. Structure and genomic organization of VH gene segments in the channel catfish: members of different VH gene families are interspersed and closely linked. Mol. Immunol. 31, 823–832 (1994).
Honjo, T., Muramatsu, M. & Fagarasan, S. AID: how does it aid antibody diversity? Immunity 20, 659–668 (2004).
Zhao, Y., Pan-Hammarstrom, Q., Zhao, Z. & Hammarstrom, L. Identification of the activation-induced cytidine deaminase gene from zebrafish: an evolutionary analysis. Dev. Comp. Immunol. 29, 61–71 (2005).
Glusman, G. et al. Comparative genomics of the human and mouse T cell receptor loci. Immunity 15, 337–349 (2001).
Rumfelt, L.L. et al. A shark antibody heavy chain encoded by a nonsomatically rearranged VDJ is preferentially expressed in early development and is convergent with mammalian IgG. Proc. Natl. Acad. Sci. USA 98, 1775–1780 (2001).
Diaz, M., Stanfield, R.L., Greenberg, A.S. & Flajnik, M.F. Structural analysis, selection, and ontogeny of the shark new antigen receptor (IgNAR): identification of a new locus preferentially expressed in early development. Immunogenetics 54, 501–512 (2002).
Havran, W.L. & Allison, J.P. Developmentally ordered appearance of thymocytes expressing different T-cell antigen receptors. Nature 335, 443–445 (1988).
Waters, W.R. & Harp, J.A. Cryptosporidium parvum infection in T-cell receptor (TCR)-α- and TCR-δ-deficient mice. Infect. Immun. 64, 1854–1857 (1996).
Hayday, A.C. γδ cells: a right time and a right place for a conserved third way of protection. Annu. Rev. Immunol. 18, 975–1026 (2000).
Ramsburg, E., Tigelaar, R., Craft, J. & Hayday, A. Age-dependent requirement for γδ T cells in the primary but not secondary protective immune response against an intestinal parasite. J. Exp. Med. 198, 1403–1414 (2003).
Rothstein, T.L. Cutting edge commentary: two B-1 or not to be one. J. Immunol. 168, 4257–4261 (2002).
Westerfield, M. The Zebrafish Book. A Guide for the Laboratory Use of Zebrafish (The University of Oregon Press, Eugene, Oregon, 1995).
Sleckman, B.P., Carabana, J., Zhong, X. & Krangel, M.S. Assessing a role for enhancer-blocking activity in gene regulation within the murine T-cell receptor α/δ locus. Immunology 104, 11–18 (2001).
Acknowledgements
We thank C. Rexroad and R. Athey of the US Department of Agriculture National Center for Cool and Cold Water Aquaculture Research Center for providing the rainbow trout cDNA clones. Supported in part by the National Institutes of Health (R01 AI08054).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Southern blot analysis of zebrafish genomic DNA with Cμ probe. (PDF 136 kb)
Supplementary Fig. 2
Classification of VH families. (PDF 216 kb)
Supplementary Fig. 3
Unrooted phylogenetic tree of VH sequences. (PDF 142 kb)
Supplementary Fig. 4
Alignment of the amino acid sequences encoded by Cμ and Cζ of zebrafish and other species. (PDF 152 kb)
Supplementary Fig. 5
A conserved sequence in the Cμ-Cδ intergenic region. (PDF 179 kb)
Supplementary Table 1
V, D and J gene segments in the zebrafish igh locus (PDF 125 kb)
Supplementary Table 2
Tools and databases used in the identification of igh genes (PDF 104 kb)
Supplementary Table 3
Comparison of CDR3 regions of μ and ζ cDNA and genomic cDNA (PDF 109 kb)
Supplementary Table 4
Primers (PDF 65 kb)
Rights and permissions
About this article
Cite this article
Danilova, N., Bussmann, J., Jekosch, K. et al. The immunoglobulin heavy-chain locus in zebrafish: identification and expression of a previously unknown isotype, immunoglobulin Z. Nat Immunol 6, 295–302 (2005). https://doi.org/10.1038/ni1166
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni1166
This article is cited by
-
Comparative analysis of the immune responses of CcIgZ3 in mucosal tissues and the co-expression of CcIgZ3 and PCNA in the gills of common carp (Cyprinus carpio L.) in response to TNP-LPS
BMC Veterinary Research (2024)
-
Molecular characterization of a new IgZ3 subclass in common carp (Cyprinus carpio) and comparative expression analysis of IgH transcripts during larvae development
BMC Veterinary Research (2021)
-
Immunoglobulins in teleosts
Immunogenetics (2021)